预约演示
更新于:2025-03-08

Razi Vaccine & Serum Research Institute
更新于:2025-03-08
概览
标签
呼吸系统疾病
感染
预防性疫苗
重组亚单位疫苗
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 疾病领域 | 数量 |
|---|---|
| 感染 | 1 |
| 排名前五的药物类型 | 数量 |
|---|---|
| 预防性疫苗 | 1 |
| 重组亚单位疫苗 | 1 |
| 排名前五的靶点 | 数量 |
|---|---|
| SARS-CoV-2 S protein(新冠病毒刺突蛋白) | 1 |
关联
1
项与 Razi Vaccine & Serum Research Institute 相关的药物作用机制 SARS-CoV-2 S protein调节剂 |
在研适应症 |
非在研适应症- |
最高研发阶段临床3期 |
首次获批国家/地区- |
首次获批日期- |
10
项与 Razi Vaccine & Serum Research Institute 相关的临床试验IRCT20201214049709N6
Immunogenicity and Safety of intranasal Razi Cov Pars as a booster dose in adult population; randomised, double blind, placebo controlled clinical trial
开始日期2022-12-03 |
IRCT20201214049709N5
Safety and Immunogenicity of Razi Cov-2 recombinant Spike protein vaccine (RAZI Cov Pars) in healthy children and adolescents aged 5-17 years; single group, open label study
开始日期2022-06-25 |
IRCT20201214049709N4
Comparison of immunogenicity and safety of Razi Cov Pars and Sinopharm booster doses in adults 18 years of age and older who have primarily vaccinated with Sinopharm: a ?parallel ??2?? arms, ?randomised, double blind clinical trial ?
开始日期2021-11-30 |
100 项与 Razi Vaccine & Serum Research Institute 相关的临床结果
登录后查看更多信息
0 项与 Razi Vaccine & Serum Research Institute 相关的专利(医药)
登录后查看更多信息
1,418
项与 Razi Vaccine & Serum Research Institute 相关的文献(医药)2025-03-03·Artificial cells, nanomedicine, and biotechnology2区 · 工程技术
Development and physicochemical, toxicity and immunogenicity assessments of recombinant hepatitis B surface antigen (rHBsAg) entrapped in chitosan and mannosylated chitosan nanoparticles: as a novel vaccine delivery system and adjuvant
2区 · 工程技术
ArticleOA
作者: Doroud, Delaram ; Mehrabi, Mohsen ; Pilehvar-Soltanahmadi, Younes ; Rezayat Sorkhabadi, Seyed Mahdi ; Ajdary, Soheila ; Dounighi, Naser Mohammadpour ; Amani, Amir ; Khoobi, Mehdi
In this study chitosan nanoparticles (CS NPs) and mannosylated chitosan nanoparticles (MCH NPs) loaded with recombinant hepatitis B surface antigen (rHBsAg) was synthesized as a vaccine delivery system and assessed toxically and immunologically. The physicochemical properties of the nanoparticles (NPs) were determined by methods including scanning electron microscope (SEM) and dynamic light scattering (DLS). The morphology of the NPs was semi spherical and the average diameter of the loaded CS and MCH NPs was found to be 189 and 239 nm, respectively. The release studies showed that after the initial burst, both of the loaded NPs provided a continuous and slow release of the antigens. 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay showed concentration and time dependent toxicity profile for both formulations, but rHBsAg loaded CS nanoparticle showed higher toxicity due to smaller particle size and larger zeta potential. Abnormal toxicity test (ATT) results showed no signs of toxicity in mice and guinea-pigs treated with loaded MCHNPs. Stability test for six months showed acceptable changes in size, surface charge, and antigenicity for loaded MCH nanoparticles. Finally, in vivo immunogenicity study revealed greater adjuvant capability of MCH nanoparticles than others formulations. Our results showed MCH NPs can be used as a controlled and targeted vaccine delivery system.
2025-03-01·PANCREAS
Current Understanding of Therapeutic and Diagnostic Applications of Exosomes in Pancreatic Cancer
Review
作者: Shakerian, Neda ; Tafazoli, Aida ; Khalili, Saeed ; Razavinia, Amir ; Sadrzadeh Aghajani, Zahra ; Khalesi, Bahman ; Hashemi, Zahra Sadat ; Bana, Nikoo ; Mard-Soltani, Maysam
ABSTRACT:
Unusual symptoms, rapid progression, lack of reliable early diagnostic biomarkers, and lack of efficient treatment choices are the ongoing challenges of pancreatic cancer. Numerous research studies have demonstrated the correlation between exosomes and various aspects of pancreatic cancer. In light of these facts, exosomes possess the potential to play functional roles in the treatment, prognosis, and diagnosis of the pancreatic cancer. In the present study, we reviewed the most recent developments in approaches for exosome separation, modification, monitoring, and communication. Moreover, we discussed the clinical uses of exosomes as less invasive liquid biopsies and drug carriers and their contribution to the control of angiogenic activity of pancreatic cancer. Better investigation of exosome biology would help to effectively engineer therapeutic exosomes with certain nucleic acids, proteins, and even exogenous drugs as their cargo. Circulating exosomes have shown promise as reliable candidates for pancreatic cancer early diagnosis and monitoring in high-risk people without clinical cancer manifestation. Although we have tried to reflect the status of exosome applications in the treatment and detection of pancreatic cancer, it is evident that further studies and clinical trials are required before exosomes may be employed as a routine therapeutic and diagnostic tools for pancreatic cancer.
2025-02-01·POULTRY SCIENCE
A new vaccination approach for Salmonellosis employing a multi-epitope vaccine based on live microbial cell factory from Lactococcus lactis
Article
作者: Jafari-Sohi, Mahnaz ; Moosavi-Kohnehsari, Reyhaneh Sadat ; Piri-Gharaghie, Tohid ; Aghassizadeh-Sherbaf, Mona ; Hosseinzadeh, Romina ; Tolou-Shikhzadeh-Yazdi, Shakiba
A major health and financial burden in the chicken sector is salmonella infection. It is difficult to create an oral vaccination that can provide strong intestinal mucosal immunity in birds, particularly cross-protection against several Salmonella serotypes. As a result, the poultry industry needs a powerful oral vaccination platform that uses live bacterial vectors to prevent various Salmonella serotypes. The genetically engineered L. lactis was given orally to birds as a vaccine after a multi-epitope vector was created using a reverse vaccinology technique. After the plasmid was digested, the target group produced a 72 kDa protein called multi-epitop. Birds that received the L. lactis/pNZ8121-Multi epitope vaccination showed increased levels of interferon (IFN-γ) and NFkB1α, increased transcription rates of cytokines, and a significant presence of IgY antibodies specific to the multi epitope gene in their serum. Salmonella infection is a severe health and economic burden in the poultry industry, according to spleen sections from the L. lactis/pNZ8121-Multi epitope. Developing an oral vaccine that can provide birds robust intestinal mucosal immunity-specifically, cross-protection against many Salmonella serotypes-is challenging. The results provide a fresh method for creating new immunological candidate multi-epitome genes by using the food-grade, non-pathogenic Lactococcus lactis as a protein cell factory. This method provides a unique technique to assess the long-term sustainability, cost, safety, and usefulness of experimental pharmaceutical products.
100 项与 Razi Vaccine & Serum Research Institute 相关的药物交易
登录后查看更多信息
100 项与 Razi Vaccine & Serum Research Institute 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2025年04月19日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
临床3期
1
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
Razi Cov Pars ( SARS-CoV-2 S protein ) | 新型冠状病毒感染 更多 | 临床3期 |
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
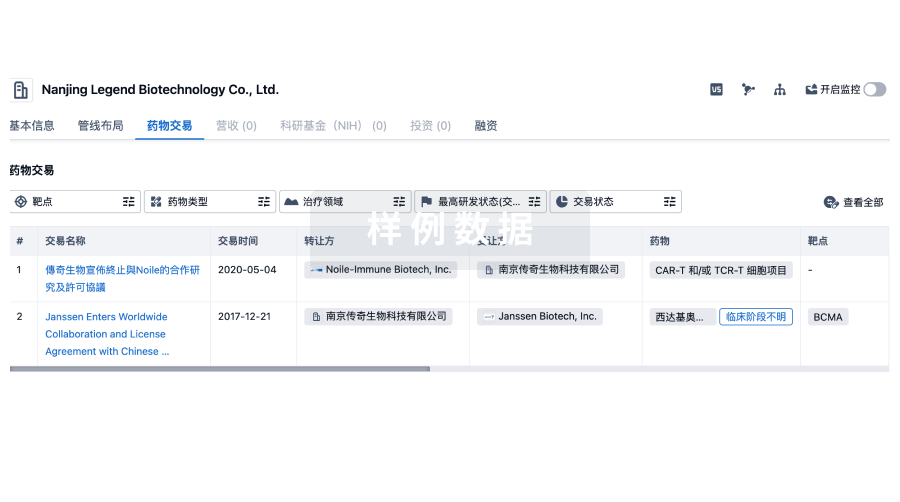
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
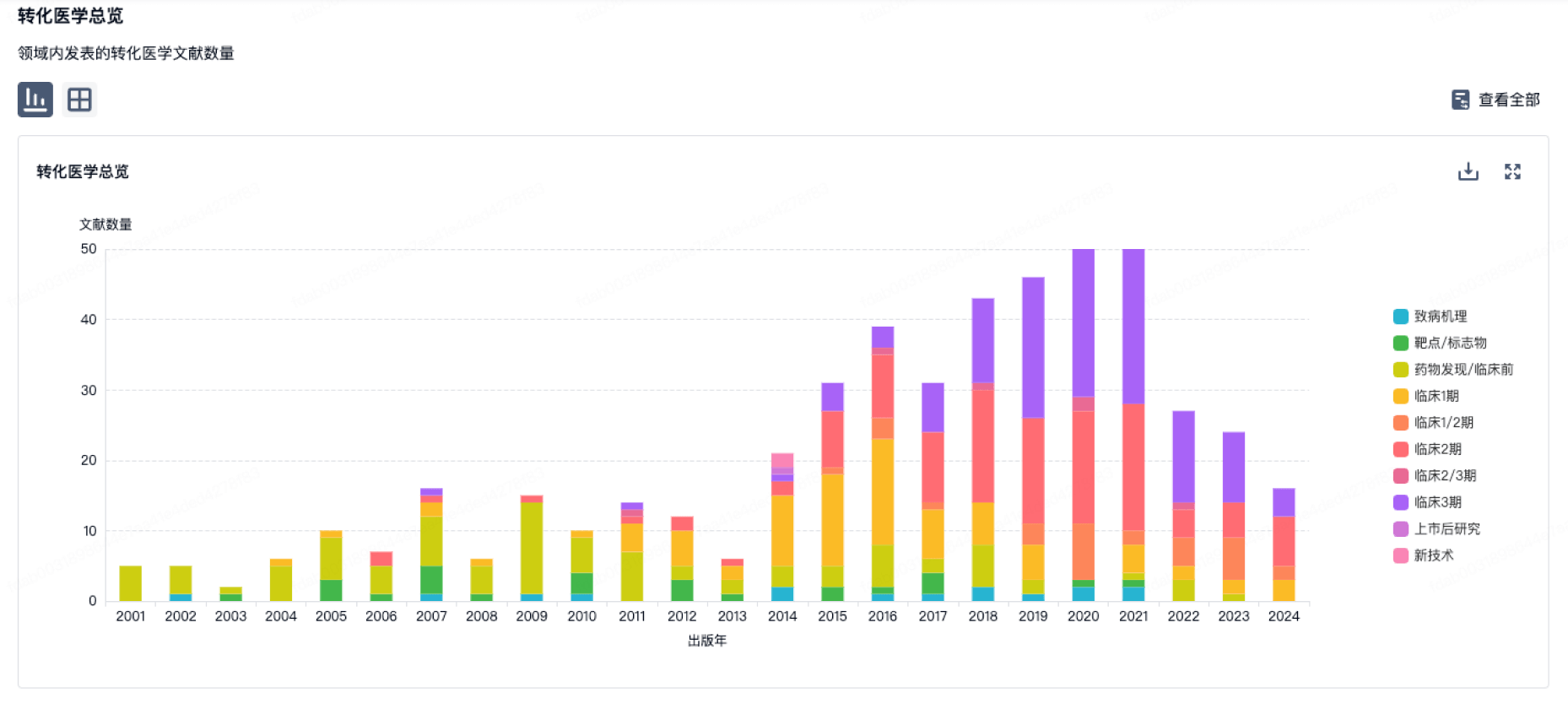
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
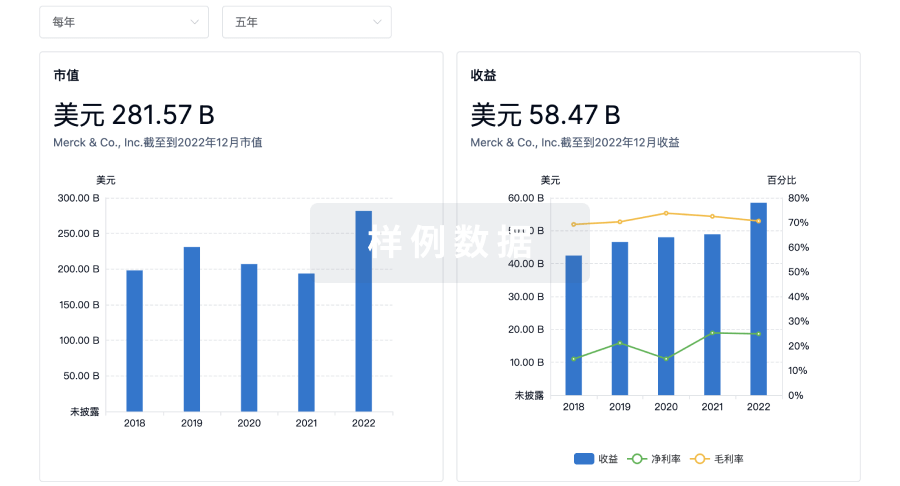
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
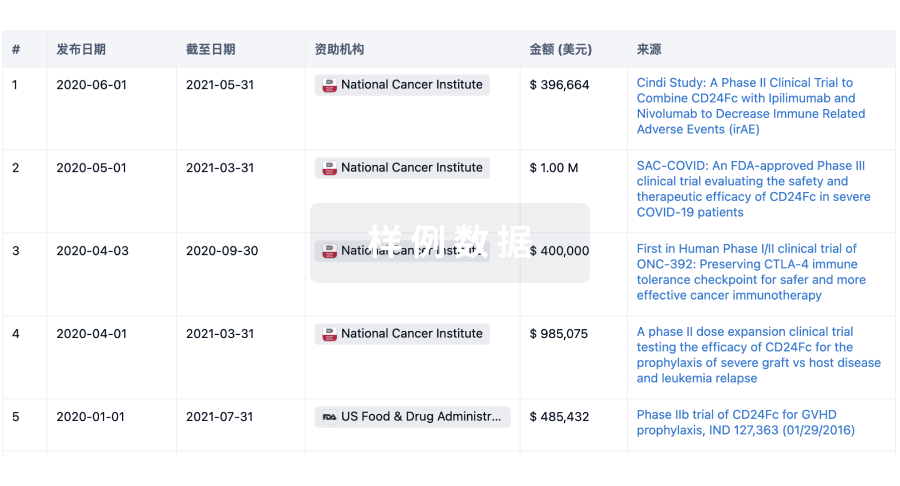
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
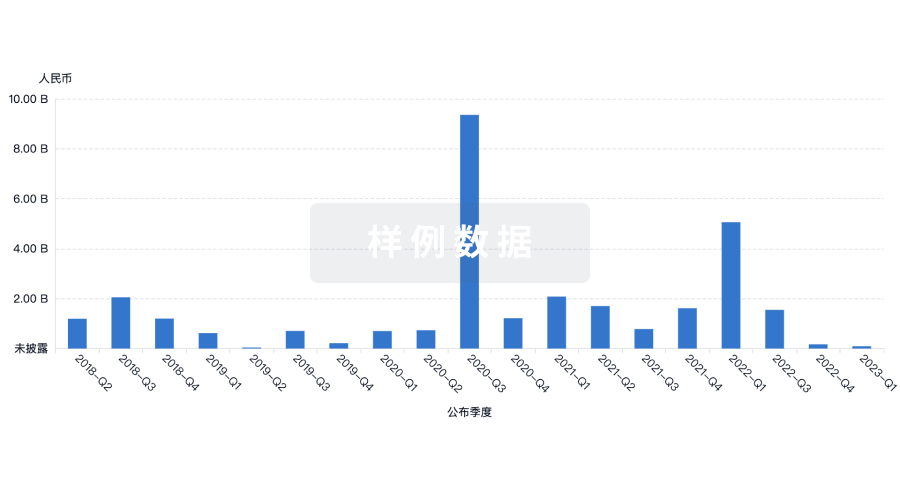
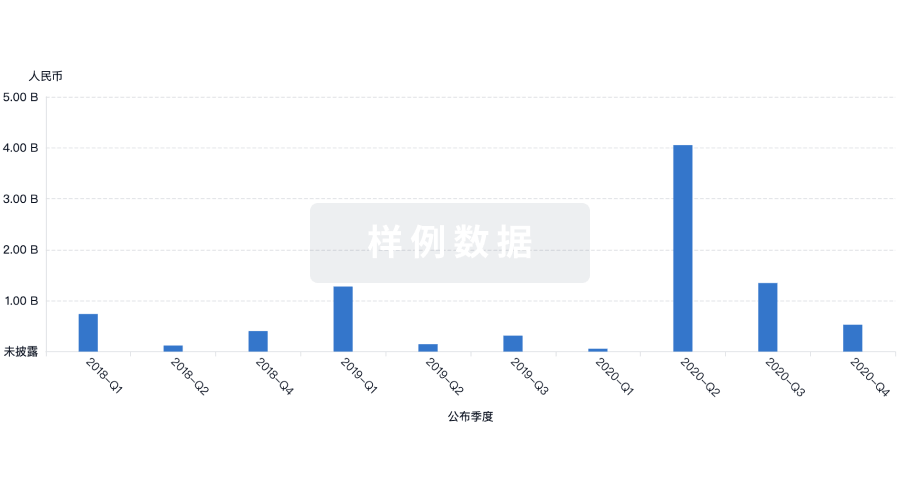
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
来和芽仔聊天吧
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用