预约演示
更新于:2025-05-07

Beam Therapeutics, Inc.
更新于:2025-05-07
概览
标签
其他疾病
遗传病与畸形
血液及淋巴系统疾病
基因疗法
基因编辑
单克隆抗体
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 基因疗法 | 6 |
| 基因编辑 | 3 |
| 单克隆抗体 | 3 |
| 造血干细胞疗法 | 2 |
| CRISPR/Cas9 | 1 |
关联
20
项与 Beam Therapeutics, Inc. 相关的药物靶点 |
作用机制 G6PC基因调节剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床1/2期 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 CD7抑制剂 [+2] |
在研适应症 |
非在研适应症- |
最高研发阶段临床1/2期 |
首次获批国家/地区- |
首次获批日期- |
作用机制 血红蛋白刺激剂 [+2] |
非在研适应症- |
最高研发阶段临床1/2期 |
首次获批国家/地区- |
首次获批日期- |
7
项与 Beam Therapeutics, Inc. 相关的临床试验NCT06934382
A Phase 1 Study of Allogeneic Anti-CD7 CAR-T Cells (BEAM-201) in Relapsed/Refractory T-cell Acute Lymphoblastic Leukemia (T-ALL) or T-cell Lymphoblastic Lymphoma (T-LLy)
This will be a Phase 1, open-label study to evaluate the safety and efficacy of BEAM-201 in patients with R/R T-ALL or T-LLy. BEAM-201 is an allogeneic anti-CD7 CART therapy.
开始日期2025-04-29 |
申办/合作机构 |
NCT06735755
A Phase 1/2, Dose-Exploration Study to Evaluate the Safety and Efficacy of BEAM-301 in Patients With Glycogen Storage Disease Type Ia (GSDIa) Homozygous or Compound Heterozygous for the G6PC1 c.247C>T (p.R83C) Variant
This is a Phase 1/2, multicenter, open-label, single-ascending-dose study to evaluate the safety, tolerability, and efficacy of BEAM-301 in adult patients with GSDIa homozygous or compound heterozygous for the G6PC1 c.247C>T (p.R83C) variant and to determine the optimal biological dose.
开始日期2024-12-06 |
申办/合作机构 |
NCT06389877
A Phase 1/2 Dose-exploration and Dose-expansion Study to Evaluate the Safety and Efficacy of BEAM-302 in Adult Patients With Alpha-1 Antitrypsin Deficiency (AATD)-Associated Lung Disease and/or Liver Disease
This is a Phase 1/2, multicenter, open-label, dose-exploration (Phase 1) and dose-expansion (Phase 2) study to evaluate the safety, tolerability, PK/PD, and efficacy of BEAM-302 in adult patients with AATD-associated lung disease and/or liver disease and to determine the optimal biological dose (OBD).
开始日期2024-06-19 |
申办/合作机构 |
100 项与 Beam Therapeutics, Inc. 相关的临床结果
登录后查看更多信息
0 项与 Beam Therapeutics, Inc. 相关的专利(医药)
登录后查看更多信息
10
项与 Beam Therapeutics, Inc. 相关的文献(医药)2025-05-01·Journal of Immunotherapy
Combined CLEC2d Expression and CD58 Loss Mitigate Rejection of Allogeneic T Cells
Article
作者: Camblin, Adam J. ; Maldini, Colby R. ; Musenge, Faith M. ; Karaca, Cisem ; Coholan, Lindsey J. ; Leboeuf, Dominique ; White, Moriah L.
2024-11-05·Blood
CD117 Antibody Conditioning and Multiplex Base Editing Enable Rapid and Robust Fetal Hemoglobin Reactivation in a Rhesus Autologous Transplantation Model
作者: Gaudelli, Nicole ; Mondal, Nandini ; Krouse, Allen E ; McDonagh, Charlotte F ; Donahue, Robert E ; Kromer, Hugh ; Butt, Henna ; Engels, Theresa ; Austin, Wayne ; Harmon, Alexander ; Linde, N Seth ; Hong, So Gun ; Demirci, Selami ; Decker, Jeremy ; Uchida, Naoya ; London, Evan ; Hartigan, Adam J. ; Le, Anh ; Essawi, Khaled ; Sathish, Shruti ; Chu, S Haihua ; Wong, Jeffrey ; Tisdale, John ; Golomb, Justin ; Zhang, Kathy ; Bonifacino, Aylin
2024-11-05·Blood
Initial Results from the BEACON Clinical Study: A Phase 1/2 Study Evaluating the Safety and Efficacy of a Single Dose of Autologous CD34+ Base Edited Hematopoietic Stem Cells (BEAM-101) in Patients with Sickle Cell Disease with Severe Vaso-Occlusive Crises
作者: DiPersio, John ; Ayala, Ernesto ; Jaroscak, Jennifer J. ; Dalal, Jignesh ; Heeney, Matthew M. ; Gardner, Beth ; Ciaramella, Giuseppe ; Lin, Ling ; Kanter, Julie ; Joseney-Antoine, Marcelyne ; Chen, Guo ; Ziga, Edward D. ; Eapen, Mary ; Rifkin-Zenenberg, Stacey ; Ianniello, Leanne ; Sharma, Akshay ; Chockalingam, Priya S. ; Simon, Amy ; Alavi, Asif ; Minella, Alex C. ; Gupta, Ashish O. ; Thompson, Alexis ; Chen, Yinzhong ; Frangoul, Haydar ; Goyal, Sunita ; Hartigan, Adam J.
362
项与 Beam Therapeutics, Inc. 相关的新闻(医药)2025-05-05
·药时空
近年来,基因编辑技术不断发展,其中 CRISPR 碱基编辑技术备受关注。如今,Beam Therapeutics 和 Verve Therapeutics 两家公司基于该技术研发的候选药物在临床试验中取得了令人期待的成果,为相关疾病的治疗带来了新希望。碱基编辑技术崭露头角,临床成果初显九年前,哈佛大学大卫・刘(David Liu)团队开创了 CRISPR 碱基编辑技术。如今,首批基于该技术的疗法临床试验已陆续公布初步结果。上个月,Beam Therapeutics 宣布其候选药物 BEAM-302 能够纠正 α-1 抗胰蛋白酶缺乏症患者的致病突变。几周后,Verve Therapeutics 发布了 VERVE-102 基因疗法的 Ib 期试验初步结果,在接受最高剂量治疗的 4 名参与者中,该疗法平均将低密度脂蛋白胆固醇(LDL)水平降低了一半。威廉・布莱尔(William Blair)分析师萨米・科温(Sami Corwin)表示,碱基编辑技术前景乐观,目前已在 α-1 抗胰蛋白酶缺乏症、家族性高胆固醇血症和镰状细胞病这三种疾病的研究中降低了风险 。技术原理:对比传统 CRISPR,凸显独特优势传统的 CRISPR/Cas9 基因编辑技术通过引导 RNA(gRNA)与目标 DNA 片段互补配对,利用 Cas9 酶在目标位点制造双链断裂,再依靠细胞自身机制进行修复。但这一过程存在风险,可能产生脱靶断裂,且断裂修复不当会导致不必要的突变 。例如,目前唯一获批的 CRISPR/Cas9 疗法 Casgevy,用于治疗镰状细胞病时,需要先提取患者骨髓细胞在体外进行编辑,过程较为复杂 。而碱基编辑技术同样使用 gRNA,不过它利用的是经过改造的 Cas9 酶,不会制造双链断裂,而是直接对 DNA 碱基进行化学修饰,将腺苷(A)转变为鸟苷(G),或将胞嘧啶(C)转变为胸腺嘧啶(T) 。这种方式更加灵活,研究人员可以针对编码 DNA 链或其互补链进行编辑,实现更多种编辑可能,且被认为比传统 CRISPR/Cas9 更安全。Beam Therapeutics:多领域探索,成果备受关注Beam Therapeutics 由大卫・刘和 CRISPR 先驱张锋共同创立,专注于碱基编辑技术的治疗应用开发。公司研发项目广泛,涉及多种疾病和递送方式,包括针对镰状细胞病、β- 地中海贫血的体内和体外项目,以及针对 α-1 抗胰蛋白酶缺乏症和糖原贮积病 1a 的体内项目 。以 BEAM-302 为例,它旨在纠正导致 α-1 抗胰蛋白酶缺乏症的突变,通过脂质纳米颗粒将碱基编辑机制传递到肝脏 。在澳大利亚、新西兰、荷兰和英国开展的试验中,不同剂量组显示出剂量依赖性的效果,总 α-1 抗胰蛋白酶增加,有害的 Z-AAT 减少,且仅出现轻微不良事件 。高剂量组患者的总循环 α-1 抗胰蛋白酶水平足以让他们不再被视为患有进行性疾病 。此外,Beam 针对镰状细胞病研发的 BEAM-101 也独具特色,通过编辑细胞,不仅开启胎儿血红蛋白的生成,还调整了一种受体,使编辑后的细胞具有生存优势,有望避免传统治疗中化疗药物的副作用 。Verve Therapeutics:聚焦心血管疾病,挑战传统治疗Verve Therapeutics 起源于一次未成功的竞赛提案。首席执行官兼联合创始人塞克・卡蒂雷桑(Sek Kathiresan)受到自然突变降低胆固醇水平的启发,希望开发模仿这种保护机制的疗法 。公司在临床前研究中对碱基编辑和传统 CRISPR/Cas9 技术进行了测试,最终选择碱基编辑技术,主要是因为其安全性更高,编辑更精确 。目前,Verve 有三款碱基编辑候选药物处于临床试验阶段。VERVE-101 的试验曾因一名患者出现 3 级药物诱导的血小板减少症而暂停,不过公司认为这可能是脂质纳米颗粒载体导致的 。VERVE-102 则基于美国以外的初步临床数据获得了 IND 批准,在 Ib 期试验中显示出良好的耐受性和剂量依赖性反应,最高剂量组 LDL 胆固醇降低了 69% 。此外,针对纯合子家族性高胆固醇血症和其他疗法无效患者的 VERVE-201 也已进入临床试验阶段 。卡蒂雷桑认为,虽然基因编辑通常用于罕见病治疗,但对于高胆固醇这种常见且可治疗的疾病,基因编辑也能满足重要的未被满足的需求,有望实现长期降低 LDL 胆固醇,降低心脏病发作风险 。CRISPR 碱基编辑技术在 Beam 和 Verve 两家公司的推动下,在临床试验中取得了重要突破。但技术仍处于发展阶段,未来能否持续带来惊喜,为更多患者带来治愈希望,还有待进一步研究和观察。参考来源:https://www.biospace.com/drug-development/safer-crispr-base-editing-breaks-through-in-the-clinic-as-beam-verve-advance识别微信二维码,可添加药时空小编请注明:姓名+研究方向!
基因疗法临床研究临床结果
2025-05-05
iStock,
PhonlamaiPhoto
Beam Therapeutics and Verve Therapeutics have each built their lead candidates on a technique billed as a safer alternative to conventional CRISPR. Clinical results have so far been promising.
Nine years ago this month, researchers led by Harvard University’s David Liu
reported
on a new gene editing technique, a variation on CRISPR/Cas9 that they dubbed base editing. Today, preliminary but encouraging results are trickling out from clinical trials on the first base editing therapies.
Just last month, Beam Therapeutics
announced
that its candidate BEAM-302 can correct the disease-causing mutation in patients with alpha-1 antitrypsin deficiency. Weeks later, Verve Therapeutics
released
initial results
from a Phase Ib trial showing that its VERVE-102 gene therapy reduced LDL cholesterol levels by half, on average, in four participants who received the highest dose.
“I think the prospects and outlook [for base editing] look very positive right now,” Sami Corwin, an analyst at William Blair, told
BioSpace
. “We’ve seen it de-risked in three indications so far”—AATD, familial hypercholesterolemia and sickle cell disease (SCD).
Beam owns the intellectual property behind base editing and, so far, has only licensed it to Verve, Beam President Giuseppe Ciaramella told
BioSpace
. But that may change: “Once you show success, I’m sure there are going to be some other companies that are going to be trying to use base editing to essentially do their own product,” he said.
Base Editing Basics
With CRISPR/Cas9 gene editing, researchers construct a guide RNA (gRNA) complementary to a targeted stretch of DNA. The gRNA is tethered to a Cas9 enzyme that snips a double-stranded break at the target, which is then repaired by the cell’s own machinery. It’s a process that carries risks, including that off-target breaks could be created and that breaks may not be properly repaired, creating unwanted mutations.
“The problem is that [with] all of the nucleases, the only thing that they could do when they landed on the spot of DNA was making a double-stranded break, and . . . basically everything else is now left to the cell to sort out,” Ciaramella said. Double-stranded breaks “need to occur only under very controlled conditions when the cells are replicating because otherwise they can lead to all sorts of unpredictable editing outcomes.”
The only CRISPR/Cas9-based therapy approved so far, Vertex and CRISPR Therapeutics’ Casgevy, relies on extracting bone marrow cells from a patient with SCD and editing them outside the body to disable a gene that suppresses production of fetal hemoglobin. The cells are then expanded and transplanted back into the patient, where they produce red blood cells with fetal hemoglobin as a stand-in for the adult hemoglobin that’s mutated in the disease. Casgevy is also
approved
for transfusion-dependent beta thalassemia.
Base editing, like the original CRISPR/Cas9, uses a gRNA. However, it leverages a modified Cas9 that, rather than creating a double-strand break in the DNA, can chemically change an adenosine (A) base to a guanine (G) or a cytosine (C) to a thymine (T).
Researchers can target either the coding DNA strand or its complementary partner for these edits, Ciaramella noted, which opens up more possibilities. “Essentially, we can achieve four edits with the two base editors that we’ve got.”
Base editing is “actually much more versatile than maybe even people anticipated at the beginning,” when it was viewed mainly as a way to correct point mutations, Ciaramella added. For example, by making a change in the regulatory rather than the coding region of a gene, researchers can activate that gene.
Beam: Running the Gamut
After his group devised base editing, Liu and CRISPR pioneer Feng Zhang co-founded Beam specifically to develop therapeutics based on the technique. The company
emerged from stealth
in 2018 with $87 million in series A funding.
The company has not confined itself to particular kinds of disorders or means of delivery. It has both
in vivo
and
ex vivo
programs for SCD and beta thalassemia alongside
in vivo
programs in alpha-1 antitrypsin deficiency (AATD) and glycogen storage disease 1a (GSD1a). Beam also has collaborations with Apellis and Pfizer, about which it’s revealed little.
The candidate at the center of Beam’s March 10
announcement
, BEAM-302, corrects a mutation that causes the body to produce a form of the AAT protein known as Z-AAT rather than the functional form, M-AAT. Z-AAT has deleterious effects on both the lungs and liver. BEAM-302 delivers base editing machinery to the liver, where AAT is made, via lipid nanoparticles.
At sites in Australia, New Zealand, the Netherlands and the U.K., the company tested single intravenous doses of 15 mg, 30 mg and 60 mg in three-patient cohorts and found dose-dependent, durable increases in total AAT and decreases in Z-AAT, along with only minor adverse events. Total circulating AAT levels were high enough in the highest-dose group that those patients can be considered to no longer have progressive disease, Ciaramella said. He added, however, that these were only interim results and that the company is testing higher doses and considering redosing some patients.
The AATD data “confirmed that the sensitivity and efficacy of the base editing technology was there,” Corwin said. Since the treatment was well-tolerated even at higher doses, the readout also suggested that an inflammatory response to another base editing candidate
reported earlier
by Verve was due to the lipid nanoparticle vehicle rather than the base editing machinery itself, she added.
Other analysts also took note. Jefferies’ Michael Yee said in a note to investors that “safety has been a major area of focus in the context of issues from other LNP products for gene editing,” but that “BEAM safety looks clean.” He added that BEAM-302 “looks to have important benefits that differentiate from RNA editing and plasma-derived AAT therapy. . . . [W]e are seeing ‘correction’ of the disease with a one-time therapy.”
With another program, Beam is seeking to set itself apart in the sickle cell disease space. Both bone marrow transplants and Casgevy treatment require patients to undergo a course of the chemotherapy drug busulfan to open up space in the marrow for the new blood-making cells that will be introduced. The treatment has side effects, including the possibility of impaired fertility.
Beam’s
ex vivo
SCD candidate, BEAM-101, edits cells so that not only is fetal hemoglobin production switched on, but a receptor is tweaked to confer resistance to an antibody Beam devised. Rather than busulfan, patients are treated with the antibody, which starves unedited bone marrow cells of a growth factor needed to proliferate.
“We can confer advantages of survival to the edited cells over unedited cells simply by changing a single amino acid in this receptor,” Ciaramella said. Beam
presented
positive safety and efficacy data on the first seven patients treated with BEAM-101 in a U.S.-based study at the American Society of Hematology meeting in December.
Verve: A Focus on the Heart
Boston-based Verve’s origin story begins with a competition entry that wasn’t selected, said CEO and co-founder Sek Kathiresan. The 2016 contest, run by the American Heart Association, promised $75 million in funding for an idea to tackle coronary heart disease. Kathiresan, then a professor at Harvard Medical School and director of Massachusetts General Hospital’s Center for Genomic Medicine, said he took inspiration from natural mutations that keep cholesterol levels low and thus protect people carrying the mutations from developing heart disease. Kathiresan wanted to develop a therapy that mimicked natural protective mutations.
“The proposal was really: well, we understand what causes disease, which is cholesterol. We know a key way to stop this disease, which is to get cholesterol down and keep it down for a long time,” he told
BioSpace
.
When the application was turned down, he forged ahead anyway, gathering the investors and initial team needed to found Verve in 2018. The company tested both base editing and traditional CRISPR/Cas9 on target genes in preclinical research, and “what ultimately led us to choose base editing was the possibility of it being a fair bit safer because it did not cut DNA in both strands, but rather just tried to make the single spelling change A to G at one spot,” he said. The editing was also precise: “we’re not really seeing any off targets elsewhere in the genome.” But for another of its programs, Verve settled on a different gene editor because it found base editing wasn’t the most effective way forward.
The company now has three base editors in clinical trials, although the trial of VERVE-101 was
paused
last April after one patient developed grade 3 drug-induced thrombocytopenia within days of dosing. Verve, like Corwin, has since
attributed
those abnormalities to the lipid nanoparticle vector. Verve said at the time that it would instead prioritize VERVE-102, which like VERVE-101 treats heterozygous familial hypercholesterolemia, a genetic propensity to high cholesterol. VERVE-102 uses a different lipid nanoparticle from VERVE-101 that targets the
PCSK9
gene for inactivation in liver cells.
Last month, Verve
announced
IND clearance for VERVE-102 based on initial clinical data from outside the U.S. showing the therapy was well-tolerated. In its April 14
press release
, the company reported a dose-dependent response and no serious adverse events in a Phase Ib trial, with a maximum LDL cholesterol reduction of 69% in the highest-dose cohort. Verve is now testing a still-higher dose and plans to begin a Phase II trial in the second half of this year, per its announcement.
Meanwhile, VERVE-201, which targets the
ANGPTL3
gene in liver cells, is in trials for the homozygous version of the condition as well as for patients whose hypercholesterolemia is not responding to other therapies. The first patient was dosed with VERVE-201 in a Phase
Ib trial
in Canada and the U.K. in November 2024.
Getting this far has required breaking new ground in a few ways, Kathiresan said, from devising the mRNA and gRNA that would go into treatments to interfacing with the FDA. “We were actually the first to go in front of the U.S. FDA and propose
in vivo
gene editing in humans,” he notes, as other companies in the space have so far run their early trials in other countries, just as Verve did for VERVE-102.
Gene editing and other gene therapies are usually associated with rare disease, often those with no other effective treatments. High cholesterol is both common and treatable, but Kathiresan argued gene editing could nonetheless fill an important unmet need. “Theoretically, current medicines can lower LDL at a single time point [by] 40, 50, 60%. . . . But the challenge is discontinuation. A year out after starting any cardiovascular medication, a third to 50% of people have discontinued,” he said. In contrast, with base editing, “you basically lower LDL and then keep it low for your whole life. And that can really have a dramatic impact on your risk for heart attack.”
Ultimately, though, Kathiresan characterized base editing not as a silver bullet but as one tool companies can consider. “I think base editing will have a place. . . . But there will be other technologies as well that I think may be even better suited for a given target,” he said. “So I think right now the focus is really on the targets and the disease indications, and then, at least for us, finding the right gene editing tool for that target.”
临床结果临床1期基因疗法
2025-04-16
·药融圈
注:本文不构成任何投资意见和建议,以官方/公司公告为准;本文仅作医疗健康相关药物介绍,非治疗方案推荐(若涉及),不代表平台立场。任何文章转载需得到授权。近日,Verve Therapeutics 宣布其靶向PCSK9的基因编辑疗法VERVE-102的1b期临床试验初步数据令人鼓舞,显著降低LDL-C水平以及安全性良好。受此消息影响,Verve Therapeutics股价飙升45%,目前市值为2.89亿美元。早在一年前,Verve因安全问题放弃了一款碱基编辑候选药物VERVE-101的一期临床试验,原因是6名接受0.45mg/kg剂量治疗的患者中,有1例出现3级肝酶升高和1例3级血小板减少。Verve指出,脂质纳米颗粒(LNP)可能是导致这些不良事件的原因。VERVE-201也是一款体内单碱基编辑基因疗法,靶向肝脏中胆固醇和甘油三酯代谢的关键调节因子ANGPTL3,能够永久性关闭ANGPTL3的表达,以治疗动脉粥样硬化性心血管疾病。但是VERVE-201使用不同的脂质纳米颗粒递送系统,包括一个附加的靶向配体。其目标是更有效地进入肝细胞,以更低的剂量获得同样的效果,现已成为Verve公司PCSK9治疗的主导项目。疗效方面,Verve 分享了 14 名患有杂合子家族性高胆固醇血症和/或早发性冠状动脉疾病 (PCA) 的患者在 1b 期临床试验中接受 VERVE-102(分三种剂量) 治疗的结果:接受 0.45 mg/kg VERVE-102治疗的6名患者中,低密度脂蛋白胆固醇 (LDL-C) 降低了 41%,PCSK9 降低了 53%。这些数据是从第 28 天到最后一次随访的平均变化。在接受 0.45 mg/kg VERVE-101 治疗的6名患者中,Verve 发现LDL-C 平均降低了 42%。接受0.6mg/kg VERVE-102治疗的4名患者中,患者LDL-C和PCSK9平均降低了53%(最高降幅达69%)和60%。单次给药后,这种降低效果持续了18个月,这支持了Verve关于疗效持久性的说法。 相比之下,1名接受VERVE-101治疗的患者LDL-C水平降低了57%。目前,该公司正在招募接受0.7mg/kg VERVE-102治疗的患者。安全性方面,未发生与治疗相关的严重不良事件。仅1名患者出现2级输液相关反应,该反应为短暂性症状,经对乙酰氨基酚治疗后缓解。Verve表示,由目前试验数据推算,接受0.7mg/kg VERVE-102治疗的患者LDL-C水平将下降更多。数据表明,VERVE-102 超过了诺华公司已获批准的 siRNA Leqvio (inclisiran) 设定的 40% 至 50% 的 LDL-C 降低目标,并通过替换 LNP 纠正了 VERVE-101 的安全隐患。Verve 计划于 2025 年下半年完成 VERVE-102 的 2 期临床试验首例患者给药。目前,该 1b 期临床试验正在英国、加拿大、以色列、澳大利亚和新西兰招募患者,但 Verve 预计将在美国的试验点开展 2 期临床试验。FDA 最近 批准Verve 在美国开展该候选药物的研究。 Verve还计划在 2025 年下半年与礼来公司共享 VERVE-102选择权,礼来公司于 2023 年从 Beam Therapeutics 公司获得了选择权。届时,礼来和 Verve 将在美国联合商业化 VERVE-102,而 Verve 拥有其他地区的所有产品权利。目前全球共上市4款PCSK9抑制剂,分别是:依洛尤单抗(Evolocumab)由安进 / 安斯泰来开发,2015 年 3 月首次在日本获批,7 月在欧洲获批。是一种 PCSK9 单克隆抗体,通过与 LDLR 竞争结合 PCSK9,抑制 LDLR 的内吞、降解,增加肝细胞膜上与 LDL-C 结合的 LDLR 数量,促进肝脏对 LDL-C 的代谢,最终降低血浆 LDL-C 水平。阿利西尤单抗(Alirocumab)由再生元 / 赛诺菲开发,2015 年 7 月被美国 FDA 批准上市。同样是 PCSK9 单克隆抗体,作用机制与依洛尤单抗类似,获批适应症与依洛尤单抗相似,适用于治疗成人原发性高胆固醇血症(杂合子家族性和非家族性)或混合型血脂异常。英克司兰钠注射液(Inclisiran)由 Alnylam / 诺华开发,于 2020 年 12 月在欧盟首次获批。这是一种小干扰 RNA(siRNA)药物,基于 Alnylam 的 GalNAc 递送系统研发,可靶向编码 PCSK9 的 mRNA,通过直接 “沉默” 肝脏中 PCSK9 蛋白的产生发挥作用,能实现长效给药,前 3 个月完成 2 次注射后,每半年打一针,降低了患者的用药频率,提高了用药便利性。托莱西单抗由信达生物自主研发,2023 年 8 月 16 日获得国家药品监督管理局批准,是中国首个获批的本土自主研发 PCSK9 抑制剂,用于治疗原发性高胆固醇血症和混合型血脂异常。他汀类药物是当前作为LDL-C降低的首选药物。然而,临床中仍有部分患者出现他汀不耐受,并且在高危患者群体中,80% 患者无法通过现有他汀类药物充分控制其LDL-C 水平。目前临床应用的PCSK9抑制剂主要通过皮下注射给药,通常与他汀类药物(治疗高胆固醇血症的标准一线疗法)一起联用,也可单独用于他汀类药物不耐受人群的治疗。哈佛医学院的Eugene Braunwald医学博士指出,VERVE-102有可能改变心血管疾病治疗方式,从每日服药转变为单剂量治疗,以实现持续降低LDL-C的效果。研发管线参考资料:https://ir.vervetx.com/news-releases/news-release-details/verve-therapeutics-announces-positive-initial-data-heart-2-phase版权声明:本文转自细胞基因治疗前沿,如不希望被转载的媒体或个人可与我们联系,我们将立即删除会议推荐 | 点击查看详情植根上海、辐射全球,药融圈作为生物医药产业级战略平台,以"让智慧与人脉无界流动"为使命,构建覆盖药物研发、生产、流通、商业化的全产业链赋能体系。通过智能化云服务、精准资源链接、产业智库与生态社群四大核心引擎,打造中国医药价值流动的基础设施。在全球化与数字化双重浪潮下,药融圈正重新定义医药资源的流通范式--这里不仅是信息与人脉的枢纽站,更是催生产业变革的化学反应器。我们以中国为原点,编织全球医药智慧网络,让每一次精准链接都成为企业跨越式增长的催化剂。点点赞点分享点推荐
临床结果基因疗法siRNA临床2期临床1期
100 项与 Beam Therapeutics, Inc. 相关的药物交易
登录后查看更多信息
100 项与 Beam Therapeutics, Inc. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2025年10月13日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
2
9
临床前
临床2期
4
5
其他
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
BEAM-301 ( G6PC1 ) | 肝肾型糖原贮积病 更多 | 临床2期 |
BEAM-302 ( A1AT ) | α1-抗胰蛋白酶缺乏症 更多 | 临床2期 |
BEAM-201 ( CD7 ) | T细胞急性淋巴细胞白血病复发 更多 | 临床2期 |
BEAM-101 ( Hemoglobins x KLKB1 x TTR ) | β地中海贫血 更多 | 临床2期 |
ESCAPE CD117 ( HBG1 x HBG2 x c-Kit ) | 镰状细胞血症 更多 | 临床前 |
登录后查看更多信息
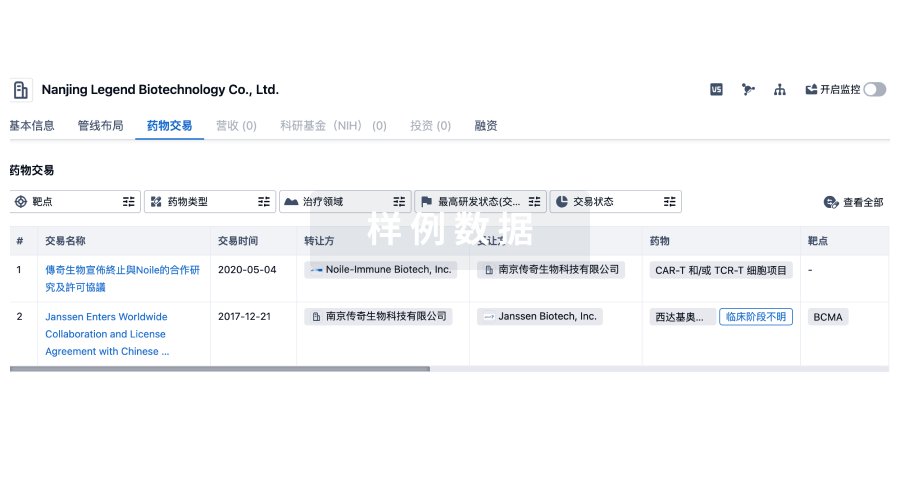
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
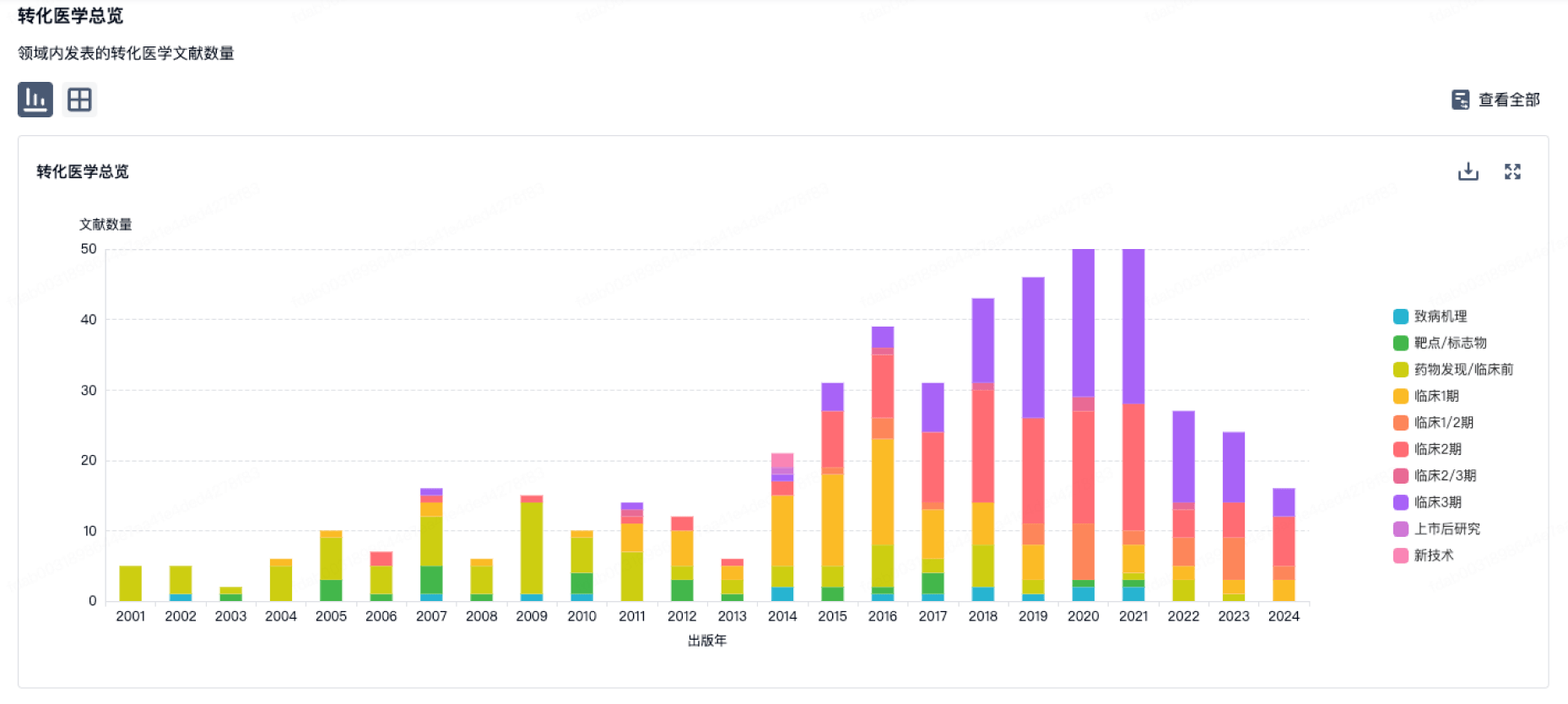
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
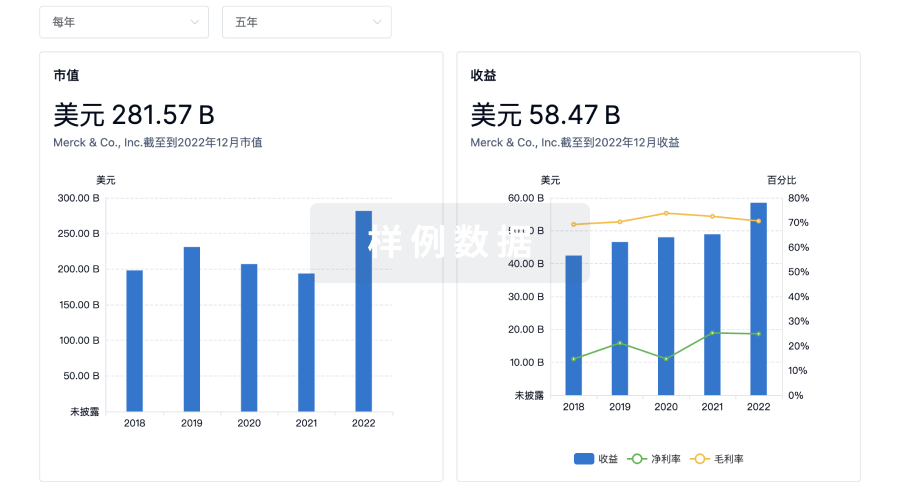
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
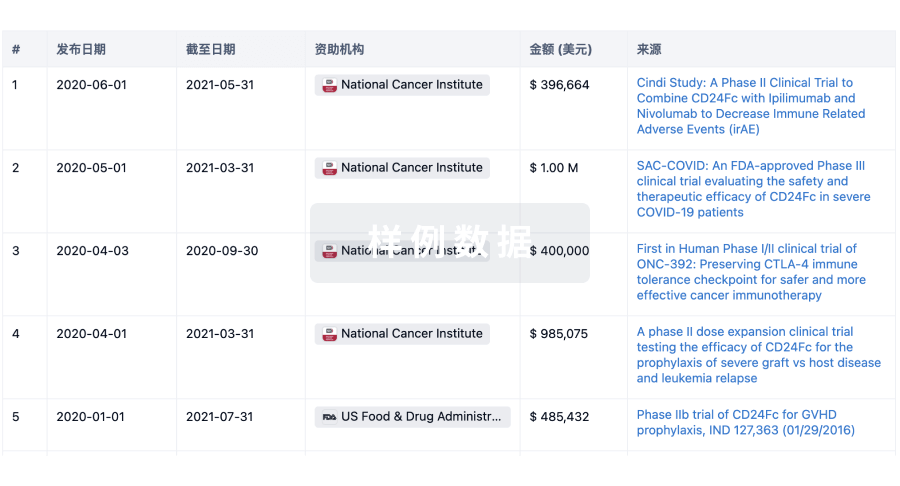
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
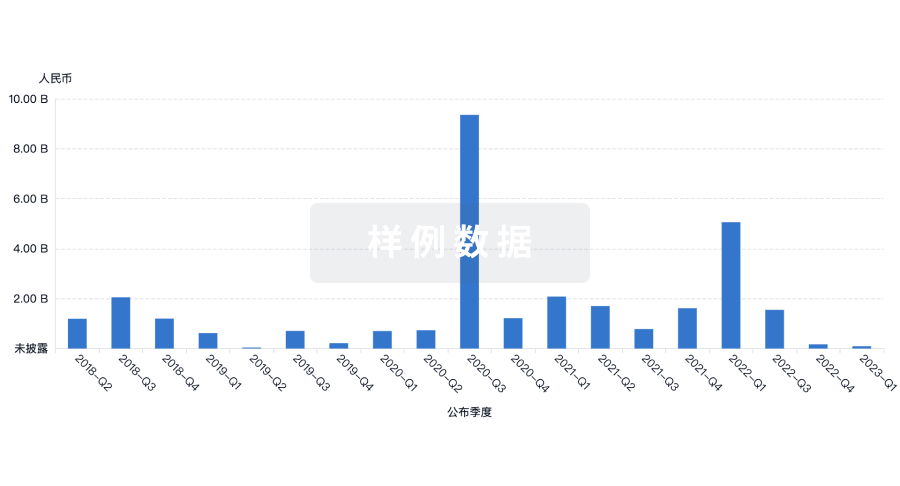
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
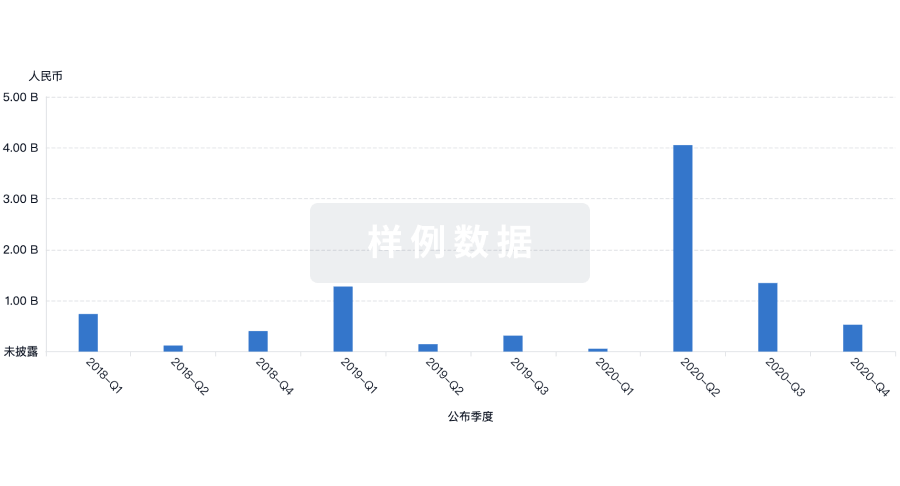
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
