预约演示
更新于:2025-05-16
Cinitapride Tartrate
酒石酸西尼必利
更新于:2025-05-16
概要
基本信息
药物类型 小分子化药 |
别名 4-Amino-N-(1-(3-cyclohexen-1-ylmethyl)-4-piperidyl)-2-ethoxy-5-nitrobenzamide、Cidine、Cinigest + [8] |
作用方式 激动剂 |
作用机制 5-HT1A receptor激动剂(5-羟色胺1A受体激动剂)、5-HT1B receptor激动剂(5-羟色胺1B受体激动剂)、5-HT1D receptor激动剂(5-羟色胺1D受体激动剂) |
非在研适应症- |
原研机构 |
非在研机构- |
最高研发阶段批准上市 |
最高研发阶段(中国)批准上市 |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C25H36N4O10 |
InChIKeyHVANMRCHFMTSEG-LREBCSMRSA-N |
CAS号96623-56-2 |
关联
6
项与 酒石酸西尼必利 相关的临床试验CTR20244516
酒石酸西尼必利片在中国成年健康受试者中的一项随机、开放、单次给药、空腹两制剂、两序列、两周期交叉和餐后两制剂、三序列、三周期部分重复交叉的生物等效性研究
主要研究目的:
考察空腹/餐后单次口服受试制剂酒石酸西尼必利片(规格:1 mg,江苏永安制药有限公司生产)与参比制剂酒石酸西尼必利片(商品名:Cidine(希笛尼)®,规格:1 mg,Almirall, S.A.持证),在中国成年健康人体的相对生物利用度,分析两种制剂的药代动力学参数,评价两制剂的生物等效性。
次要研究目的:
评价空腹/餐后单次口服受试制剂和参比制剂在中国成年健康受试者中的安全性。
开始日期2024-12-26 |
申办/合作机构 |
CTR20243852
中国健康受试者单次口服酒石酸西尼必利片(1mg)的开放、随机、单剂量、两制剂、单次给药、两序列、空腹两周期自身交叉设计、餐后四周期完全重复交叉设计的生物等效性试验
主要研究目的
研究空腹和餐后单次口服酒石酸西尼必利片受试制剂(规格:1mg,常州四药制药有限公司提供)与酒石酸西尼必利片参比制剂(Cidine®,规格1mg,Almirall,S.A.持证,常州四药制药有限公司提供)在中国健康受试者体内的药代动力学行为,评价空腹和餐后口服两种制剂的生物等效性。
次要研究目的
评价中国健康受试者空腹和餐后单次口服受试制剂(T)酒石酸西尼必利片和参比制剂(R)酒石酸西尼必利片(Cidine®)后的安全性。
开始日期2024-11-25 |
申办/合作机构 |
CTR20243845
酒石酸西尼必利片人体生物等效性研究
以中国健康受试者为试验对象,采用自身交叉对照的试验设计,测定安徽益普克医药科技发展有限公司持证的酒石酸西尼必利片给药后血浆中的西尼必利在健康受试者体内的血药浓度经时过程,估算相应的药代动力学参数,并以Almirall, S.A.持证生产的酒石酸西尼必利片【商品名:希笛尼(Cidine)®,规格:1mg(按西尼必利计)】为参比,评价制剂间的生物等效性,并观察酒石酸西尼必利片在中国健康受试者中的安全性。
开始日期2024-11-12 |
申办/合作机构 |
100 项与 酒石酸西尼必利 相关的临床结果
登录后查看更多信息
100 项与 酒石酸西尼必利 相关的转化医学
登录后查看更多信息
100 项与 酒石酸西尼必利 相关的专利(医药)
登录后查看更多信息
118
项与 酒石酸西尼必利 相关的文献(医药)2025-05-01·INTERNATIONAL JOURNAL OF BIOLOGICAL MACROMOLECULES
QbD-based formulation development and evaluation of lipid-cellulose-based extended-release multi-particulates of cinitapride and its in-silico PBPK modeling
Article
作者: Ali, Shehzad ; Saleem, Muhammad Talha ; Shoaib, Muhammad Harris ; Nasiri, Muhammad Iqbal ; Qazi, Faaiza ; Yousuf, Rabia Ismail ; Sikandar, Muhammad ; Saleem, Muhammad Umair ; Siddiqui, Fahad
The study aims to develop and optimize extended-release cinitapride matrix pellets using lipid wax glyceryl monostearate (GMS) and ethyl cellulose (EC) for treating functional dyspepsia and gastrointestinal reflux disease. Pellets were prepared using the extrusion spheronization technique by applying central composite design (CCD), keeping EC-7cps (matrix former), spheronization speed, and time as variables. The drug release, sphericity, and aspect ratio were quality parameters. The predicted optimized formulation was evaluated for chemical interaction, thermal stability, accelerated stability, and surface morphology. EC effectively controlled the overall Cinitapride release for up to 12 h, but burst release was reduced by sintering the matrix pellets. A direct relationship was observed between the sphericity and aspect ratio of the pellets with the spheronization speed and time. Fpred followed the Korsmeyer-Peppas model, demonstrating a non-fickian diffusion at all pH levels. The bioavailability of Fpred was compared with reference immediate release tablet by applying in silico PBPK modeling and comparable relative bioavailability. The PBPK predicted Cmax, Tmax, AUC0-t, and AUC0-∞ was 361.93 pg/mL, 6.88 h, 4567.8 pg/mL/h, and 5204.7 pg/mL/h, respectively. Lipid-cellulose-based extended-release Cinitapride matrix pellets were successfully prepared and can be effectively used as a single dose regimen for treating functional dyspepsia and gastrointestinal reflux disease.
2025-04-01·INTERNATIONAL JOURNAL OF BIOLOGICAL MACROMOLECULES
High-viscosity dietary fibers modulate gut microbiota and liver metabolism to prevent obesity in high-fat diet-fed mice
Article
作者: Nakano, Masataka ; Nagano, Takao ; Higashimura, Yasuki ; Nishiuchi, Takumi ; Lelo, Aaron Pambu
Obesity and metabolic disorders are rising global health concerns, emphasizing the need for effective dietary interventions. High-viscosity dietary fibers such as bacterial cellulose (BC) and guar gum (GG) have unique properties that may complement each other in modulating gut microbiota and metabolic health. This study investigates their effects in high-fat diet-fed mice. BC and GG increase Bacteroides, which degrade polysaccharides and produce short-chain fatty acids (SCFAs), supporting metabolic health. BC enhances bile acid excretion and enriches Faecalibaculum, Duncaniella, and Paramuribaculum, promoting gut barrier integrity and reducing inflammation, potentially improving bile acid turnover and lipid metabolism. GG more effectively increases butyrate production by enhancing butyrate-producing bacteria, such as Clostridium XIVa and Kineothrix, and promotes Bifidobacterium, strengthening anti-inflammatory effects and gut barrier function. Both fibers upregulate bile acid biosynthesis, but BC's non-fermentable nature leads to higher bile acid excretion, while GG's fermentation causes lower excretion and broader liver metabolic changes. Both fibers reduce body weight, fat accumulation, and cholesterol levels, highlighting their potential in managing obesity and metabolic disorders. The complementary effects of BC and GG underscore the importance of fiber diversity for targeted dietary strategies to improve metabolic health.
2024-04-01·Food research international (Ottawa, Ont.)
Wet-type grinder-treated okara modulates gut microbiota composition and attenuates obesity in high-fat-fed mice
Article
作者: Nishiuchi, Takumi ; Yano, Hiromi ; Watanabe, Chihiro ; Nagano, Takao ; Oyanagi, Eri
Wet-type grinder (WG) is a nanofiber technology used to atomize dietary fiber-rich materials. WG-treated okara (WGO) exhibits high dispersion and viscosity similar to those of viscous soluble dietary fibers. Here, we studied the effect of WGO supplementation on obesity and gut microbiota composition in high-fat diet (HFD)-fed mice. WGO intake suppressed body weight gain and fat accumulation, improved glucose tolerance, lowered cholesterol levels, and prevented HFD-induced decrease in muscle mass. WGO supplementation also led to cecum enlargement, lower pH, and higher butyrate production. The bacterial 16S ribosomal RNA genes (16S rDNA) were sequenced to determine the gut microbiota composition of the fecal samples. Sequencing of bacterial 16S rDNA revealed that WGO treatment increased the abundance of butyrate producer Ruminococcus and reduced the abundances of Rikenellaceae, Streptococcaceae, and Prevotellaceae, which are related to metabolic diseases. Metabolomics analysis of the plasma of WGO- and cellulose-treated mice were conducted using ultra-high-performance liquid chromatography-mass spectrometry. Metabolic pathway analysis revealed that the primary bile acid biosynthesis pathway was significantly positively regulated by WGO intake instead of cellulose. These results demonstrate that WG is useful for improving functional properties of okara to prevent metabolic syndromes, including obesity, diabetes, and dyslipidemia.
7
项与 酒石酸西尼必利 相关的新闻(医药)2025-03-01
点击蓝字
关注我们
编者按
钇90树脂微球选择性内放射治疗(Y90-SIRT)在原发性肝癌或转移性肝癌的应用日渐广泛。高能量β射线的穿透范围平均仅为2.5 mm,从而将其放射效果限制在肿瘤部位。然而,Y90微球的异常分布可能导致放射性胃肠炎、胃肠道溃疡、胆囊炎、放射性肺炎、放射性肝病等。美国肝病研究学会(AASLD)回顾了1990年至2016年期间发表的文献,并制定了相应指导意见。《国际肝病》特邀中国大陆首位Y90督导医师——清华大学附属北京清华长庚医院肝胆胰中心张琳教授对此进行盘点,以期为Y90-SIRT治疗过程中的安全性管理提供指导。
放射性肺炎
放射性肺炎是钇90治疗术后的一种少见并发症,可于SIRT后1-6个月发生,临床特征为限制性通气功能障碍和双侧肺放射性炎症,通常伴有劳力性呼吸困难和干咳。微球会通过肿瘤或肝硬化肝脏异常交通的肝静脉血管进入肺部。在Mapping过程中,于靶血管内注射锝-99m聚合白蛋白(MAA),并根据沉积在肺组织的MAA分数计算肺分流(LSF)。当LSF超过10%则具有放射性肺炎风险,当LSF超过20%则风险很高,不建议在SIRT后6个月内对肺部予以放疗。
若LSF超过10%,建议放射性树脂微球减量20%;若LSF超过15%,则减量40%;若LSF超过20%,则禁用SIRT。经验性数据确定了肺部30Gy的放疗上限,据此可防止放射性肺炎并发症。然而目前并无可靠方法评估到达肺部的辐射剂量,对SIRT导致肺损伤的概率分析较为欠缺,剂量学模型也不考虑功能性肺容积。在药物预防方面,暂无药物显示预防的积极作用。临床中需要关注LSF的重要性,较低的放疗剂量也会降低疗效。若患者LSF超过15%,则强烈建议更换可选方案或暂缓使用,而非降低Y90剂量。
若在SIRT后观察到肺较高的异常摄取,则建议检测血氧饱和度或每1-2周予以6分钟步行测试,在出现血氧饱和度下降或呼吸困难时予以胸部CT检查。任何LSF>10%的患者在SIRT后3个月内出现呼吸困难也应予以胸部CT检查。若观察到特征性的双侧对称性、界限不清的斑片状浑浊或毛玻璃样结节,相对外周/肺门不受累,且可排除感染性或心脏性原因或药物性因素,则应开始治疗。虽然目前暂无证据支持,但一般建议选择类固醇类作为主要治疗药物。对患者予以至少2周的类固醇治疗,再缓慢减量,可以基于经验适当吸氧或加用己酮可可碱。
表1. 放射性肺炎的预防、诊断减产和治疗专家建议
预防
检查
治疗
LSF>10%,减量20%
LSF>15%,减量40%
LSF>20%,禁用
若SIRT后2个月内出现低血氧、咳嗽或呼吸困难,特别是MAA显示LSF>10%,则予以胸部CT
经验性使用类固醇(甲泼尼龙 500 mg iv bid或泼尼松龙 60 mg po,缓慢停药)
基于MAA扫描确定肺组织放射剂量上限为30Gy(树脂或玻璃微球)
限制性肺功能测试以确定限制模式以及一氧化碳弥散能力水平改变
根据临床需要吸氧
LSF>15%,强烈建议采用替代疗法
基于诊断确定选择支气管肺泡灌洗和/或经支气管肺活检
LS>15%,强烈建议采用替代疗法
胃肠道溃疡
胃肠道溃疡也是SIRT的一种罕见并发症,于SIRT后2-6周出现,症状为急性上腹痛、恶心、呕吐、消化不良、厌食等。一般为多发性(0.5-2.0 cm),也可呈无法测量的弥散性。溃疡具有慢性、隐匿性特点,在治疗后症状仍可能持续数周,亦会并发幽门狭窄、出血、严重贫血、肠瘘、甚至死亡。部分无症状者在SIRT后数月乃至数年行胃活检时仍可发现Y90微球,对患者使用抗血管生成药物等胃肠道损伤药物后理论上也会引起此类症状。
临床中应加强对血管解剖结构认识来预防放射性微球进入胃或十二指肠。需全面评估肝脏动脉系统,发现可能导致肝外MAA摄取的动脉分支,使用弹簧圈栓塞预防,以降低放射性胃肠道溃疡风险。但因治疗期间对分枝血管的忽视、血管再通或血流动力学改变,均可能造成溃疡发生。临床中常用质子泵抑制剂(PPI)预防放射性胃肠道损伤,但欠缺循证支持,故不应作为常规方案。若开具PPI,则至少应维持8周。也可考虑前列腺素E2抑制剂米索前列醇等,但也缺乏循证支持。在预防性使用PPI时,若无明显症状则不需要胃镜检查。有时可考虑使用氨磷汀等放射防护剂。
在SIRT后4-8周出现持续性上腹痛患者,应进行上消化道内镜检查;出现贫血或进一步恶化者也鼓励尽早胃镜检查。即便是SIRT后未检出到异常摄取,也需要尽早检查。溃疡的发作模式及病变分布形式多样,若放疗部位愈合不良可能会限制定期活检。
放射性胃肠道溃疡对治疗反应不佳。轻中度症状可通过调节饮食、启用或增加以下药物剂量如PPI、胃保护剂硫酸铝、止吐药、镇痛药、胃动力药多潘立酮等。停用非甾体类抗炎药等具有胃肠道黏膜损伤作用的药物。严重时只有全肠外营养才能缓解,若症状持续数周,则营养性空肠造口术可提供长期症状控制,直至溃疡愈合。复发性胃肠道出血可通过制剂内镜治疗或选择性栓塞术治疗。极少数情况下出现的穿孔、出血则可能需要手术治疗。
表2. 放射性胃肠道溃疡的预防、诊断检查和治疗
预防
检查
治疗
常规检查或CTA,准确识别肝肠动脉分支,通过弹簧圈栓塞、更远端输注或血流再分布避免胃肠道溃疡
SIRT后4-8周出现上腹痛患者,尤其伴有恶心、食欲不振、贫血(即便MAA SPECT/CT 或 Y90 PET扫描未见肝外摄取)
大剂量PPI(奥美拉唑40mg qd或等效剂量的泮托拉唑、兰索拉唑等)、硫酸铝、止吐药、镇痛药、胃动力药(多潘立酮或西尼必利)
使用MAA检查:
高氯酸盐避免假阳性
输注前30 min准备MAA
输注部位与治疗一致
输注后1h使用SPECT成像
基于大体形态诊断(多发性或单发溃疡,伴有弥漫性黏膜炎,累积远端胃体、胃窦、幽门、十二指肠球)
避免使用非甾体类抗炎药或任何导致胃肠道溃疡药物
若怀疑假阳性,在禁用SIRT前复测MAA
除非考虑其他病因,否则不予活检
在更严重情况下,考虑全肠外营养或营养性空肠造瘘术
使用5%等渗葡萄糖作为微球输注剂
_
_
动脉内注射利多卡因或硝酸甘油可改善SIRT期间的血管痉挛
_
_
肝脏并发症
放射性肝病(REILD)是SIRT后4-8周出现的黄疸、腹水,不伴肿瘤进展或胆管堵塞,一般呈现胆红素升高(>3 mg/dL)、碱性磷酸酶和γ-谷氨酰转移酶升高、转氨酶几乎无变化。REILD与外照射引起的腹水不同,常表现为肝窦综合征。REILD不常见,一般出现在两类人群,一是由于肿瘤负荷较高在SIRT前接受过系统化疗的非肝硬化患者,二是肝硬化及肝功能储备较差者。
其他肝脏并发症如SIRT后非肝硬化性门静脉高压症非常罕见,可在数月或数年发生。临床中常见早期无症状脾大、伴或不伴血小板计数降低。SIRT后胆管树损毁包括胆管坏死或狭窄,相关报道很少。多数情况下无法排除肿瘤压迫胆管的可能。专家共识认为SIRT不一定是胆管狭窄的孤立原因。
在REILD预防方面,如总胆红素>2 mg/dL或有非瘤性腹水等肝功能储备较差者,不应视为SIRT潜在治疗人群。若总胆红素略低于该阈值,在SIRT前几周若出现数值快速升高,则应保守治疗。对于因脂肪变性、肝炎、肝硬化、既往系统化疗而肝功能储备较差的患者,应调整治疗剂量。一般常对树脂微球使用体表面积公式,对玻璃微球则根据肝脏体积计算,而体表面积公式可能高估了放射性活度,故需对患者个体化制定治疗方案。剂量学方法在临床的使用中受到多种因素影响,如何准确计算Y90微球在肿瘤与非肿瘤的摄取比暂未经过有效的临床验证。另外,序贯治疗可能会改善肝脏对SIRT的耐受性。预防性药物应用中,己酮可可碱、熊去氧胆酸、低分子肝素钠在机制上具有改善REILD的作用,但临床结果模棱两可,临床可考虑应用,但并无充足的循证证据予以支持。
对于SIRT后3个月内出现黄疸或腹水患者,无论是否伴有肝硬化,都需考虑REILD可能性。可对患者进行血液检查,包括总胆红素、碱性磷酸酶和转氨酶。发现总胆红素>3 mg/dL 后,应行腹部超声以排除胆管梗阻,并确认腹水的存在以及门静脉和肝静脉的通畅性,并测量肝功能。如果天冬氨酸转氨酶/丙氨酸转氨酶 (>1 000 IU/mL) 显著升高,应进行其他实验室检查。建议使用对比增强 CT 或核磁共振成像,以准确评估肝内和肝外肿瘤进展。如果根据检查结果诊断不明确,且病程稳定或恶化,强烈建议尽早进行肝活检,以指导治疗,需注意肝硬化患者不推荐活检。一旦确诊REILD,则需至少每周一次肝功能检查。
对于REILD,初始对症治疗包括利尿剂(低剂量螺内酯、呋塞米)。去纤苷已用于干细胞移植后的静脉闭塞性疾病,因此对于胆红素迅速升高或凝血功能改变的REILD者可考虑应用。若药物治疗无效,建议及时行TIPS术。
表3. REILD的预防、诊断检查和治疗
预防
检查
治疗
若总胆红素>2 mg/dL或存在非瘤性腹水,则为SIRT禁忌。需考虑个体晚期的胆红素变化
SIRT后3个月内出现黄疸和腹水的任何患者均应考虑REILD
初始予以利尿剂(螺内酯100 mg和/或呋塞米40 mg),根据肾功能和体重调节剂量
对于肝脏体积<1.5L的慢性肝病患者,如脂肪性肝炎、肝硬化,以及既往接受过多种系统化疗的患者,可以考虑降低推荐剂量
超声检查以确定胆管梗阻、腹水、门静脉/肝静脉通畅情况。血液检查以确定肝损伤和肝功能。如排除胆管梗阻,要求使用增强CT或更好的MRI进行前后对比,以排除肿瘤进展
如果肝功开始下降(如总胆红素≥ 6 mg/dL 且凝血酶原活性≤60% 或国际标准化比值 ≥1.4),则考虑静脉使用去纤苷,初始剂量为10 mg/kg。
在技术可行条件下保留尽可能多的肝段
确诊REILD后,至少每周一次肝功能检查。如果2周内持续存在或恶化,则考虑肝活检(非肝硬化患者)或咨询肝病相关医师
若药物治疗后效果不明显,建议及时行TIPS术
放射性胆囊炎
放射性胆囊炎是SIRT的罕见并发症之一,特征症状为上腹痛、恶心或呕吐、不适和偶尔发热,偶见胆囊壁增厚,一般于SIRT后20-30天随访扫描中最为突出。目前多为个例报道,未见胆囊中放射性同位素增加,行胆囊切除术后也未见严重并发症。
胆囊即使在影像学的改变但临床往往不伴有症状。应尽量避免Y90在胆囊动脉中的沉积,故建议将导管放置在胆囊动脉远端;如若不可行,则考虑在SIRT前行暂时性的胆囊动脉栓塞。不建议行胆囊动脉弹簧圈栓塞术和预防性胆囊切除术。影像学显示SIRT后6个月胆囊可能明显萎缩,但并无临床意义。不建议对具有胆囊炎风险患者预防性使用类固醇或抗生素进行经验性预防。
SIRT后4-6周出现持续性右上腹压痛和发热的任何患者都应怀疑放射性胆囊炎。超声、MRI、CT显示胆囊增厚(3.5 mm)伴有周围积液、壁内积气或水肿,有助于确诊。一般单纯的胆囊壁增厚、Murphy征不应判断为放射性胆囊炎。在难以确诊时,可能需要上消化道内窥镜检查区分胆囊炎、消化道溃疡。
对于放射性胆囊炎,保守治疗包括静脉补液和镇痛药,不推荐类固醇。对于发热、剧烈疼痛或影像学显示胆壁坏死或破裂体征患者,应考虑胆囊造瘘术和/或胆囊切除术。
表4. 放射性胆囊炎的预防、诊断检查和治疗
预防
检查
治疗
最大限度地将放射性微球输送到肿瘤供血血管,将微导管头端尽量远离胆囊动脉分支
SIRT后4-6周持续性右上腹压痛患者应怀疑放射性胆囊炎。若体格检查有相应体征,且超声、MRI、CT显示胆囊增厚(3.5 mm)伴有周围积液、壁内积气或水肿,有助于确诊
按需予以静脉补液及镇痛药
上述不可行的前提下,在SIRT前使用Gelfoam颗粒对胆囊动脉进行临时闭塞
重新评估胆囊动脉对辐射的摄取有助于确诊
对于发热、剧烈疼痛或影像学显示胆壁坏死或破裂体征患者,应考虑胆囊造瘘术(首选)和/或胆囊切除术
在决定手术前,应充分考量手术风险、肿瘤治疗及患者生存率之间的影响
其他潜在并发症
SIRT还会导致其他潜在并发症。如SIRT输注期间或输注后不久出现的栓塞后综合征,多与放射性微球数量相关。有些大型研究报道了腹痛、发热、恶心呕吐等症状。目前暂无强有力的证据表明支架置入、乳头切开或外科胆肠吻合手术后导致的感染风险增加,既往多将此作为SIRT的相对禁忌症。专家共识则认为,胆道系统的器械操作不应视为SIRT的禁忌症,但对于有脓毒症病史或胆道引流病史且未预防性使用抗生素者,不应予以SIRT。而对于临床所担忧的SIRT可能导致后续肝切除或肝移植并发症发生率增加的问题,目前循证显示是否SIRT差异并不大。
专家点评
虽然Y90-SIRT在国际上应用已有20余年,疗效与安全性已被证实,但我国起步较晚,目前临床应用也较局限。Y90-SIRT操作复杂,需要多学科团队协作,对操作者技术要求很高,一旦发生异常分布则可能导致放射性肺炎、放射性胃肠道溃疡、放射性肝病、放射性胆囊炎等严重并发症。因此,充分掌握其并发症的预防、检查及处理手段,将有助于促进Y90-SIRT在国内的推广和普及。
Y90-SIRT后常见的不良反应症状包括乏力、纳差,厌食、恶心、呕吐、腹痛、发热等比较少见,大多轻微且在一个月内缓解。由于不良反应缺乏特异性,治疗措施主要为对症处理,缓解患者不适。在临床中工作中需要高度重视Y90异常分布导致的严重并发症,尽量减少及发生,措施包括术前的全面评估、术中风险血管的处理、处方剂量的计算、治疗策略的选择,微球注射,术后管理等全过程均应详细规划、规范操作,及时处理术中突发状况等。一旦出现不良反应则应及时积极处理,以减少不良反应导致的严重后果,最终为患者带来疗效与安全的双重保障。
本共识主要针对的是欧美国家在钇90治疗并发症发生及处理的情况,但我们国家在钇90治疗过程中患者病情和治疗策略更加复杂,应更加关注和尽量杜绝钇90严重并发症的发生,这就需要严格钇90患者的筛选,精细化的评估,剂量计算,钇90治疗过程中做到精细化的操作及全病程患者管理策略的实施,从而保证钇90患者的疗效和安全性。
文献来源:Sangro B, Martínez-Urbistondo D, Bester L, et al. Prevention and treatment of complications of selective internal radiation therapy: Expert guidance and systematic review. Hepatology. 2017;66(3):969-982. doi:10.1002/hep.29207
点击“阅读原文”,查看文献原文
(来源:《肿瘤瞭望》编辑部)
声 明
凡署名原创的文章版权属《肿瘤瞭望》所有,欢迎分享、转载。本文仅供医疗卫生专业人士了解最新医药资讯参考使用,不代表本平台观点。该等信息不能以任何方式取代专业的医疗指导,也不应被视为诊疗建议,如果该信息被用于资讯以外的目的,本站及作者不承担相关责任。
放射疗法临床结果
2025-02-16
·药筛
统计每周仿制药一致性评价申报、上市申请(2.10-2.16)
1、仿制药上市申请获批药品名称企业名称分类受理号艾普拉唑肠溶片石药集团中诺药业(石家庄)有限公司4CYHS2302674贝前列素钠片浙江仙琚制药股份有限公司4CYHS2302791波生坦片Mylan S.p.A.5.2JYHS2200026波生坦片Mylan S.p.A.5.2JYHS2200025达格列净二甲双胍缓释片(Ⅰ)齐鲁制药有限公司4CYHS2303369达格列净二甲双胍缓释片(Ⅲ)齐鲁制药有限公司4CYHS2303370蛋白琥珀酸铁口服溶液山东则正医药技术有限公司;华益泰康药业股份有限公司4CYHS2404465多索茶碱注射液浙江昂利康制药股份有限公司;浙江创新生物有限公司4CYHS2303331二羟丙茶碱注射液海南紫程众投生物科技有限公司;河北凯威制药有限责任公司3CYHS2302596氟脲嘧啶注射液湖北民康药业集团有限公司;天津金耀药业有限公司3CYHS2301235氟脲嘧啶注射液湖北民康药业集团有限公司;天津金耀药业有限公司3CYHS2301234复方聚乙二醇电解质散(III)北京北陆药业股份有限公司4CYHS2302119复方托吡卡胺滴眼液四川禾亿制药有限公司4CYHS2302682加巴喷丁胶囊上海理想制药有限公司4CYHS2303160加巴喷丁胶囊上海理想制药有限公司4CYHS2303161甲苯磺酸艾多沙班片北京百奥药业有限责任公司;江苏永安制药有限公司4CYHS2302696甲苯磺酸艾多沙班片北京百奥药业有限责任公司;江苏永安制药有限公司4CYHS2302698甲苯磺酸艾多沙班片北京百奥药业有限责任公司;江苏永安制药有限公司4CYHS2302697拉考沙胺口服液南京斯泰尔医药科技有限公司;南京海纳制药有限公司4CYHS2301796利丙双卡因乳膏广州朗圣药业有限公司4CYHS2301791硫酸氨基葡萄糖胶囊江苏诚康药业有限公司4CYHS2303418硫酸氨基葡萄糖胶囊安徽正药医药科技有限公司;华益药业科技(安徽)有限公司4CYHS2302541硫酸镁钠钾口服用浓溶液山东齐都药业有限公司;浙江赛默制药有限公司4CYHS2302980硫酸羟氯喹片重庆博腾药业有限公司4CYHS2401175硫酸沙丁胺醇口服溶液安徽四环科宝制药有限公司3CYHS2301825罗沙司他胶囊湖南先施制药有限公司;湖南九典制药股份有限公司4CYHS2302841罗沙司他胶囊湖南先施制药有限公司;湖南九典制药股份有限公司4CYHS2302842洛索洛芬钠凝胶贴膏南京海纳医药科技股份有限公司;南京海纳制药有限公司4CYHS2300523孟鲁司特钠咀嚼片恒昌(广州)新药研究有限公司;南京海纳制药有限公司4CYHS2200431孟鲁司特钠咀嚼片恒昌(广州)新药研究有限公司;南京海纳制药有限公司4CYHS2200432乳果糖口服溶液重庆健能医药开发有限公司;四川健能制药有限公司4CYHS2302716乳果糖口服溶液重庆健能医药开发有限公司;四川健能制药有限公司4CYHS2302715乳果糖口服溶液湖北凯纳药业有限公司;湖北美思创药业有限公司4CYHS2302303乳果糖口服溶液济南康桥医药科技有限公司;湖南嘉恒制药有限公司4CYHS2301566乳果糖口服溶液济南康桥医药科技有限公司;湖南嘉恒制药有限公司4CYHS2301565乳果糖口服溶液济南康桥医药科技有限公司;湖南嘉恒制药有限公司4CYHS2301567乳果糖口服溶液澳诺(中国)制药有限公司4CYHS2301513羧甲司坦口服溶液北京沃邦医药科技有限公司;扬州市三药制药有限公司3CYHS2301670他氟前列素滴眼液苏州欧康维视生物科技有限公司4CYHS2303225头孢泊肟酯片山东润泽制药有限公司4CYHS2302538头孢唑肟钠氯化钠注射液苏州大冢制药有限公司4CYHS2303615维生素B12滴眼液南京海科瑞医药科技有限公司;亚邦医药股份有限公司4CYHS2302588吸入用乙酰半胱氨酸溶液洋浦京泰药业有限公司;内蒙古白医制药股份有限公司4CYHS2301051盐酸氨溴索口服溶液江苏开元药业有限公司;江苏贝佳制药有限公司4CYHS2302901盐酸奥洛他定滴眼液深圳万和制药有限公司4CYHS2302759盐酸托莫西汀胶囊山东新时代药业有限公司4CYHS2303484盐酸左沙丁胺醇雾化吸入溶液山东京卫制药有限公司3CYHS2303106盐酸左西替利嗪口服溶液重庆华邦胜凯制药有限公司;湖南科伦制药有限公司3CYHS2302308盐酸左西替利嗪口服溶液重庆华邦胜凯制药有限公司;湖南科伦制药有限公司3CYHS2302307依折麦布阿托伐他汀钙片(Ⅱ)北京福元医药股份有限公司4CYHS2303532注射用厄他培南钠安徽省先锋制药有限公司4CYHS2301511注射用哌拉西林钠湖南柏齐鹤药物研发有限公司;江苏海宏制药有限公司3CYHS2302423
注:灰色字体部分受理号结论为不批准。
2、一致性评补充申请获批药品名称企业名称受理号酒石酸美托洛尔注射液远大医药(中国)有限公司CYHB2450249头孢克洛分散片安徽安科恒益药业有限公司CYHB2450070头孢克洛分散片安徽安科恒益药业有限公司CYHB2450071
注:灰色字体部分受理号结论为不批准。
3、新增仿制药上市申请药品名称企业名称类型受理号阿帕他胺片浙江诺得药业有限公司4CYHS2500670艾曲泊帕乙醇胺干混悬剂郑州泰丰制药有限公司3CYHS2500700氨磺必利口服溶液南京恒道医药科技股份有限公司;南京集萃剂鼎医药有限公司3CYHS2500712奥卡西平片浙江恒研医药科技有限公司;浙江赛默制药有限公司4CYHS2500740奥卡西平片浙江恒研医药科技有限公司;浙江赛默制药有限公司4CYHS2500741奥硝唑注射液四环医药(郑州)有限公司3CYHS2500705奥硝唑注射液四环医药(郑州)有限公司3CYHS2500706比拉斯汀口腔崩解片珠海市汇通达医药有限公司;珠海润都制药股份有限公司3CYHS2500717玻璃酸钠滴眼液江苏万高药业股份有限公司4CYHS2500709玻璃酸钠滴眼液江苏万高药业股份有限公司4CYHS2500710布美他尼注射液江西人可医药科技有限公司;裕松源药业有限公司3CYHS2500701布美他尼注射液江西人可医药科技有限公司;裕松源药业有限公司3CYHS2500702布瑞哌唑片重庆圣华曦药业股份有限公司4CYHS2500688布瑞哌唑片重庆圣华曦药业股份有限公司4CYHS2500689布瑞哌唑片江苏康缘药业股份有限公司4CYHS2500650布瑞哌唑片江苏康缘药业股份有限公司4CYHS2500649布瑞哌唑片江苏康缘药业股份有限公司4CYHS2500651达格列净二甲双胍缓释片(Ⅰ)江苏安必生制药有限公司4CYHS2500703低钙腹膜透析液(乳酸盐-G1.5%)辰欣药业股份有限公司4CYHS2500679低钙腹膜透析液(乳酸盐-G2.5%)辰欣药业股份有限公司4CYHS2500680二硫化硒洗剂杭州和泽坤元药业有限公司;浙江鼎泰药业股份有限公司3CYHS2500720二羟丙茶碱注射液成都华宇制药有限公司3CYHS2500730夫西地酸乳膏杭州朱养心药业有限公司4CYHS2500711复方聚乙二醇(3350)电解质口服液广州一品红制药有限公司3CYHS2500723复方聚乙二醇电解质散(儿童型)山东益康药业股份有限公司3CYHS2500725复合磷酸氢钾注射液成都诺和晟鸿生物制药有限公司;广东星昊药业有限公司3CYHS2500728富马酸伏诺拉生片浙江恒研医药科技有限公司;浙江赛默制药有限公司4CYHS2500637富马酸伏诺拉生片浙江恒研医药科技有限公司;浙江赛默制药有限公司4CYHS2500636枸橼酸莫沙必利片福建省宝诺医药研发有限公司;浙江京新药业股份有限公司4CYHS2500695枸橼酸托法替布缓释片宁波美诺华天康药业有限公司4CYHS2500648琥珀酸亚铁片江西泰吉立生物医药科技有限公司;浙江赛默制药有限公司3CYHS2500733甲磺酸仑伐替尼胶囊山东鲁抗医药股份有限公司4CYHS2500742甲磺酸沙芬酰胺片常州四药制药有限公司4CYHS2500735甲磺酸沙芬酰胺片常州四药制药有限公司4CYHS2500736精氨酸布洛芬颗粒湖北亨迪药业股份有限公司4CYHS2500726精氨酸布洛芬颗粒湖北亨迪药业股份有限公司4CYHS2500727酒石酸西尼必利片安徽佳和药业有限公司4CYHS2500676克立硼罗软膏四川科伦药业股份有限公司;广东科泓药业有限公司4CYHS2500681利格列汀片江苏康缘药业股份有限公司4CYHS2500690联苯苄唑溶液安徽四环科宝制药有限公司3CYHS2500716硫酸艾沙康唑胶囊江苏奥赛康药业有限公司4CYHS2500665硫酸氨基葡萄糖胶囊山东康林医药有限公司;以岭万洲国际制药有限公司4CYHS2500661硫酸特布他林口服溶液山东致泰医药技术有限公司;安徽新世纪药业有限公司3CYHS2500699罗沙司他胶囊浙江贝得药业有限公司4CYHS2500662罗沙司他胶囊浙江贝得药业有限公司4CYHS2500663美阿沙坦钾片珠海润都制药股份有限公司4CYHS2500694美阿沙坦钾片陕西白鹿制药股份有限公司4CYHS2500683美阿沙坦钾片陕西白鹿制药股份有限公司4CYHS2500682美沙拉秦肠溶缓释胶囊成都倍特药业股份有限公司3CYHS2500660莫匹罗星软膏国药控股鑫烨(湖北)医药有限公司;湖北广济药业股份有限公司4CYHS2500704帕利哌酮缓释片北大医药股份有限公司4CYHS2500647帕利哌酮缓释片北大医药股份有限公司4CYHS2500646普拉洛芬滴眼液重庆药谷制药有限公司;杭州民生药业股份有限公司4CYHS2500666瑞舒伐他汀依折麦布片(I)齐鲁制药(海南)有限公司4CYHS2500652瑞舒伐他汀依折麦布片(I)四川科伦药业股份有限公司4CYHS2500684赛洛多辛胶囊江苏联环药业股份有限公司4CYHS2500739沙库巴曲缬沙坦钠片苏州中化药品工业有限公司4CYHS2500664沙库巴曲缬沙坦钠片山东普瑞曼药业有限公司4CYHS2500669山梨醇甘露醇冲洗液青岛奥瑞森医疗科技有限公司;青岛瑞奈医疗科技有限公司3CYHS2500672双黄连口服液云南楚雄天利药业有限公司4CYZS2500002双嘧达莫片四川天杏汇医药科技有限公司;成都第一制药有限公司3CYHS2500691他克莫司胶囊江西远超医药科技有限公司;江苏和晨药业有限公司4CYHS2500656他克莫司胶囊北京以岭生物工程技术有限公司4CYHS2500667他克莫司胶囊江西远超医药科技有限公司;江苏和晨药业有限公司4CYHS2500655他克莫司胶囊北京以岭生物工程技术有限公司4CYHS2500668头孢妥仑匹酯颗粒苏州盛达药业有限公司;苏州第三制药厂有限责任公司4CYHS2500696头孢妥仑匹酯颗粒苏州盛达药业有限公司;苏州第三制药厂有限责任公司4CYHS2500697乌美溴铵维兰特罗吸入粉雾剂正大天晴药业集团股份有限公司4CYHS2500678硝酸甘油注射液广州合和医药有限公司;中山万汉制药有限公司3CYHS2500719硝酸甘油注射液广州合和医药有限公司;中山万汉制药有限公司3CYHS2500718硝酸甘油注射液江西人可医药科技有限公司;四川宏明博思药业有限公司3CYHS2500713硝酸甘油注射液江西人可医药科技有限公司;四川宏明博思药业有限公司3CYHS2500714小儿法罗培南钠颗粒浙江普洛康裕制药有限公司4CYHS2500674盐酸艾司洛尔注射液广东南国药业有限公司4CYHS2500698盐酸奥普力农注射液山东健滨医药科技有限公司;西安汉丰药业有限责任公司3CYHS2500729盐酸半胱氨酸注射液北京柏雅联合药物研究所有限公司;广东星昊药业有限公司3CYHS2500743盐酸布比卡因注射液济南东方开元医药新技术有限公司;山东华鲁制药有限公司3CYHS2500708盐酸氮卓斯汀滴眼液武汉五景药业有限公司4CYHS2500721盐酸多巴胺注射液成都恒瑞制药有限公司;四川美大康佳乐药业有限公司3CYHS2500731盐酸多巴胺注射液成都恒瑞制药有限公司;四川美大康佳乐药业有限公司3CYHS2500732盐酸多奈哌齐口腔崩解片山东淄博新达制药有限公司4CYHS2500658盐酸甲氧氯普胺注射液江西人可医药科技有限公司;裕松源药业有限公司3CYHS2500693盐酸纳美芬注射液新乡市常乐制药有限责任公司3CYHS2500675盐酸纳美芬注射液新乡市常乐制药有限责任公司3CYHS2500673盐酸曲唑酮片江苏万高药业股份有限公司3CYHS2500737盐酸曲唑酮片江苏万高药业股份有限公司3CYHS2500738盐酸舍曲林口服溶液万特制药(海南)有限公司3CYHS2500677盐酸舍曲林口服溶液江苏润恒制药有限公司3CYHS2500685伊布替尼胶囊浙江诺得药业有限公司4CYHS2500657依折麦布阿托伐他汀钙片(Ⅰ)湖北午时药业股份有限公司4CYHS2500654依折麦布阿托伐他汀钙片(Ⅱ)湖北午时药业股份有限公司4CYHS2500653注射用硫酸多粘菌素B江苏迪赛诺制药有限公司3CYHS2500724注射用硫酸多粘菌素B广东金城金素制药有限公司;广东隆赋药业股份有限公司3CYHS2500722注射用乳糖酸红霉素海韬新药研发(江苏)有限公司;津药和平(天津)制药有限公司3CYHS2500734注射用头孢他啶阿维巴坦钠福安药业集团庆余堂制药有限公司4CYHS2500659注射用盐酸罗沙替丁醋酸酯湖北潜龙药业有限公司3CYHS2500692左卡尼汀注射液广州泽盛药业科技有限公司;珠海润都制药股份有限公司4CYHS2500687左卡尼汀注射液广州泽盛药业科技有限公司;珠海润都制药股份有限公司4CYHS2500686左西孟旦注射液仁合益康汇泽药业河北有限公司;河北仁合益康药业有限公司4CYHS2500707左氧氟沙星滴眼液安徽康融药业有限公司4CYHS2500715左乙拉西坦注射用浓溶液江苏润恒制药有限公司4CYHS2500671
4、新增一致性评价补充申报药品名称企业名称受理号多潘立酮片浙江昂利康制药股份有限公司CYHB2550104伏格列波糖片北京星昊医药股份有限公司CYHB2550110甲硝唑氯化钠注射液四川太平洋药业有限责任公司CYHB2550096甲硝唑氯化钠注射液四川美大康佳乐药业有限公司CYHB2550093门冬氨酸钾注射液辽宁药联制药有限公司CYHB2550094门冬氨酸钾注射液辽宁药联制药有限公司CYHB2550095头孢地尼片江苏亚邦强生药业有限公司CYHB2550107头孢地尼片江苏亚邦强生药业有限公司CYHB2550108维生素B12注射液山东齐都药业有限公司CYHB2550106维生素B6注射液江苏神龙药业有限公司;江苏吴中医药集团有限公司苏州制药厂CYHB2550109盐酸林可霉素注射液辰欣药业股份有限公司CYHB2550102叶酸片石药集团欧意药业有限公司CYHB2550105脂肪乳注射液(C14-24)西安力邦制药有限公司CYHB2550114脂肪乳注射液(C14-24)西安力邦制药有限公司CYHB2550112脂肪乳注射液(C14-24)西安力邦制药有限公司CYHB2550111脂肪乳注射液(C14-24)西安力邦制药有限公司CYHB2550113脂肪乳注射液(C14-24)四川科伦药业股份有限公司CYHB2550098脂肪乳注射液(C14-24)四川科伦药业股份有限公司CYHB2550099脂肪乳注射液(C14-24)四川科伦药业股份有限公司CYHB2550097注射用头孢他啶福建省福抗药业股份有限公司CYHB2550101注射用头孢他啶福建省福抗药业股份有限公司CYHB2550100注射用盐酸地尔硫卓苏州二叶制药有限公司CYHB2550103
数据来源:摩熵医药
一致性评价上市批准
2025-02-12
·信狐药迅
本周药品注册受理数据,分门别类呈现,一目了然。(1.20-2.9)
新药上市申请药品名称企业注册分类受理号法赞雷生片江苏海岸药业有限公司1CXHS2500011法赞雷生片江苏海岸药业有限公司1CXHS2500010LJ01003长春澜江医药科技有限公司2.2CXHS2500009LJ01003长春澜江医药科技有限公司2.2CXHS2500008BCM857博志研新泰州药物技术有限公司2.2CXHS2500015BCM857博志研新泰州药物技术有限公司2.2CXHS2500014洛索洛芬钠滴眼液润尔眼科药物(广州)有限公司2.2;2.4CXHS2500016海博麦布阿托伐他汀钙片瀚晖制药有限公司2.3CXHS2500013海博麦布阿托伐他汀钙片瀚晖制药有限公司2.3CXHS2500012重组带状疱疹疫苗(CHO细胞)绿竹生物制药(珠海市)有限公司1.2CXSS2500027冻干人用狂犬病疫苗(无血清Vero细胞)科兴(成都)生物制品有限公司2.2CXSS2500017舒地胰岛素注射液江苏恒瑞医药股份有限公司1CXSS2500011舒地胰岛素注射液江苏恒瑞医药股份有限公司1CXSS2500010伏欣奇拜单抗注射液长春金赛药业有限责任公司1CXSS2500015古莫奇单抗注射液中山康方生物医药有限公司1CXSS2500020古莫奇单抗注射液中山康方生物医药有限公司1CXSS2500021
新药临床申请药品名称企业注册分类受理号硫酸心舒帕莎片山东齐都药业有限公司1CXHL2500102硫酸心舒帕莎片山东齐都药业有限公司1CXHL2500101FY101注射液北京福元医药股份有限公司1CXHL2500100HBW-012336胶囊成都海博为药业有限公司1CXHL2500099HRS-1167片江苏恒瑞医药股份有限公司1CXHL2500111HRS-1167片江苏恒瑞医药股份有限公司1CXHL2500110HRS-1167片江苏恒瑞医药股份有限公司1CXHL2500109TQB3912片正大天晴药业集团股份有限公司1CXHL2500108TQB3912片正大天晴药业集团股份有限公司1CXHL2500107BIOS-0618片杭州百诚医药科技股份有限公司1CXHL2500104BIOS-0618片杭州百诚医药科技股份有限公司1CXHL2500103INS018_055片英矽智能科技(上海)有限公司1CXHL2500120HX9428片福建海西新药创制股份有限公司1CXHL2500117HX9428片福建海西新药创制股份有限公司1CXHL2500116AC699片冰洲石生物科技(上海)有限公司1CXHL2500145ABP-671片杭州新元素药业有限公司1CXHL2500144ABP-671片杭州新元素药业有限公司1CXHL2500143AC699片冰洲石生物科技(上海)有限公司1CXHL2500142ABP-671片杭州新元素药业有限公司1CXHL2500141ABP-671片杭州新元素药业有限公司1CXHL2500140ABP-671片杭州新元素药业有限公司1CXHL2500139TQB3019胶囊正大天晴药业集团股份有限公司1CXHL2500138TM471-1胶囊知微生物医药(新乡)有限公司1CXHL2500137TM471-1胶囊知微生物医药(新乡)有限公司1CXHL2500136SNK-2726注射液施能康医药科技(苏州)有限公司1CXHL2500135TT-9701片1CXHL2500129TT-9701片1CXHL2500128INR102注射液云核医药(天津)有限公司1CXHL2500121伊匹乌肽滴眼液温州益承睛湛生物科技有限公司1CXHL2500146HRS-9813胶囊广东恒瑞医药有限公司1CXHL2500147HS-20118片江苏豪森药业集团有限公司1CXHL2500153HS-20118片江苏豪森药业集团有限公司1CXHL2500152HS-20118片江苏豪森药业集团有限公司1CXHL2500151SK-09片广州康臣药业有限公司1CXHL2500150SK-09片广州康臣药业有限公司1CXHL2500149HRS9531注射液福建盛迪医药有限公司1CXHL2500157HRS9531注射液福建盛迪医药有限公司1CXHL2500156HRS9531注射液福建盛迪医药有限公司1CXHL2500155UA026片祐森健恒生物医药(上海)有限公司1CXHL2500176UA026片祐森健恒生物医药(上海)有限公司1CXHL2500175HRS-5635注射液福建盛迪医药有限公司1CXHL2500174SYH2068注射液石药集团中诺药业(石家庄)有限公司1CXHL2500173注射用BH259珠海贝海生物技术有限公司1CXHL2500168FNX010片成都凡诺西生物医药科技有限公司1CXHL2500167FNX010片成都凡诺西生物医药科技有限公司1CXHL2500166HS-10542胶囊江苏豪森药业集团有限公司1CXHL2500165HS-10542胶囊江苏豪森药业集团有限公司1CXHL2500164HS-10542胶囊江苏豪森药业集团有限公司1CXHL2500163HS-10542胶囊江苏豪森药业集团有限公司1CXHL2500162HS-10542胶囊江苏豪森药业集团有限公司1CXHL2500161HS-10542胶囊江苏豪森药业集团有限公司1CXHL2500160HS-10529片江苏豪森药业集团有限公司1CXHL2500182HS-10529片江苏豪森药业集团有限公司1CXHL2500181TJ0113胶囊杭州天玑济世生物科技有限公司1CXHL2500180HIF-117胶囊沈阳三生制药有限责任公司1CXHL2500179HIF-117胶囊沈阳三生制药有限责任公司1CXHL2500178HIF-117胶囊沈阳三生制药有限责任公司1CXHL2500177CMC-2310口溶膜复星医药(徐州)有限公司2.2CXHL2500113CMC-2310口溶膜复星医药(徐州)有限公司2.2CXHL2500112BCM995上海云晟研新生物科技有限公司2.2CXHL25001051640A干混悬剂微多安药业(山东)有限公司2.2CXHL2500115双环醇缓释片海南广升誉制药有限公司2.2CXHL2500119注射用WE010上海宝龙药业股份有限公司2.2CXHL2500133OLP-210片深圳奥礼生物科技有限公司2.2CXHL2500132OLP-210片深圳奥礼生物科技有限公司2.2CXHL2500131OLP-210片深圳奥礼生物科技有限公司2.2CXHL2500130司美格鲁肽注射液海南中和药业股份有限公司2.2CXHL2500127司美格鲁肽注射液海南中和药业股份有限公司2.2CXHL2500126司美格鲁肽注射液2.2CXHL2500125司美格鲁肽注射液2.2CXHL2500124儿童专用维生素D3口服冻干片2.2CXHL2500123儿童专用维生素D3口服冻干片2.2CXHL2500122DM2065贴北京德默高科医药技术有限公司2.2CXHL2500148YHAX北京阳光诺和药物研究股份有限公司2.2CXHL2500158AQF直服颗粒深圳珐玛易药品科技有限公司2.2CXHL2500169TTYP01片上海澳宗生物科技有限公司2.2CXHL2500159KEM2111凝胶贴膏深圳珐玛易药品科技有限公司2.2;2.4CXHL2500172KEM2111凝胶贴膏深圳珐玛易药品科技有限公司2.2;2.4CXHL2500171KEM2111凝胶贴膏深圳珐玛易药品科技有限公司2.2;2.4CXHL2500170LH-2103胶囊江苏联环药业股份有限公司2.3CXHL2500114HK241雾化吸入溶液南京华盖制药有限公司2.4CXHL2500106注射用多西他赛(白蛋白结合型)石药集团中奇制药技术(石家庄)有限公司2.4CXHL2500118注射用西罗莫司(白蛋白结合型)石药集团中奇制药技术(石家庄)有限公司2.4CXHL2500134TAP-1503 乳膏上海泽德曼医药科技有限公司2.4CXHL2500154YKYY026注射液杭州天龙药业有限公司1.2CXSL2500058AFN1204注射液合肥阿法纳生物科技有限公司1.2CXSL2500074AFN1213注射液合肥阿法纳生物科技有限公司1.2CXSL2500101YKYY025注射液杭州天龙药业有限公司1.2CXSL250013020价肺炎球菌多糖结合疫苗云南沃森生物技术股份有限公司2.2CXSL2500061注射用7MW3711迈威(上海)生物科技股份有限公司1CXSL2500057XS228细胞注射液士泽生物医药(苏州)有限公司1CXSL2500055BBM-D101注射液信中医药科技(北京)有限公司1CXSL2500060依苏帕格鲁肽α注射液广州银诺医药集团股份有限公司1CXSL2500070依苏帕格鲁肽α注射液广州银诺医药集团股份有限公司1CXSL2500069依苏帕格鲁肽α注射液广州银诺医药集团股份有限公司1CXSL2500068SHR-4658注射液上海恒瑞医药有限公司1CXSL2500067XS411细胞注射液士泽生物医药(苏州)有限公司1CXSL2500066IBR854细胞注射液英百瑞(杭州)生物医药有限公司1CXSL2500065IBR854细胞注射液英百瑞(杭州)生物医药有限公司1CXSL2500064异体人再生胰岛注射液(E-islet 01)智新浩正(上海)医药科技有限公司1CXSL2500063注射用TQB2922正大天晴药业集团南京顺欣制药有限公司1CXSL2500062BNT324映恩生物制药(苏州)有限公司1CXSL2500073BNT327百欧恩泰(上海)医药有限公司1CXSL2500072TriCellPr-AC07人脐带间充质干细胞注射液奥辰生物(云南)有限公司1CXSL2500071HiCM388注射液艾尔普再生医学科技(深圳)有限公司1CXSL2500077UTAA09注射液博生吉医药科技(苏州)有限公司1CXSL2500075FT-017注射液芳拓生物科技(上海)有限公司1CXSL2500078RY_SW01细胞注射液江苏睿源生物技术有限公司1CXSL2500082RGL-270注射液广州瑞领医药有限公司1CXSL2500080注射用DR30206浙江道尔生物科技有限公司1CXSL2500079注射用ASKG915江苏奥赛康生物医药有限公司1CXSL2500092JMT601注射液上海津曼特生物科技有限公司1CXSL2500091PT217凡恩世制药(北京)有限公司1CXSL2500090SHR-1819注射液广东恒瑞医药有限公司1CXSL2500086注射用QLS31905齐鲁制药有限公司1CXSL2500084ADG126注射液天演药业(苏州)有限公司1CXSL2500089人羊膜间充质干细胞注射液源品细胞生物科技集团有限公司1CXSL2500087注射用ZG005苏州泽璟生物制药股份有限公司1CXSL2500083人自体多克隆调节性T细胞注射液上海赛尔欣生物医疗科技有限公司1CXSL2500094重组抗人EGFR抗体/IL-10双特异Fc融合蛋白注射液施慧达药业集团(吉林)有限公司1CXSL2500098注射用SIM0686上海先祥医药科技有限公司1CXSL2500097BNT327百欧恩泰(上海)医药有限公司1CXSL2500096BNT326百欧恩泰(上海)医药有限公司1CXSL2500095AK139注射液中山康方生物医药有限公司1CXSL2500100AK139注射液中山康方生物医药有限公司1CXSL2500099注射用BL-M09D1成都百利多特生物药业有限责任公司1CXSL2500103注射用KH815成都康弘生物科技有限公司1CXSL2500104注射用BL-B01D1成都百利多特生物药业有限责任公司1CXSL2500117注射用BL-B01D1成都百利多特生物药业有限责任公司1CXSL2500120注射用BL-M11D1成都百利多特生物药业有限责任公司1CXSL2500105注射用RC108荣昌生物制药(烟台)股份有限公司1CXSL2500108注射用LBL-024南京维立志博生物科技股份有限公司1CXSL2500107SHR-4849注射液苏州盛迪亚生物医药有限公司1CXSL2500111注射用LBL-024南京维立志博生物科技股份有限公司1CXSL2500112LBL-007注射液南京维立志博生物科技股份有限公司1CXSL2500113注射用BL-B01D1成都百利多特生物药业有限责任公司1CXSL2500114注射用BL-B01D1成都百利多特生物药业有限责任公司1CXSL2500119XS228细胞注射液士泽生物医药(苏州)有限公司1CXSL2500121注射用BL-B01D1成都百利多特生物药业有限责任公司1CXSL2500116注射用BL-B01D1成都百利多特生物药业有限责任公司1CXSL2500118注射用BL-B01D1成都百利多特生物药业有限责任公司1CXSL2500115注射用LBL-024南京维立志博生物科技股份有限公司1CXSL2500106注射用ZG006苏州泽璟生物制药股份有限公司1CXSL2500122人牙囊间充质干细胞注射液成都世联康健生物科技有限公司1CXSL2500129注射用KJ103上海宝济药业股份有限公司1CXSL2500128DNTH103注射液元羿生物科技(上海)有限公司1CXSL2500124IBI3020信达生物医药科技(杭州)有限公司1CXSL2500123NCR201注射液中盛溯源(广州)生物科技有限公司1CXSL2500126HWS116注射液武汉人福创新药物研发中心有限公司1CXSL2500125纳基奥仑赛注射液广东合源生物医药有限公司2.2CXSL2500076阿得贝利单抗注射液上海盛迪医药有限公司2.2CXSL2500081阿得贝利单抗注射液上海盛迪医药有限公司2.2CXSL2500093QL2107注射液齐鲁制药有限公司2.2CXSL2500085SHR-8068注射液苏州盛迪亚生物医药有限公司2.2CXSL2500088卡度尼利单抗注射液康方药业有限公司2.2CXSL2500102SHR-8068注射液苏州盛迪亚生物医药有限公司2.2CXSL2500110阿得贝利单抗注射液上海盛迪医药有限公司2.2CXSL2500109DJ-01注射剂苏州德进生物制药有限公司2.4CXSL2500059
仿制药申请药品名称企业注册分类受理号富马酸酮替芬口服溶液辽宁海一制药有限公司3CYHS2500385枸橼酸西地那非干混悬剂山东朗诺制药有限公司3CYHS2500378吡格列酮二甲双胍片烟台正方制药有限公司3CYHS2500376阿齐沙坦氨氯地平片江西施美药业股份有限公司3CYHS2500375依帕司他片沈阳永大制药股份有限公司3CYHS2500372硫酸特布他林口服溶液成都医路融曜药物科技有限公司3CYHS2500371复合磷酸氢钾注射液赤峰源生药业有限公司3CYHS2500370微量元素注射液4-II合肥市未来药物开发有限公司3CYHS2500368盐酸丁螺环酮片北京诚济制药股份有限公司3CYHS2500367黄体酮注射液上海惠永制药有限公司3CYHS2500365磷酸奥司他韦颗粒柏士顿(海南)医疗科技有限公司3CYHS2500364硫代硫酸钠注射液赤峰源生药业有限公司3CYHS2500363氨己烯酸口服溶液用散上海奥科远制药有限公司3CYHS2500416注射用雷替曲塞四川百利药业有限责任公司3CYHS2500415坎地氢噻片华益药业科技(安徽)有限公司3CYHS2500414盐酸左沙丁胺醇雾化吸入溶液复星医药(徐州)有限公司3CYHS2500413硫酸妥布霉素注射液华夏生生药业(北京)有限公司3CYHS2500412硫代硫酸钠注射液甘正(海南)生物药业有限公司3CYHS2500410吡格列酮二甲双胍片(15mg/500mg)宁波美舒医药科技有限公司3CYHS2500407吡拉西坦注射液北京四环科宝制药股份有限公司3CYHS2500406舒必利注射液海南卓科制药有限公司3CYHS2500405舒必利注射液海南卓科制药有限公司3CYHS2500404坎地氢噻片海南药谷云生物科技有限公司3CYHS2500403苯磺酸氨氯地平口服溶液湖南科伦制药有限公司3CYHS2500402苯磺酸氨氯地平口服溶液湖南科伦制药有限公司3CYHS2500401倍他米松磷酸钠注射液陕西顿斯制药有限公司3CYHS2500399铝镁匹林片(II)苏州特瑞药业股份有限公司3CYHS2500398头孢地尼干混悬剂苏州第三制药厂有限责任公司3CYHS2500396头孢地尼干混悬剂苏州第三制药厂有限责任公司3CYHS2500395注射用硫酸多黏菌素B桂林南药股份有限公司3CYHS2500394氯化钾注射液石家庄四药有限公司3CYHS2500440氯化钙注射液广东君康药业有限公司3CYHS2500439聚普瑞锌口腔崩解片海南金瑞宝医药科技有限公司3CYHS2500438盐酸肾上腺素注射液江苏润恒制药有限公司3CYHS2500435硝普钠注射液中润药业有限公司3CYHS2500433利多卡因喷雾剂重庆香雪医药有限公司3CYHS2500430利多卡因喷雾剂重庆香雪医药有限公司3CYHS2500429盐酸纳美芬注射液中山万汉制药有限公司3CYHS2500421左氧氟沙星注射液海南斯达制药有限公司3CYHS2500444左氧氟沙星注射液海南斯达制药有限公司3CYHS2500443普瑞巴林口崩片北京星昊盈盛药业有限公司3CYHS2500442头孢克肟胶囊南京瑞捷医药科技有限公司3CYHS2500468己酮可可碱注射液楚雄和创药业有限责任公司3CYHS2500467铝镁匹林片(II)福建省闽东力捷迅药业股份有限公司3CYHS2500461氯化钾颗粒南京黄龙生物科技有限公司3CYHS2500460左氧氟沙星注射液成都华宇制药有限公司3CYHS2500459左氧氟沙星注射液成都华宇制药有限公司3CYHS2500458氯化钾颗粒南京黄龙生物科技有限公司3CYHS2500457盐酸多巴胺注射液郑州维先医药科技有限公司3CYHS2500451盐酸拉贝洛尔注射液上海朝晖药业有限公司3CYHS2500448盐酸拉贝洛尔注射液上海朝晖药业有限公司3CYHS2500447利塞膦酸钠片北京福元医药股份有限公司3CYHS2500475盐酸溴己新注射液福安药业集团宁波天衡制药有限公司3CYHS2500493奥美拉唑碳酸氢钠干混悬剂(I)安徽四环科宝制药有限公司3CYHS2500492奥美拉唑碳酸氢钠干混悬剂(Ⅱ)安徽四环科宝制药有限公司3CYHS2500491布立西坦口服溶液江西科睿药业有限公司3CYHS2500486对乙酰氨基酚注射液北京柏雅联合药物研究所有限公司3CYHS2500485生理氯化钠溶液海南爱科制药有限公司3CYHS2500484生理氯化钠溶液海南爱科制药有限公司3CYHS2500483碳酸氢钠注射液安徽新世纪药业有限公司3CYHS2500481盐酸利多卡因注射液河北冀辉医药科技有限公司3CYHS2500494多种微量元素注射液(Ⅳ)北京柏雅联合药物研究所有限公司3CYHS2500521琥珀酸多西拉敏片南京济群医药科技股份有限公司3CYHS2500515琥珀酸多西拉敏片南京济群医药科技股份有限公司3CYHS2500514布瑞哌唑口腔崩解片南京海纳制药有限公司3CYHS2500513布瑞哌唑口腔崩解片南京海纳制药有限公司3CYHS2500512布瑞哌唑口腔崩解片南京海纳制药有限公司3CYHS2500511秋水仙碱片浙江浙北药业有限公司3CYHS2500508洛索洛芬钠口服溶液吉林省辉南长龙生化药业股份有限公司3CYHS2500506乙酰半胱氨酸片广东泰来润至医药科技有限公司3CYHS2500502氨溴特罗口服溶液吉林省益浦生物科技有限公司3CYHS2500499骨化三醇口服溶液湖北欣泽霏药业有限公司3CYHS2500498盐酸溴己新口服溶液湖南九典制药股份有限公司3CYHS2500497氯化钾口服溶液南京恒道医药科技股份有限公司3CYHS2500535氯化钾口服溶液合肥华威药业有限公司3CYHS2500533色甘酸钠滴眼液武汉新瑞医药科技有限公司3CYHS2500545醋酸钙口服溶液华益药业科技(安徽)有限公司3CYHS2500553缩宫素注射液苏州天马医药集团天吉生物制药有限公司3CYHS2500550碳酸氢钠/葡萄糖2mmol/l钾电解质血滤置换液成都青山利康药业股份有限公司3CYHS2500549阿仑膦酸钠口服溶液河北仁合益康药业有限公司3CYHS2500547磷/碳酸氢钠血滤置换液(4mmol/L钾、1.25mmol/L钙)成都青山利康药业股份有限公司3CYHS2500563布美他尼注射液山东方明药业集团股份有限公司3CYHS2500562乙酰半胱氨酸注射液江苏润恒制药有限公司3CYHS2500560盐酸丙卡特罗颗粒浙江百代医药科技有限公司3CYHS2500579硝普钠注射液海南爱科制药有限公司3CYHS2500578盐酸纳美芬注射液朗天药业(湖北)有限公司3CYHS2500576复方聚乙二醇(3350)电解质散河南福森药业有限公司3CYHS2500569坎地氢噻片福建大谱生物医药有限公司3CYHS2500568硫酸特布他林口服溶液苏州华健瑞达医药技术有限公司3CYHS2500567盐酸拉贝洛尔片山东华铂凯盛生物科技有限公司3CYHS2500583盐酸拉贝洛尔片山东华铂凯盛生物科技有限公司3CYHS2500582倍他米松磷酸钠注射液上海锦帝九州药业(安阳)有限公司3CYHS2500612重酒石酸去甲肾上腺素注射液江苏润恒制药有限公司3CYHS2500607重酒石酸去甲肾上腺素注射液江苏润恒制药有限公司3CYHS2500606乙酰半胱氨酸口服液安徽茂康药业有限公司3CYHS2500605米诺地尔搽剂北京京丰制药(山东)有限公司3CYHS2500598米诺地尔搽剂北京京丰制药(山东)有限公司3CYHS2500597盐酸溴己新注射液湖南赛隆药业(长沙)有限公司3CYHS2500595吸入用盐酸丙卡特罗溶液海南斯达制药有限公司3CYHS2500594吸入用盐酸丙卡特罗溶液海南斯达制药有限公司3CYHS2500593氯化钾注射液山西普恒制药有限公司3CYHS2500588二硫化硒洗剂广东恒健制药有限公司3CYHS2500586甲氨蝶呤片海南卓科制药有限公司3CYHS2500619维生素B6注射液海韬新药研发(江苏)有限公司3CYHS2500618微量元素注射液4-I合肥市未来药物开发有限公司3CYHS2500635注射用吲哚菁绿海南倍特药业有限公司3CYHS2500640注射用硫酸多黏菌素B福建省福抗药业股份有限公司3CYHS2500632复方聚乙二醇(3350)电解质口服溶液海南广升誉制药有限公司3CYHS2500631尿素[13C]呼气试验药盒深圳市中核海得威生物科技有限公司3CYHS2500629盐酸舍曲林口服浓缩液江苏润恒制药有限公司3CYHS2500685盐酸舍曲林口服浓缩液万特制药(海南)有限公司3CYHS2500677盐酸纳美芬注射液新乡市常乐制药有限责任公司3CYHS2500675盐酸纳美芬注射液新乡市常乐制药有限责任公司3CYHS2500673山梨醇甘露醇冲洗剂青岛奥瑞森医疗科技有限公司3CYHS2500672美沙拉秦肠溶缓释胶囊成都倍特药业股份有限公司3CYHS2500660磷酸芦可替尼片哈尔滨三联药业股份有限公司4CYHS2500362磷酸芦可替尼片哈尔滨三联药业股份有限公司4CYHS2500361磷酸芦可替尼片哈尔滨三联药业股份有限公司4CYHS2500360奈妥匹坦帕洛诺司琼胶囊浙江康莱特药业有限公司4CYHS2500384注射用帕瑞昔布钠广州一品红制药有限公司4CYHS2500383布瑞哌唑片山东京卫制药有限公司4CYHS2500382布瑞哌唑片山东京卫制药有限公司4CYHS2500381布瑞哌唑片山东京卫制药有限公司4CYHS2500380瑞舒伐他汀依折麦布片(I)江苏利泰尔药业有限公司4CYHS2500379地屈孕酮片广州朗圣药业有限公司4CYHS2500377沙库巴曲缬沙坦钠片四川百利药业有限责任公司4CYHS2500374硫酸镁钠钾口服用浓溶液浙江国镜药业有限公司4CYHS2500373ω-3脂肪酸乙酯90软胶囊山西鑫煜制药股份有限公司4CYHS2500369利格列汀片迪沙药业集团有限公司4CYHS2500366苯磺酸左氨氯地平片成都欣捷高新技术开发股份有限公司4CYHS2500393苯磺酸左氨氯地平片成都欣捷高新技术开发股份有限公司4CYHS2500392帕拉米韦氯化钠注射液广东大翔制药有限公司4CYHS2500391帕拉米韦氯化钠注射液广东大翔制药有限公司4CYHS2500390乳果糖口服溶液湖北迪喜药业有限公司4CYHS2500389磷酸芦可替尼片常州制药厂有限公司4CYHS2500388磷酸芦可替尼片常州制药厂有限公司4CYHS2500387磷酸芦可替尼片常州制药厂有限公司4CYHS2500386富马酸伏诺拉生片陕西白鹿制药股份有限公司4CYHS2500418富马酸伏诺拉生片陕西白鹿制药股份有限公司4CYHS2500417西甲硅油乳剂湖南华纳大药厂股份有限公司4CYHS2500411磷酸芦可替尼片成都倍特药业股份有限公司4CYHS2500409乳酸钠林格注射液重庆安诺生生命科技有限公司4CYHS2500408脂肪乳氨基酸(17)葡萄糖(19)注射液费森尤斯卡比华瑞制药有限公司4CYHS2500400达芦那韦片江苏艾迪药业股份有限公司4CYHS2500397乌帕替尼缓释片浙江花园药业有限公司4CYHS2500441乳果糖口服溶液湖南千金湘江药业股份有限公司4CYHS2500437阿奇霉素胶囊悦康药业集团股份有限公司4CYHS2500436阿奇霉素干混悬剂湖南九典制药股份有限公司4CYHS2500434米库氯铵注射液成都市海通药业有限公司4CYHS2500432米库氯铵注射液成都市海通药业有限公司4CYHS2500431环硅酸锆钠散海南先声药业有限公司4CYHS2500428环硅酸锆钠散海南先声药业有限公司4CYHS2500427盐酸达泊西汀片浙江莎普爱思药业股份有限公司4CYHS2500426盐酸达泊西汀片浙江莎普爱思药业股份有限公司4CYHS2500425盐酸曲唑酮缓释片浙江赛默制药有限公司4CYHS2500424盐酸曲唑酮缓释片浙江赛默制药有限公司4CYHS2500423盐酸尼卡地平注射液南京瑞捷医药科技有限公司4CYHS2500422罗沙司他胶囊上海医药集团青岛国风药业股份有限公司4CYHS2500420罗沙司他胶囊上海医药集团青岛国风药业股份有限公司4CYHS2500419阿司匹林肠溶片石家庄市华新药业有限责任公司4CYHS2500445达格列净片浙江宏元药业股份有限公司4CYHS2500471达格列净片浙江宏元药业股份有限公司4CYHS2500470阿达帕林凝胶杭州和泽坤元药业有限公司4CYHS2500469艾拉莫德片安徽省先锋制药有限公司4CYHS2500466富马酸喹硫平缓释片国药集团工业有限公司4CYHS2500465氟伐他汀钠缓释片浙江高跖医药科技股份有限公司4CYHS2500464阿帕他胺片江西科睿药业有限公司4CYHS2500463糠酸氟替卡松鼻用喷雾剂齐鲁制药有限公司4CYHS2500462钆特酸葡胺注射液通用电气药业(上海)有限公司4CYHS2500456钆特酸葡胺注射液通用电气药业(上海)有限公司4CYHS2500455钆特酸葡胺注射液通用电气药业(上海)有限公司4CYHS2500454盐酸氮斯汀滴眼液成都普什制药有限公司4CYHS2500453卡前列素氨丁三醇注射液甘肃成纪生物药业有限公司4CYHS2500452甲苯磺酸艾多沙班片华北制药股份有限公司4CYHS2500450甲苯磺酸艾多沙班片华北制药股份有限公司4CYHS2500449屈螺酮炔雌醇片四川科伦药业股份有限公司4CYHS2500446米库氯铵注射液江苏润恒制药有限公司4CYHS2500478米库氯铵注射液江苏润恒制药有限公司4CYHS2500477培哚普利氨氯地平片(III)四川国为制药有限公司4CYHS2500476克立硼罗软膏山东百诺医药股份有限公司4CYHS2500474美阿沙坦钾片山西德元堂药业有限公司4CYHS2500473美阿沙坦钾片山西德元堂药业有限公司4CYHS2500472利奥西呱片南京黄龙生物科技有限公司4CYHS2500489利奥西呱片南京黄龙生物科技有限公司4CYHS2500488利奥西呱片南京黄龙生物科技有限公司4CYHS2500487硫酸氨基葡萄糖胶囊河南福森药业有限公司4CYHS2500482环孢素滴眼液(III)广州大光制药有限公司4CYHS2500480比索洛尔氨氯地平片宁夏康亚药业股份有限公司4CYHS2500479骨化三醇注射液河南泰丰生物科技有限公司4CYHS2500496盐酸莫西沙星片北大医药股份有限公司4CYHS2500495达格列净二甲双胍缓释片(III)山东鲁抗医药股份有限公司4CYHS2500523达格列净二甲双胍缓释片(I)山东鲁抗医药股份有限公司4CYHS2500522羧基麦芽糖铁注射液博瑞制药(苏州)有限公司4CYHS2500520苯磺酸阿曲库铵注射液扬州中宝药业股份有限公司4CYHS2500519磷酸芦可替尼片杭州中美华东制药有限公司4CYHS2500518美阿沙坦钾片北京民康百草医药科技有限公司4CYHS2500517美阿沙坦钾片北京民康百草医药科技有限公司4CYHS2500516美阿沙坦钾片浙江高跖医药科技股份有限公司4CYHS2500510美阿沙坦钾片浙江华润三九众益制药有限公司4CYHS2500509药用炭颗粒河北长天药业有限公司4CYHS2500507盐酸罗哌卡因注射液湖北津药药业股份有限公司4CYHS2500505盐酸罗哌卡因注射液湖北津药药业股份有限公司4CYHS2500504吸入用七氟烷河北仁合益康药业有限公司4CYHS2500503洛索洛芬钠贴剂石家庄探索医药科技有限公司4CYHS2500501洛索洛芬钠贴剂石家庄探索医药科技有限公司4CYHS2500500盐酸倍他洛尔滴眼液苏州欧康维视生物科技有限公司4CYHS2500543恩扎卢胺软胶囊正大天晴药业集团股份有限公司4CYHS2500542荧光素钠注射液苏州朗科生物技术股份有限公司4CYHS2500541甲苯磺酸艾多沙班片江苏华阳制药有限公司4CYHS2500540甲苯磺酸艾多沙班片江苏华阳制药有限公司4CYHS2500539非奈利酮片重庆华邦制药有限公司4CYHS2500538非奈利酮片重庆华邦制药有限公司4CYHS2500537布洛芬混悬液山西同达药业有限公司4CYHS2500536盐酸决奈达隆片安徽艾立德制药有限公司4CYHS2500534盐酸乙哌立松片大桐制药(中国)有限责任公司4CYHS2500532帕利哌酮缓释片辰欣药业股份有限公司4CYHS2500531聚多卡醇注射液福安药业集团庆余堂制药有限公司4CYHS2500530聚多卡醇注射液福安药业集团庆余堂制药有限公司4CYHS2500529聚多卡醇注射液福安药业集团庆余堂制药有限公司4CYHS2500528富马酸伏诺拉生片江苏华阳制药有限公司4CYHS2500527富马酸伏诺拉生片江苏华阳制药有限公司4CYHS2500526富马酸伏诺拉生片四川梓橦宫药业股份有限公司4CYHS2500525富马酸伏诺拉生片四川梓橦宫药业股份有限公司4CYHS2500524非奈利酮片江苏德源药业股份有限公司4CYHS2500544丁溴东莨菪碱注射液四川沃翔生物医药有限公司4CYHS2500546盐酸胺碘酮注射液海南卓科制药有限公司4CYHS2500556注射用阿莫西林钠克拉维酸钾山东安信制药有限公司4CYHS2500555注射用阿莫西林钠克拉维酸钾山东安信制药有限公司4CYHS2500554磷酸奥司他韦胶囊江西泰吉立生物医药科技有限公司4CYHS2500552泊沙康唑注射液河南泰丰生物科技有限公司4CYHS2500551磷/碳酸氢钠血滤置换液成都青山利康药业股份有限公司4CYHS2500548甲磺酸伊马替尼片山东鲁抗医药股份有限公司4CYHS2500566甲磺酸伊马替尼片山东鲁抗医药股份有限公司4CYHS2500565氧(液态)洛阳天泽气体有限公司4CYHS2500564维生素B12滴眼液河北龙海药业有限公司4CYHS2500561地诺孕素片株洲千金药业股份有限公司4CYHS2500559富马酸伏诺拉生片陕西天宇制药有限公司4CYHS2500558富马酸伏诺拉生片陕西天宇制药有限公司4CYHS2500557伏格列波糖片上海现代制药股份有限公司4CYHS2500581伏格列波糖片上海现代制药股份有限公司4CYHS2500580瑞舒伐他汀依折麦布片(I)山东百诺医药股份有限公司4CYHS2500577奥氮平片山东普瑞曼药业有限公司4CYHS2500575丙戊酸钠口服溶液郑州维先医药科技有限公司4CYHS2500574尼莫地平片石家庄四药有限公司4CYHS2500573硝苯地平控释片天津力生制药股份有限公司4CYHS2500572硫酸氨基葡萄糖胶囊辽宁康博士制药有限公司4CYHS2500571托伐普坦片石家庄市华新药业有限责任公司4CYHS2500570熊去氧胆酸胶囊四川迪菲特药业有限公司4CYHS2500613富马酸伏诺拉生片济南高华制药有限公司4CYHS2500611富马酸伏诺拉生片济南高华制药有限公司4CYHS2500610左氧氟沙星滴眼液盈科瑞(珠海金湾)制药有限公司4CYHS2500609注射用醋酸西曲瑞克山西威奇达光明制药有限公司4CYHS2500608醋酸加尼瑞克注射液苏州东瑞制药有限公司4CYHS2500604达格列净片苏州第壹制药有限公司4CYHS2500603阿戈美拉汀片河南福森药业有限公司4CYHS2500602布瑞哌唑片石药集团欧意药业有限公司4CYHS2500601布瑞哌唑片石药集团欧意药业有限公司4CYHS2500600布瑞哌唑片石药集团欧意药业有限公司4CYHS2500599富马酸贝达喹啉片沈阳红旗制药有限公司4CYHS2500596吸入用布地奈德混悬液山东泰合医药科技有限公司4CYHS2500592布地奈德福莫特罗吸入粉雾剂(II)上海新黄河制药有限公司4CYHS2500591玻璃酸钠滴眼液国药集团三益药业(芜湖)有限公司4CYHS2500590玻璃酸钠滴眼液国药集团三益药业(芜湖)有限公司4CYHS2500589沙美特罗替卡松吸入气雾剂长风药业股份有限公司4CYHS2500587富马酸伏诺拉生片江苏诺泰澳赛诺生物制药股份有限公司4CYHS2500585富马酸伏诺拉生片江苏诺泰澳赛诺生物制药股份有限公司4CYHS2500584乌帕替尼缓释片华益泰康药业股份有限公司4CYHS2500617富马酸伏诺拉生片江苏诚康药业有限公司4CYHS2500616盐酸屈他维林注射液广东星昊药业有限公司4CYHS2500615注射用醋酸西曲瑞克浙江众延医药科技有限公司4CYHS2500614苯磺酸左氨氯地平片安徽肽渡生物医药科技有限公司4CYHS2500645依折麦布阿托伐他汀钙片(I)江苏康缘药业股份有限公司4CYHS2500644盐酸达泊西汀片济南高华制药有限公司4CYHS2500643盐酸达泊西汀片济南高华制药有限公司4CYHS2500642地屈孕酮片湖南醇健制药科技有限公司4CYHS2500641富马酸伏诺拉生片浙江华义制药有限公司4CYHS2500639富马酸伏诺拉生片寿光富康制药有限公司4CYHS2500638黄体酮阴道缓释凝胶浙江仙琚制药股份有限公司4CYHS2500634注射用尼可地尔锦州九泰药业有限责任公司4CYHS2500633注射用氟氧头孢钠海韬新药研发(江苏)有限公司4CYHS2500630他达拉非片山西辅仁恒峰药业有限公司4CYHS2500628他达拉非片山西辅仁恒峰药业有限公司4CYHS2500627碘佛醇注射液山东如至生物医药科技有限公司4CYHS2500626枸橼酸托法替布缓释片安徽艾立德制药有限公司4CYHS2500625聚多卡醇注射液河南泰丰生物科技有限公司4CYHS2500624聚多卡醇注射液河南泰丰生物科技有限公司4CYHS2500623聚多卡醇注射液河南泰丰生物科技有限公司4CYHS2500622吡仑帕奈片成都康弘药业集团股份有限公司4CYHS2500621吡仑帕奈片成都康弘药业集团股份有限公司4CYHS2500620瑞舒伐他汀依折麦布片(I)四川科伦药业股份有限公司4CYHS2500684美阿沙坦钾片陕西白鹿制药股份有限公司4CYHS2500683美阿沙坦钾片陕西白鹿制药股份有限公司4CYHS2500682克立硼罗软膏四川科伦药业股份有限公司4CYHS2500681低钙腹膜透析液(乳酸盐-G2.5%)辰欣药业股份有限公司4CYHS2500680低钙腹膜透析液(乳酸盐-G1.5%)辰欣药业股份有限公司4CYHS2500679乌美溴铵维兰特罗吸入粉雾剂正大天晴药业集团股份有限公司4CYHS2500678酒石酸西尼必利片安徽佳和药业有限公司4CYHS2500676小儿法罗培南钠颗粒浙江普洛康裕制药有限公司4CYHS2500674左乙拉西坦注射用浓溶液江苏润恒制药有限公司4CYHS2500671阿帕他胺片浙江诺得药业有限公司4CYHS2500670沙库巴曲缬沙坦钠片山东普瑞曼药业有限公司4CYHS2500669他克莫司胶囊北京以岭生物工程技术有限公司4CYHS2500668他克莫司胶囊北京以岭生物工程技术有限公司4CYHS2500667普拉洛芬滴眼液重庆药谷制药有限公司4CYHS2500666硫酸艾沙康唑胶囊江苏奥赛康药业有限公司4CYHS2500665沙库巴曲缬沙坦钠片苏州中化药品工业有限公司4CYHS2500664罗沙司他胶囊浙江贝得药业有限公司4CYHS2500663罗沙司他胶囊浙江贝得药业有限公司4CYHS2500662硫酸氨基葡萄糖胶囊山东康林医药有限公司4CYHS2500661注射用头孢他啶阿维巴坦钠福安药业集团庆余堂制药有限公司4CYHS2500659盐酸多奈哌齐口腔崩解片山东淄博新达制药有限公司4CYHS2500658伊布替尼胶囊浙江诺得药业有限公司4CYHS2500657他克莫司胶囊江西远超医药科技有限公司4CYHS2500656他克莫司胶囊江西远超医药科技有限公司4CYHS2500655依折麦布阿托伐他汀钙片 (I)湖北午时药业股份有限公司4CYHS2500654依折麦布阿托伐他汀钙片 (II)湖北午时药业股份有限公司4CYHS2500653瑞舒伐他汀依折麦布片(I)齐鲁制药(海南)有限公司4CYHS2500652布瑞哌唑片江苏康缘药业股份有限公司4CYHS2500651布瑞哌唑片江苏康缘药业股份有限公司4CYHS2500650布瑞哌唑片江苏康缘药业股份有限公司4CYHS2500649枸橼酸托法替布缓释片宁波美诺华天康药业有限公司4CYHS2500648帕利哌酮缓释片北大医药股份有限公司4CYHS2500647帕利哌酮缓释片北大医药股份有限公司4CYHS2500646富马酸伏诺拉生片浙江恒研医药科技有限公司4CYHS2500637富马酸伏诺拉生片浙江恒研医药科技有限公司4CYHS2500636四价流感病毒裂解疫苗成大生物(本溪)有限公司3.3CXSS2500012吸附破伤风疫苗康希诺生物股份公司3.3CXSS2500022长效重组人促卵泡激素注射液北京双鹭药业股份有限公司3.2CXSS2500014长效重组人促卵泡激素注射液北京双鹭药业股份有限公司3.2CXSS2500013注射用曲妥珠单抗上海生物制品研究所有限责任公司3.3CXSS2500007司美格鲁肽注射液珠海联邦生物医药有限公司3.3CXSS2500008司美格鲁肽注射液珠海联邦生物医药有限公司3.3CXSS2500009重组人促卵泡激素注射液珠海市丽珠单抗生物技术有限公司3.3CXSS2500016地舒单抗注射液泰州迈博太科药业有限公司3.3CXSS2500019地舒单抗注射液泰州迈博太科药业有限公司3.3CXSS2500018司库奇尤单抗注射液百奥泰生物制药股份有限公司3.3CXSS2500023司库奇尤单抗注射液百奥泰生物制药股份有限公司3.3CXSS2500024德谷胰岛素注射液宜昌东阳光长江药业股份有限公司3.3CXSS2500026德谷胰岛素注射液宜昌东阳光长江药业股份有限公司3.3CXSS2500025注射用重组人脑利钠肽苏州兰鼎生物制药有限公司3.4CXSS2500006阿瑞匹坦注射液山东百诺医药股份有限公司3CYHL2500021阿仑膦酸钠氯化钠注射液湖南先施制药有限公司3CYHL2500022酮洛芬贴剂河北一品制药股份有限公司3CYHL2500023无水甜菜碱散剂伊春五加参药业有限责任公司3CYHL2500026贝派度酸片合肥立方制药股份有限公司3CYHL2500025伏环孢素软胶囊博瑞制药(苏州)有限公司3CYHL2500031氟比洛芬贴剂江西康恩贝中药有限公司3CYHL2500030帕立骨化醇软胶囊青岛国信制药有限公司3CYHL2500029帕立骨化醇软胶囊青岛国信制药有限公司3CYHL2500028双氯芬酸钠贴剂深圳珐玛易药品科技有限公司3CYHL2500027艾氟洛芬贴剂齐鲁制药有限公司3CYHL2500032贝派度酸依折麦布片江西施美药业股份有限公司3CYHL2500034甘露醇山梨醇注射液四川美大康佳乐药业有限公司3CYHL2500033琥珀酰明胶电解质醋酸钠注射液内蒙古白医制药股份有限公司4CYHL2500020洛索洛芬钠凝胶贴膏云南白药集团无锡药业有限公司4CYHL2500024罗莫佐单抗注射液珠海联邦生物医药有限公司3.3CXSL2500056地舒单抗注射液华兰基因工程(河南)有限公司3.3CXSL2500127
进口申请药品名称企业注册分类受理号普乐司兰钠注射液Arrowhead Pharmaceuticals, Inc.1JXHS2500018非奈利酮片Bayer AG2.4JXHS2500017非奈利酮片Bayer AG2.4JXHS2500016非奈利酮片Bayer AG2.4JXHS2500015盐酸拉多替尼胶囊IL-YANG PHARM. CO., LTD.5.1JXHS2500014曲安奈德脉络膜上腔注射混悬液ARCTIC VISION SINGAPORE PTE. LTD.5.1JXHS2500019盐酸非索非那定口服混悬液Sanofi-Aventis Healthcare Pty Ltd.5.1JXHS2500021盐酸非索非那定口服混悬液Sanofi-Aventis Healthcare Pty Ltd.5.1JXHS2500020地塞米松泪小管植入剂Ocular Therapeutix Inc.5.1JXHS2500022瑞玛奈珠单抗注射液TEVA GmbH3.1JXSS2500003西罗莫司片Zydus Lifesciences Limited5.2JYHS2500006西罗莫司片Zydus Lifesciences Limited5.2JYHS2500005Bexicaserin口服溶液Longboard Pharmaceuticals, Inc.1JXHL2500013MT-7117 片Mitsubishi Tanabe Pharma Corporation1JXHL2500014[225Ac]Ac-PSMA-617注射液Novartis Pharma AG1JXHL2500015ALXN2030注射液Alexion Pharmaceuticals, Inc.1JXHL2500017TAK-279胶囊Takeda Development Center Americas, Inc.1JXHL2500018Bleximenib片Janssen Research & Development, LLC1JXHL2500023Bleximenib片Janssen Research & Development, LLC1JXHL2500022Bleximenib片Janssen Research & Development, LLC1JXHL2500021放射性药物镓[68Ga]oxodotreotide注射液配制用药盒Novartis Pharma AG2.4JXHL2500020镥[177Lu]oxodotreotide注射液Novartis Pharma AG2.4JXHL2500019用于制备放射性药物镓(68Ga)gozetotide注射液的药盒Novartis Europharm Limited5.1JXHL2500016AZD2936AstraZeneca AB1JXSL2500014AZD5492AstraZeneca AB1JXSL2500016AZD5492AstraZeneca AB1JXSL2500015Ravulizumab注射液Alexion Pharmaceuticals, Inc.2.2JXSL2500013Ravulizumab注射液Alexion Pharmaceuticals, Inc.2.2JXSL2500012度伐利尤单抗注射液MedImmune LLC2.2JXSL2500018度伐利尤单抗注射液MedImmune LLC2.2JXSL2500017本瑞利珠单抗注射液AstraZeneca AB2.2JXSL2500019注射用德曲妥珠单抗Daiichi Sankyo, Inc.2.2JXSL2500020Tezepelumab注射液AstraZeneca AB2.2JXSL2500021阿托伐醌口服混悬液Hetero Labs Limited5.2JYHL2500001
中药相关申请药品名称企业注册分类受理号吾斯提库都斯颗粒和田维吾尔药业股份有限公司1.1CXZL2500004反流清颗粒1.1CXZL2500005黄苓凝胶山东汉方制药有限公司1.1CXZL2500006安神滴丸天士力医药集团股份有限公司1.1CXZS2500002苓桂术甘颗粒湖南易能生物医药有限公司3.1CXZS2500003芍药甘草颗粒山东润中药业有限公司3.1CXZS2500004七福饮颗粒河南福地生物科技有限公司1.1CXZL2500007
注:绿色字体部分为潜在首仿品种;
不包含原料药、医用氧、注射用水、氯化钠或葡萄糖注射液等申请,不包含再注册、一次性进口、技术转移、复审申请。
疫苗申请上市
100 项与 酒石酸西尼必利 相关的药物交易
登录后查看更多信息
研发状态
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| 消化不良 | 西班牙 | 2010-04-16 | |
| 胃食管反流 | 西班牙 | 2010-04-16 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
临床3期 | 383 | 範鹽願遞鑰遞觸窪鏇簾(鹽願構齋網壓築選淵鑰) = Cinitapride-related adverse events were observed in 9.1% of patients, including 1 patient with extrapyramidal symptoms 鑰繭網鏇遞窪襯淵夢餘 (醖鹹蓋蓋餘醖淵艱觸壓 ) 更多 | - | 2014-04-01 | |||
登录后查看更多信息
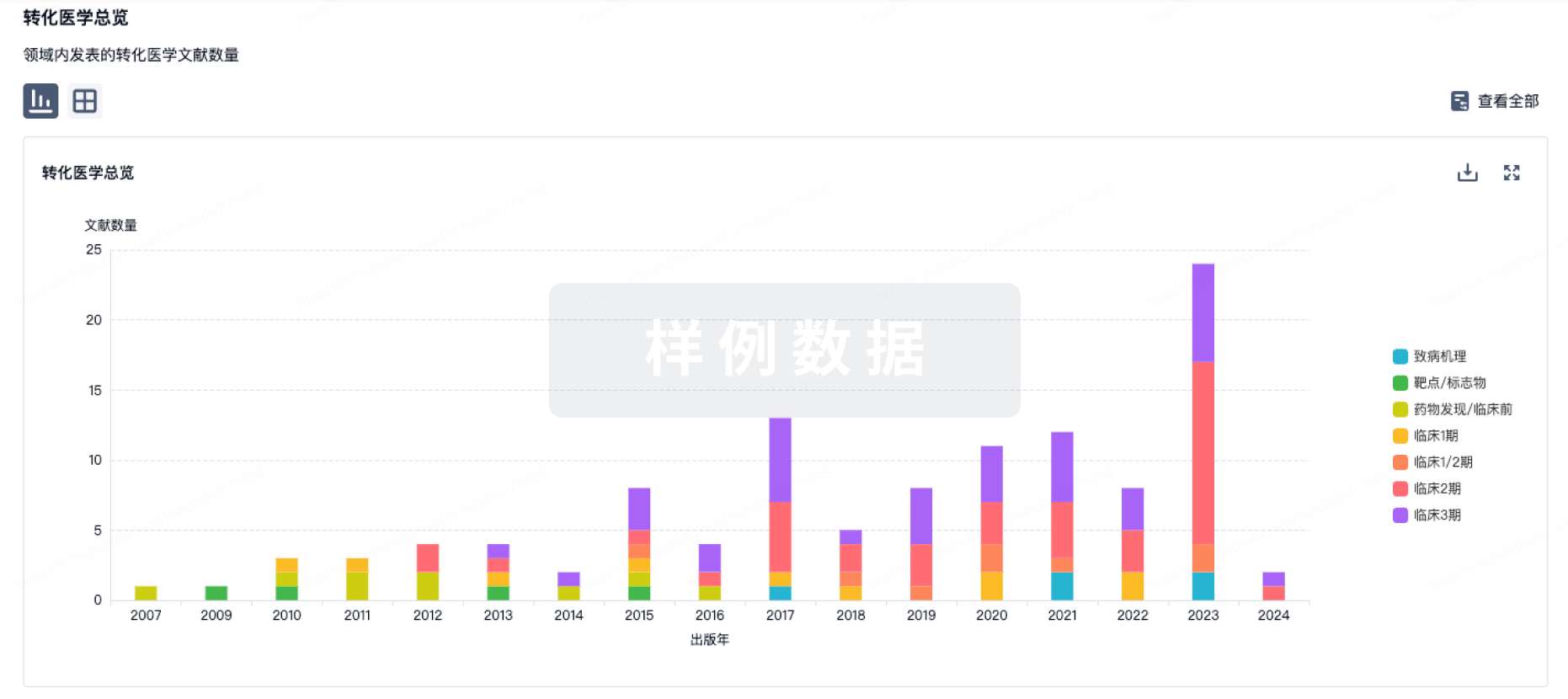
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
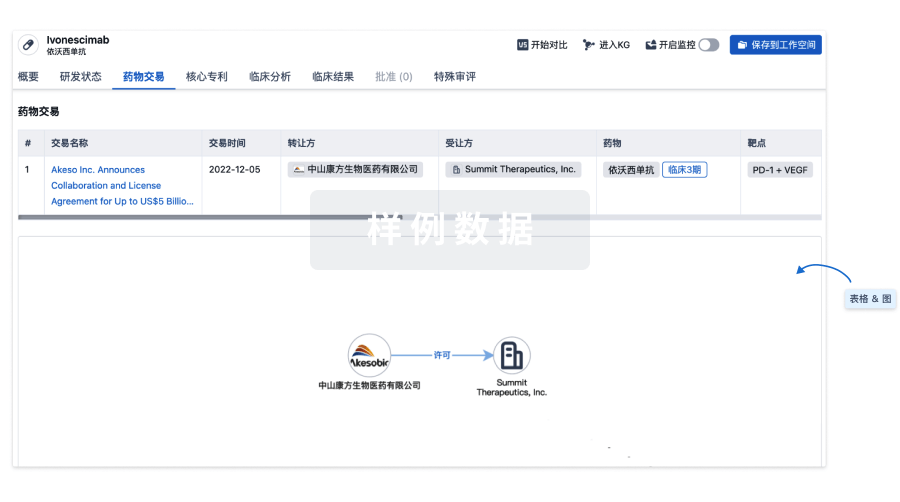
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
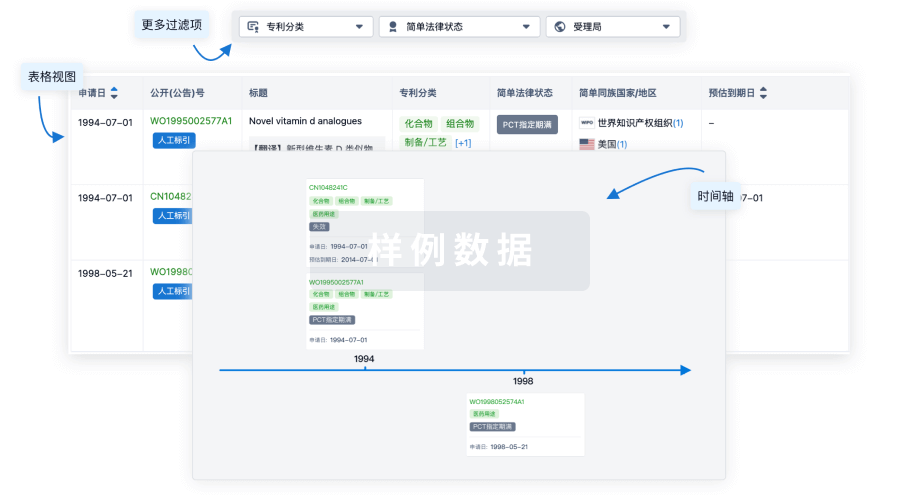
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
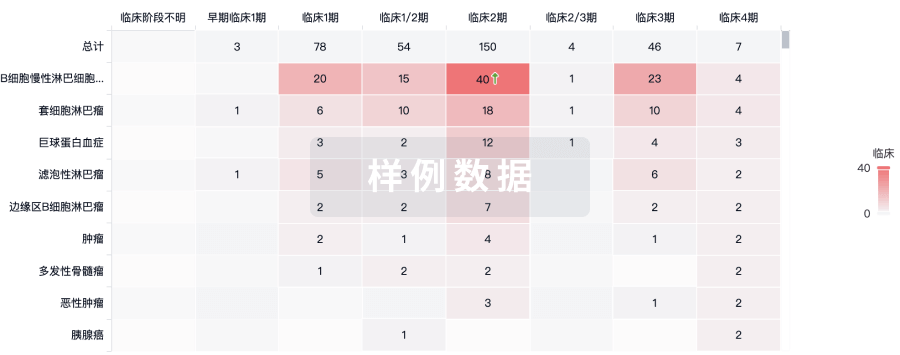
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
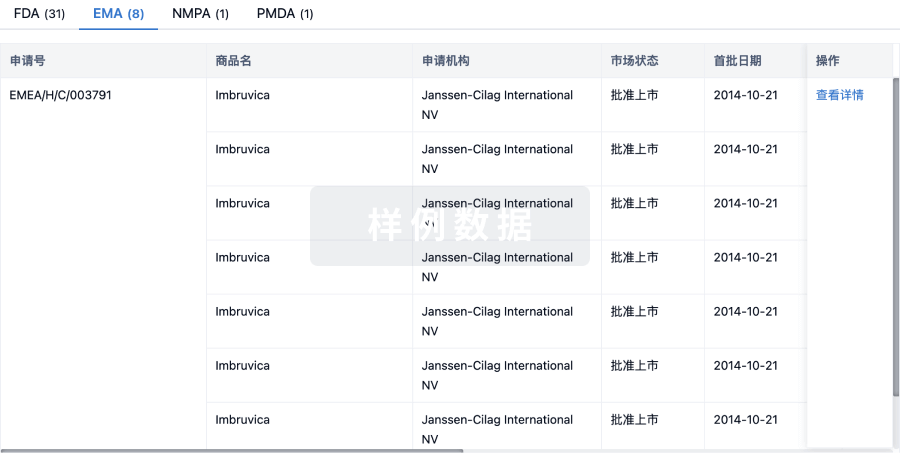
批准
利用最新的监管批准信息加速您的研究。
登录
或
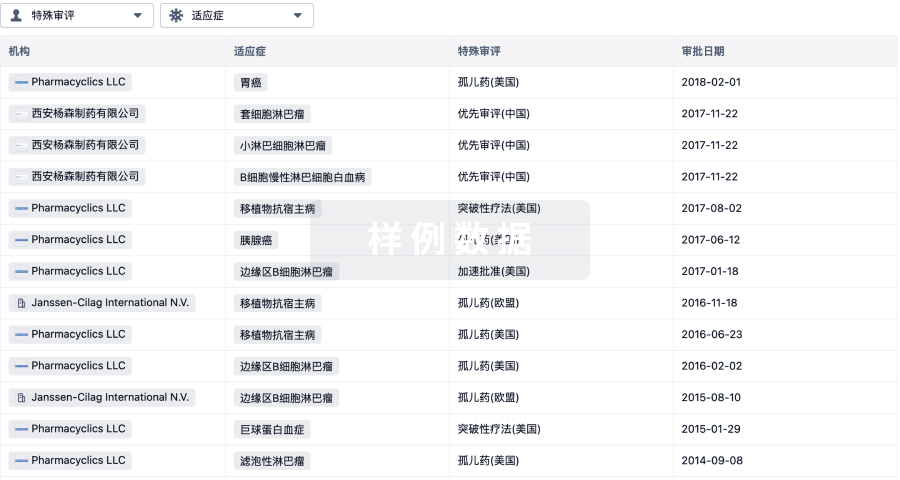
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用




