预约演示
更新于:2025-08-09

Daan Gene Co., Ltd.
更新于:2025-08-09
概览
标签
消化系统疾病
肿瘤
mRNA
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 疾病领域 | 数量 |
|---|---|
| 肿瘤 | 1 |
| 排名前五的药物类型 | 数量 |
|---|---|
| mRNA | 1 |
| 排名前五的靶点 | 数量 |
|---|---|
| LRP1(LDL receptor related protein 1) | 1 |
关联
1
项与 广州达安基因股份有限公司 相关的药物CN116004623
专利挖掘1
项与 广州达安基因股份有限公司 相关的临床试验DRKS00030899
Clinical Performance Evaluation of Detection Kit for Monkeypox Virus DNA (PCR-Fluorescence Probing) - REQA_22023_DaAn
开始日期2023-02-09 |
申办/合作机构 |
100 项与 广州达安基因股份有限公司 相关的临床结果
登录后查看更多信息
0 项与 广州达安基因股份有限公司 相关的专利(医药)
登录后查看更多信息
40
项与 广州达安基因股份有限公司 相关的文献(医药)2025-11-01·ANALYTICAL BIOCHEMISTRY
Oxford Nanopore third generation sequencing for analysis of FMR1 5′UTR CGG repeat expansions
Article
作者: Wang, Wenyu ; Huang, Jie ; Zhang, Li ; Ouyang, Guojun ; Zhang, Min ; Weng, Shi ; Chen, Juanjuan ; Yang, Xuexi ; Han, Bowei ; Du, Juan ; Han, Wanqing ; Yang, Xu ; Wu, Yingsong
OBJECTIVE:
Fragile X syndrome is mainly caused by the expansion of GC-rich cytosine-guanine-guanine (CGG) repeat in FMR1 5'UTR region, as well as rare gene point mutations or deletions in its open reading frame. Currently, third-generation long-read sequencing is a potential technology for simultaneously detecting CGG repeat expansions, point mutations, and deletions. However, a major challenge remains in obtaining the target long-fragment CGG repeat region with ultra-high GC content for sequencing.
METHODS:
We developed a novel approach combining long-fragment ultra-high GC polymerase chain reaction (PCR) amplification with Oxford Nanopore sequencing to detect the full spectrum of FMR1 5'UTR CGG repeat mutations. The method was validated using 10 standard cell line samples (males: nnormal = 1, nintermediate = 1, npre-mutation = 1, and nfull mutation = 2; females: nnormal = 1, nintermediate = 1, npre-mutation = 2, and nfull mutation = 1) and 53 retrospective clinical blood samples (males: nnormal = 7, npre-mutation = 3, nfull mutation = 15, and nmosaicmutaion = 2; females: nnormal = 9, npre-mutation = 13, and nfull mutation = 4).
RESULTS:
Our method demonstrated that the 100 % concordance with the triplet repeat-primed PCR and Southern blot analysis in genotyping 10 cell line samples and 53 clinical samples. Additionally, CGG repeat numbers showed strong correlation with reference mehods (male cell lines, n = 5, R2 = 0.9996; female cell lines, n = 5, R2 = 0.9972; clinical male samples, n = 26, R2 = 1.0000; clinical female samples, n = 25, R2 = 0.9854).
CONCLUSION:
This study presents a simple and cost-effective strategy for preparing FMR1 5'UTR CGG repeat regions with long-fragment ultra-high GC content for third-generation sequencing. The approach could serve as a model for detecting other challenging disorders caused by short tandem repeat expansions, such as myotonic dystrophy and Huntington' s disease.
2025-07-01·Microbiology Spectrum
Simultaneous detection and differentiation of
Mycobacterium tuberculosis
and nontuberculous mycobacteria in smear-negative sputum by a multiplex PCR assay: a clinical feasibility study
Article
作者: Chen, Li ; Yang, Yong ; Chen, Zhong ; Jiang, Shao-Long ; Xie, Long ; Zhang, Jia ; Jiang, Xi-Wen ; Tan, Jin-Lin ; Xu, Da-Yong ; Ruan, Ze-Fan ; Cao, Jing ; Huang, Ming-Xiang
ABSTRACT:
Accurate and rapid differentiation of
Mycobacterium tuberculosis
(
M. tuberculosis
) and nontuberculous mycobacteria (NTM) remains challenging, especially in smear-negative samples. Routine diagnostic methods often lack specificity and rapidity, highlighting the urgency of developing applicative molecular diagnostics to guide clinical management and improve patient outcomes. We developed and validated a multiplex PCR assay employing TaqMan probes targeting the
IS6110
and
rpoB
genes to simultaneously detect
M. tuberculosis
and 23 NTM species in a single reaction, while avoiding cross-reactivity. Analytical performance was assessed using reference strains, and clinical feasibility was evaluated in a prospective, single-center pilot study involving 96 smear-negative presumptive tuberculosis patients. The assay’s diagnostic performance was compared to mycobacterial culture, DNA sequencing, and the Xpert MTB/RIF assay. The multiplex PCR assay demonstrated a wide linear dynamic range (10
8
–10
3
copies/mL), low limit of detection (10
3
copies/mL), high precision (intra-assay CV 0.05–1.18%, inter-assay CV 0.08–2.57%), and complete specificity for both
M. tuberculosis
and NTM. In the clinical feasibility study, using a combination of clinical diagnosis and mycobacterial culture as reference standards, it achieved sensitivities of 73.33% (
M. tuberculosis
) and 100% (NTM), and specificities of 100% (
M. tuberculosis
) and 94.33% (NTM). Notably, it reliably identified
M. tuberculosis
/NTM co-infections. The Multiplex PCR assay fills a critical gap in the rapid, accurate diagnosis of
M. tuberculosis
and NTM in challenging clinical specimens. With a cost of $5 per test and a turnaround time of 3 h, our assay has the potential to improve patient care and optimize mycobacterial infection treatment strategies in resource-limited settings.
IMPORTANCE:
Rapid and accurate differentiation between
Mycobacterium tuberculosis
(
M. tuberculosis
) and nontuberculous mycobacteria (NTM) is essential for ensuring timely and appropriate treatment, especially in high tuberculosis (TB) burden regions. Conventional diagnostic methods, such as smear microscopy and culture, often lack the sensitivity or speed needed for reliable results in smear-negative cases, risking misdiagnosis and delayed care. In this study, we developed a novel multiplex real-time PCR assay capable of simultaneously detecting
M. tuberculosis
and up to 23 clinically relevant NTM species with high specificity and sensitivity. By targeting distinct genetic markers for
M. tuberculosis
and NTM, our assay provides a cost-effective, 3-h diagnostic solution that enhances diagnostic accuracy in challenging samples. This innovation addresses a critical gap in mycobacterial diagnostics, supporting improved patient outcomes and aligning with global health priorities for the control and elimination of TB.
2025-01-01·PLACENTA
Gene expression profiles based on maternal plasma cfDNA nucleosome footprints indicate fetal development and maternal immunity changes during pregnancy progress
Article
作者: Zhou, Huiling ; Yang, Xuexi ; Huang, Xiang ; Tan, Jiayu ; Guo, Zhiwei ; Ouyang, Guojun ; Wei, Xingyu ; Li, Fenxia ; Zhang, Min ; Hao, Wenbo ; Weng, Shi ; Li, Kun ; Liu, Yuming
BACKGROUND:
Pregnancy significantly alters the maternal immune system, affecting fetal development. The collection of tissues from the human placenta and fetus is not ethically or practically feasible at various gestational stages, thus limiting the study of gene expression in the fetus and placenta. Recent studies have shown that plasma cell-free DNA (cfDNA) nucleosome patterns can predict gene expression in the source tissue, offering insights into an individual's health status. This study aimed to identify pregnancy-related gene expression changes across gestational periods using cfDNA nucleosome distribution to understand fetal development and maternal immune changes.
METHODS:
Plasma samples were collected from 150 healthy pregnant women in different trimesters (early, mid, and late) and 32 healthy nonpregnant women. The correlation between gene expression and physiological changes during pregnancy was evaluated by inferring differential expression profiles around the transcription start site (TSS) using cfDNA nucleosome distribution patterns obtained through whole-genome sequencing. We utilized Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses to annotate differentially expressed genes with the mother and fetus.
RESULTS:
We identified gene expression changes that support the regulation of fetal development and immune system function during pregnancy. Differential coverage genes were mainly enriched in pathways related to transcription and translation, organic compound metabolism, and immune regulation. In addition, differentially expressed genes with significant temporal trends were identified. Among them, the upregulated differential genes were mainly related to development, whereas those with downregulated trends were mainly related to the immune system response. This indicates that differential changes of the placenta and maternal are significantly correlated with the pregnancy status.
DISCUSSION:
This study demonstrated the differential gene expression represented by the characteristic distribution of cfDNA nucleosome in maternal peripheral blood can effectively capture significant changes in maternal immunity and fetal development throughout pregnancy stages. It may help identify abnormal gene expression patterns associated with complications in pregnancy and childbirth, enhancing the quality of life and safety for both mother and fetus.
305
项与 广州达安基因股份有限公司 相关的新闻(医药)2025-08-07
来源:IVD资讯整理自广东财政厅
8月7日,广东省财政厅发布基孔肯雅病毒核酸检测试剂紧急采购项目结果公告。广州意达斯生物科技有限公司,以单价1.48元,总价29.6万元的报价中标。
该标是目前基孔肯雅核酸试剂盒第一大标,原预算280万,此次以29.6万元中标,单价打到1.48元,是目前该类试剂盒中标最低价。
达安上次3.9元中标,同行都说别活了(基孔肯雅热试剂,达安中标,3.9元!)今天这个标真的是低到尘埃了。这次关键性的采购,参与投标的IVD企业真是多,大厂报价已经很低了,但最终还是以最低价中标了。
这家中标的企业,行业内听过的人并不多,去年底才成立的,参保人数只有3人。
声明:本微信注明来源的稿件均为转载,仅用于分享,不代表平台立场,如涉及版权等问题,请尽快联系我们,我们第一时间更正。
诊断试剂带量采购
2025-08-07
·生物谷
葡萄糖代谢是肿瘤细胞维持快速增殖的重要代谢通路,因此抑制糖代谢(如低碳水化合物饮食)被视为一种潜在的抗肿瘤策略。但肿瘤的致死主要源自远处转移,而非原发瘤负荷。代谢干预是否可能在某些条件下反而促进肿瘤远端转移,一直缺乏系统研究。
7月15日,中山大学生命科学学院院邝栋明教授、魏瑗副教授团队在《Cell》期刊发表最新研究成果。研究结合多队列临床样本、多组学分析与小鼠模型,系统阐明了葡萄糖剥夺如何通过外泌体介导的免疫调控机制,塑造肺部促转移的免疫微环境,为肿瘤代谢与免疫互作研究提供了重要理论基础。
通过对15种肿瘤类型的大样本数据分析发现,肿瘤组织中葡萄糖代谢活性低的患者术后2年内复发风险明显升高。在多个小鼠肿瘤模型中,无论采用低碳水化合物饮食还是肿瘤糖酵解通路基因敲除,均观察到显著增强的肺部转移现象,且与原发瘤体积无关。
进一步研究发现,这一促转移效应并非源自代谢抑制增强了肿瘤细胞本身的转移能力,而是由葡萄糖代谢缺陷肿瘤细胞通过“旁观者效应”改变肺部免疫微环境,间接促进邻近代谢正常细胞转移。
机制研究显示,葡萄糖剥夺诱导肿瘤细胞发生内质网应激反应,激活泛素连接酶HRD1介导TRAIL的K63泛素化,并通过ESCRT复合体将其包装入外泌体中。释放至循环的外泌体-TRAIL在肺部促使PVR⁺巨噬细胞富集与极化,通过PVR–TIGIT信号轴耗竭NK细胞功能,削弱肺部先天免疫监视,从而建立有利于肿瘤定植的“转移前生态位”。
功能实验进一步证实,阻断TIGIT信号轴不仅可显著抑制葡萄糖剥夺诱导的肺转移,还能增强NK细胞的活性,并协同延缓原位肿瘤生长,提示该通路具有治疗干预潜力。
研究还发现,血浆中外泌体TRAIL水平可作为预测肝癌术后早期肺转移的生物标志物,其预测性能优于传统标志物AFP与肿瘤体积,有望用于高风险患者的早期筛查与干预评估。
研究从肿瘤代谢应激出发,揭示了代谢状态改变可通过外泌体介导的固有免疫细胞功能重塑,在转移前阶段驱动远端免疫生态位的建立,建立了代谢–免疫–转移之间的因果链条。团队负责人表示,相关发现不仅丰富了对肿瘤转移机制的理解,也为代谢-免疫联合调控策略提供了理论依据。
对免疫学领域而言,研究强调了巨噬细胞-NK细胞互作轴在肺转移前免疫塑形中的关键作用,也提示免疫检查点分子如TIGIT在代谢相关转移背景下具有重要干预价值。
值得一提的是,该研究是邝栋明团队在《Cancer Cell》(2024年12月)揭示抗原交叉呈递驱动免疫治疗超进展机制、以及在《Immunity》(2025年6月)报道肝癌中多胺代谢重编程驱动免疫抑制的新机制后的又一重要成果,系统阐明了肿瘤糖代谢干预与免疫微环境间的复杂互作机制。
原文链接:
https://www.sysu.edu.cn/info/2121/50602.htm
本文仅用于学术分享,转载请注明出处。若有侵权,请联系微信:bioonSir 删除或修改!
免疫疗法细胞疗法
2025-08-06
IVD资讯整理自采招网8月5日,广州市疾病预防控制中心(广州市卫生监督所)2025年基孔肯雅病毒核酸检测试剂紧急采购发布项目结果公告。达安基因中标,价格3.9元/人份。
报价汇总
感觉有了新冠核酸试剂的价格在前,基孔肯雅热核酸试剂价格也就这样了。广东疾控采购的20万人份还没开标,估计价格也在3块多吧。那些还在拼命研发基孔试剂的企业,散了吧,可能成本都收不回来。
声明:本微信注明来源的稿件均为转载,仅用于分享,不代表平台立场,如涉及版权等问题,请尽快联系我们,我们第一时间更正。
带量采购诊断试剂
100 项与 广州达安基因股份有限公司 相关的药物交易
登录后查看更多信息
100 项与 广州达安基因股份有限公司 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2025年08月12日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
1
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
CN116004623 ( LRP1 )专利挖掘 | 胃肠道肿瘤 更多 | 药物发现 |
登录后查看更多信息
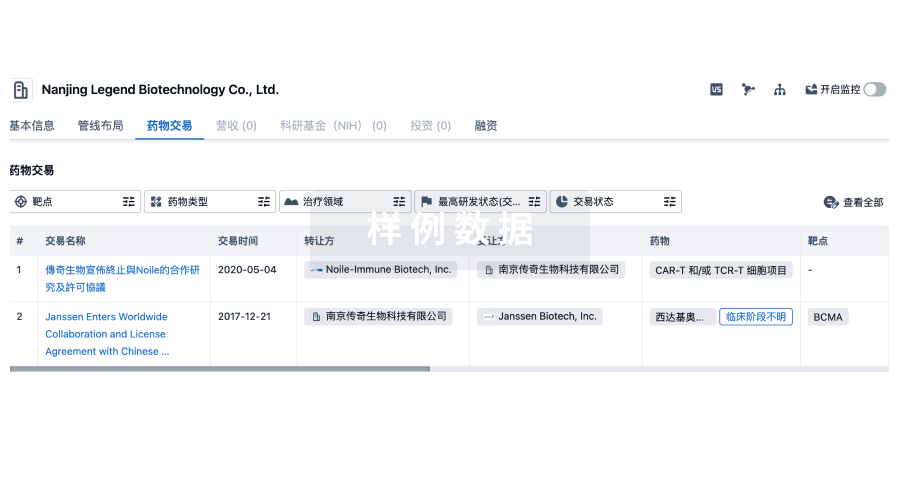
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
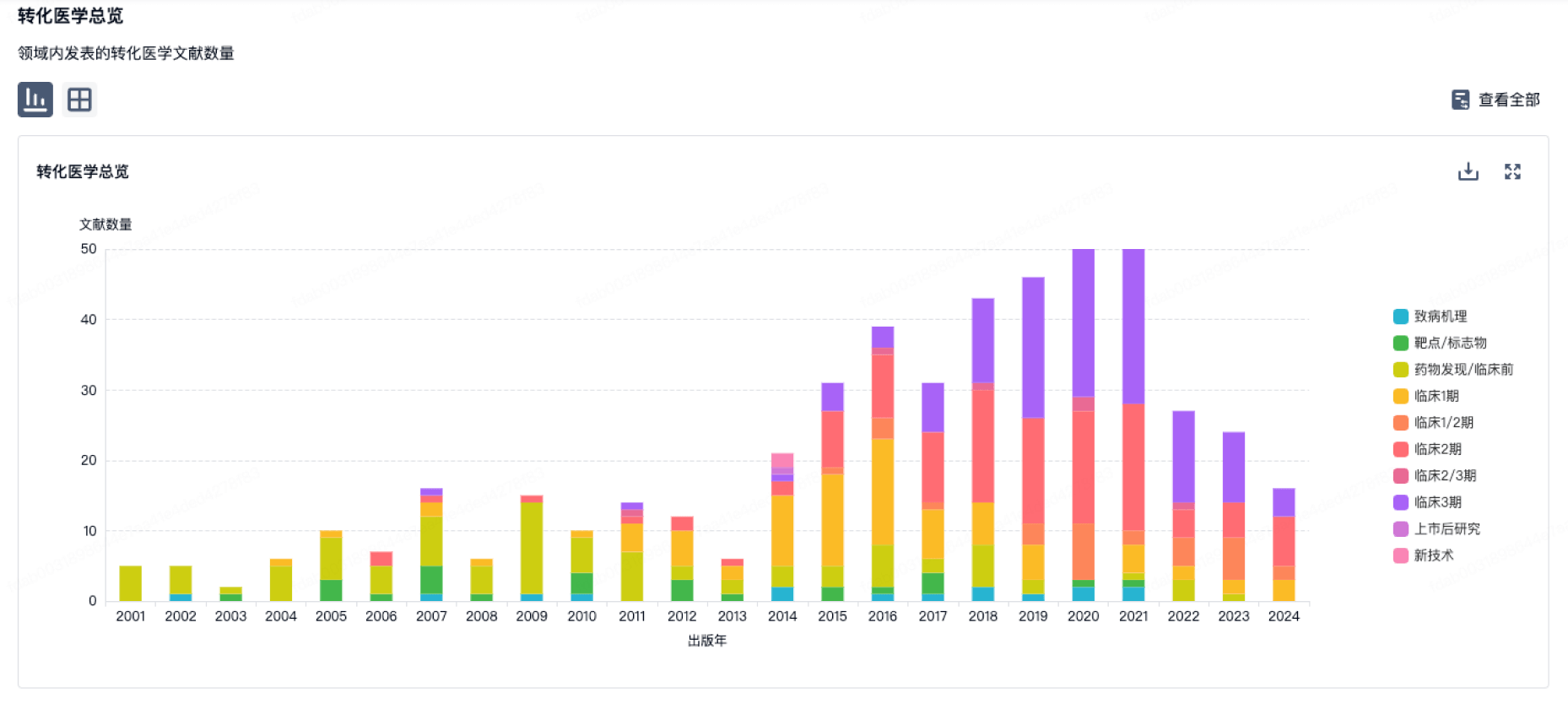
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
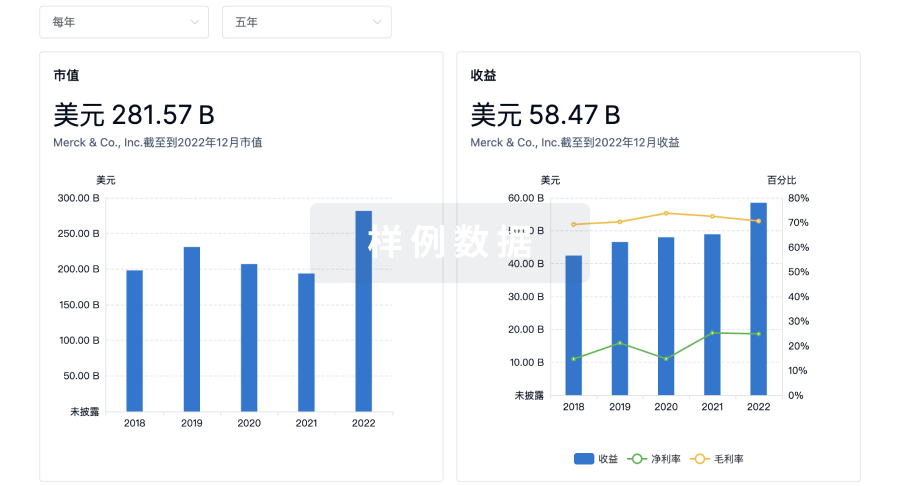
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
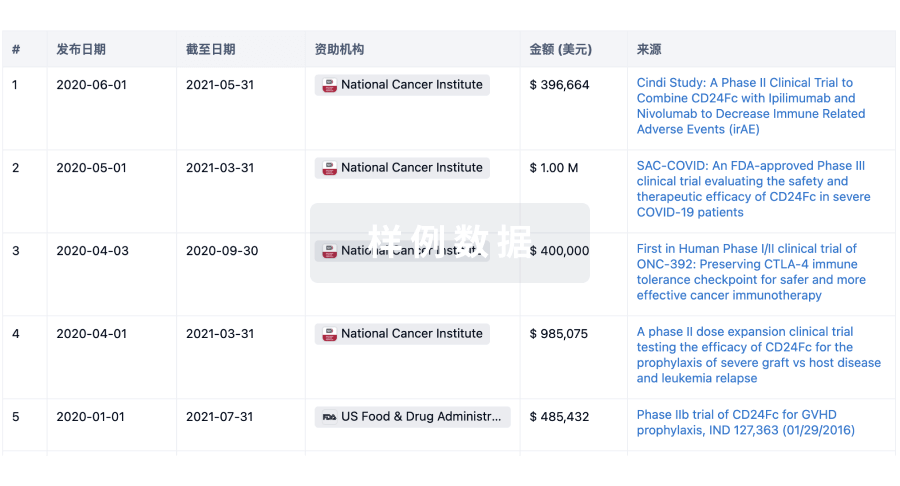
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
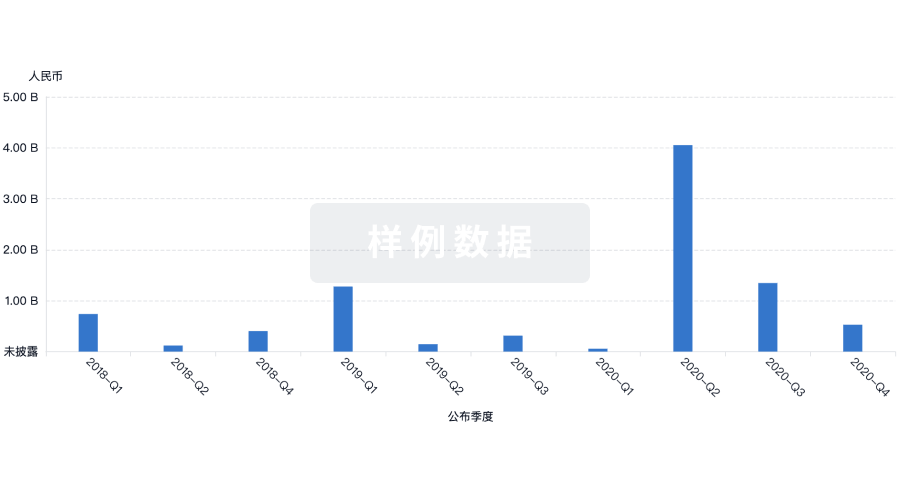
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
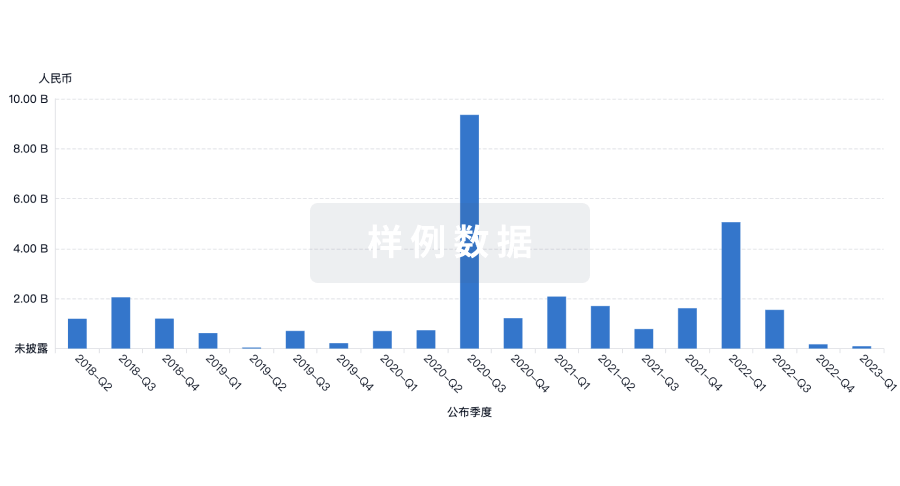
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用

