预约演示
更新于:2025-08-29

Beijing Tiankeya Biotechnology Co., Ltd
更新于:2025-08-29
概览
标签
肿瘤
神经系统疾病
口颌疾病
TCR-T细胞疗法
自体CAR-T
合成多肽
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| TCR-T细胞疗法 | 3 |
| 自体CAR-T | 1 |
| 合成多肽 | 1 |
| 治疗性疫苗 | 1 |
关联
5
项与 北京天科雅生物科技有限公司 相关的药物靶点 |
作用机制 PD-1调节剂 |
在研机构 |
原研机构 |
在研适应症 |
非在研适应症- |
最高研发阶段临床1/2期 |
首次获批国家/地区- |
首次获批日期- |
靶点- |
作用机制- |
在研机构 |
原研机构 |
非在研适应症- |
最高研发阶段临床1/2期 |
首次获批国家/地区- |
首次获批日期- |
靶点- |
作用机制- |
在研机构 |
原研机构 |
在研适应症 |
非在研适应症- |
最高研发阶段临床1/2期 |
首次获批国家/地区- |
首次获批日期- |
7
项与 北京天科雅生物科技有限公司 相关的临床试验NCT06355908
Safety, Tolerability and Efficacy of Enhanced IL-13Rα2 Targeted Chimeric Antigen Receptor T Cell Immunotherapy Against Recurrent/Refractory Grade 4 Glioma
This is a dose exploration clinical trial to assess the safety and feasibility of the IL13Ra2-targeted CAR-T in glioma.
开始日期2024-03-21 |
申办/合作机构  首都医科大学附属北京天坛医院 首都医科大学附属北京天坛医院 [+1] |
NCT05881525
A Phase I/II Clinical Study of TC-N201 Injection for the Treatment of Advanced Solid Tumors With HLA-A2 Expression and Positive NY-ESO-1.
New York Esophageal Squamous Cell Carcinoma 1 (NY-ESO-1) is a cancer-testis antigen (CTA) which is expressed in various tumors. In TCR-T therapy, researchers take the blood of a certain patient, select T cells and insert genes into the cell that expressing a kind of protein that targeting NY-ESO-1. The genetically engineered cells are called NY-ESO-1 TCR-T cells. Then the engineered cells are re-infused to the cancer patients to cure the disease or prolong life.
开始日期2023-06-01 |
申办/合作机构  北京天科雅生物科技有限公司 北京天科雅生物科技有限公司 [+1] |
NCT04509726
Single-Arm Trial of EBV Specific Cytokine Secreting TCR-T Cells in the Treatment of EBV-Positive Head and Neck Carcinoma Metastatic/Refractory Nasopharyngeal Carcinoma
Epstein-Barr virus (EBV) infections is known to be a high-risk factor to induce cervical cancers. To date, EBV-related nasopharyngeal carcinoma (NPC) is still a major concern in east Asia, especially in China. Concurrent therapies for NPC have limited response rate and high chance of relapse. However, EBV-induced cancers provided an ideal target for T cell-based immunotherapy due to the non-self origins. Engineered T cells bearing a TCR (TCR-T) that can specifically recognize the presented EBV-epitope become a viable approach to treat this type of cancer. Though engineered T therapies have been well-recognized in hematological cancers, solid cancer treatment has been a major hurdle due to the immune-suppressive tumor microenvironment. Cytokine seemed to represent the ideal candidate for tumor immunotherapy, due to its ability to activate both innate (NK cells) and adaptive immunities. therefore, TCR-T cells armed with a cytokine -secretion element could further enhance the efficacy of TCR-T in solid cancers.
开始日期2023-03-01 |
申办/合作机构 新桥医院 [+1] |
100 项与 北京天科雅生物科技有限公司 相关的临床结果
登录后查看更多信息
0 项与 北京天科雅生物科技有限公司 相关的专利(医药)
登录后查看更多信息
4
项与 北京天科雅生物科技有限公司 相关的文献(医药)2024-02-01·MOLECULAR BIOTECHNOLOGY
Development of a Novel Pyroptosis-Associated lncRNA Biomarker Signature in Lung Adenocarcinoma
Article
作者: Peng, Ling ; Stebbing, Justin ; Zhu, Liping ; Castellano, Leandro ; Lin, Yanke ; Wang, Zhiqiang ; Wang, Peng
Pyroptosis is a novel type of cell death observed in various diseases. Our study aimed to investigate the relationship between pyroptosis-associated-long non-coding RNAs (lncRNAs), immune infiltration, and expression of immune checkpoints in the setting of lung adenocarcinoma and the prognostic value of pyroptosis-related lncRNAs. RNA-seq transcriptome data and clinical information from The Cancer Genome Atlas (TCGA) were downloaded, and consensus clustering analysis was used to separate the samples into two groups. Least absolute shrinkage and selection operator (LASSO) analyses were conducted to construct a risk signature. The association between pyroptosis-associated lncRNAs, immune infiltration, and expression of immune checkpoints were analysed. The cBioPortal tool was used to discover genomic alterations. Gene set enrichment analysis (GSEA) was utilized to investigate downstream pathways of the two clusters. Drug sensitivity was also examined. A total of 43 DEGs and 3643 differentially expressed lncRNAs were identified between 497 lung adenocarcinoma tissues and 54 normal samples. A signature consisting of 11 pyroptosis-related lncRNAs was established as prognostic for overall survival. Patients in the low-risk group have a significant overall survival advantage over those in the high-risk group in the training group. Immune checkpoints were expressed differently between the two risk groups. Risk scores were validated to develop an independent prognostic model based on multivariate Cox regression analysis. The area under time-dependent receiver operating characteristic curve (AUC of the ROC) at 1-, 3-, and 5-years measured0.778, 0.757, and 0.735, respectively. The high-risk group was more sensitive to chemotherapeutic drugs than the low-risk group. This study demonstrates the association between pyroptosis-associated lncRNAs and prognosis in lung adenocarcinoma and enables a robust predictive signature of 11 lncRNAs to inform overall survival.
2023-01-01·Journal of hepatocellular carcinoma
Identification of a Novel Prognostic Signature Based on N-Linked Glycosylation and Its Correlation with Immunotherapy Response in Hepatocellular Carcinoma.
Article
作者: Zhang, Honghua ; Lin, Shusheng ; Zhu, Xiangping ; Zhu, Ke ; Yang, Caini ; Zhang, Rui ; Cao, Yi
Background:
The complex tumor microenvironment of hepatocellular carcinoma (HCC) has led to a low response to immune checkpoints inhibitors (ICIs) and a poor prognosis. PD-L1, as one of the indications for ICIs, is rich in glycosylation modifications, which result in untimely ICIs. Our study constructed a prognostic model based on N-linked glycosylation related genes for predicting the prognosis and the response to ICIs.
Methods:
The list of N-linked glycosylation related genes is from the AmiGO2 database. The patients in The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO) cohorts were enrolled. The Cox regression was performed to develop a prognostic model and patients were divided into a low- and high-risk subgroups. The role of signature in HCC was well investigated by prognostic analysis, gene set enrichment analysis, and immune infiltration analysis. 21 recurrent HCC patients who received postoperative adjuvant ICIs were recruited to evaluate the relationship between immunotherapy response and the signature. In vitro studies were conducted to investigate the oncogenic effects of DDOST, STT3A and TMEM165 in HCC.
Results:
59 N-linked glycosylation related differentially expressed genes were screened from HCC and normal tissues in the TCGA cohort. The prognostic model was developed with DDOST, STT3A and TMEM165. The risk score could be an independent prognostic factor. Patients in the high-risk subgroup showed a worse prognosis than patients in the low-risk one. ssGSEA showed that patients in the low-risk subgroup tended to be in the immune-activated state, with higher levels of B cell and macrophage cell infiltrations and lower levels of regulatory T cell (Treg) infiltrations in both TCGC and GEO cohorts. Immunohistochemistry studies showed that DDOST, STT3A and TMEM165 are highly expressed in tumor tissues and patients with a high-risk score correlated with poor progression free survival and worse immunotherapeutic response. Furthermore, the proliferation of HCC cells was reduced after the knockdown of DDOST, as well as upon the knockdown of STT3A and TMEM165.
Conclusion:
In this study, we establish that the risk model based on N-linked glycosylation related genes could efficiently predict the prognosis and tumor microenvironment immune state of HCC patients, and the risk score could serve as a novel indicator of immunotherapy.
2020-01-01·Oncoimmunology2区 · 医学
Potential lung attack and lethality generated by EpCAM-specific CAR-T cells in immunocompetent mouse models
2区 · 医学
ArticleOA
作者: Chen, Yue ; Alexander, Peter B. ; Li, Dan ; Wang, Yongsheng ; Li, Xiaoyu ; Zhang, Benxia ; Qin, Diyuan ; Liao, Xuelian ; Li, Qi-Jing
The tumoricidal efficiency of human CAR-T cells is generally evaluated using immune-deficient mouse models; however, due to their immune-incompetency and the species-specific reactivity of a target antigen, these models are problematic to imitate CAR-T-induced adverse effects in the clinic. Epithelial cell adhesion molecule (EpCAM) is a tumor-associated antigen overtly presented on the cell surface of various carcinomas, making it an attractive target for CAR-T therapy. Here, we developed an anti-mouse EpCAM CAR to evaluate its safety and efficacy in immunocompetent mouse models. As previously reported for their human equivalents, murine EpCAM CAR-T cells exhibit promising anti-tumor efficacy in vitro and in vivo. However, after CAR-T infusion, various dose-depended toxicities including body weight loss, cytokine-release syndrome (CRS), and death were observed in both tumor-bearing and tumor-free mice. Pathological examination revealed unexpected and severe pulmonary immunopathology due to basal EpCAM expression in normal lung. While our study validates EpCAM CAR-T's potent anti-tumor efficacy, it also reveals that EpCAM CAR-T cells used for the treatment of solid tumors may cause lethal toxicity and should, therefore, be evaluated in patients with caution.
63
项与 北京天科雅生物科技有限公司 相关的新闻(医药)2025-06-09
·今日头条
近一半晚期肝癌患者肿瘤消退,23.5% 的患者甚至实现了 HBsAg (血清乙肝表面抗原)完全消失和持续清除!中国研发新型TCR-T疗法为肝癌患者开启治愈新可能!
肝癌,尤其是肝细胞癌(HCC),一直是全球癌症领域的难题。慢性乙型或丙型肝炎、大量饮酒、脂肪肝和其他黄曲霉素等毒性食物引起的肝损伤是导致肝癌的元凶,其中最为严峻的是HBV病毒性肝癌。据统计,全球每年有超过 42万人发生与 HBV 相关的 肝癌,至少占 50%全球病例,而亚洲最为严重,至少80%的患者是HBV 相关的肝癌!
尽管近年来针对晚期肝细胞癌的系统性治疗取得一定进展,靶向联合免疫治疗体系逐步建立,但 HBV 相关肝细胞癌的精准治疗仍存在巨大缺口。HBV 相关肝细胞癌患者体内的 HBV 特异性 T 细胞往往功能性耗竭,导致自身免疫系统无法有效清除感染肝细胞和肝癌细胞,治疗陷入困境。临床上迫切需要针对这一广大人群的新疗法!
近日,2025 年欧洲肝病研究协会大会上揭晓的一项 I 期临床试验(NCT06617000)数据显示,SCG101—— 一种自体乙肝病毒(HBV)特异性 T 细胞疗法,能够促使患者肿瘤缩小,并实现 HBV - DNA 和血清乙肝表面抗原(HBsAg)的持续清除的双重突破,引起巨大轰动!
双重突破!47%完全缓解+病毒持续清除!国研TCR-T开启肝癌治愈新可能
SCG101 是一种自体 T 细胞受体 (TCR) T 细胞疗法,是专门针对乙肝表面抗原 (HBsAg) 特定表位而设计的,这种全新的疗法可以特异性地靶向 HBV-HCC 感染细胞,能够触发溶细胞和非溶细胞两种机制,不仅能够高效清除 HBV 感染的肝细胞,还能对癌前病变细胞和 HBV - HCC 细胞进行精准打击,一箭双雕!
2023年国际细胞与基因治疗学会 (ISCT) 会议上,同类首创自体 HBsAg 特异性 T 细胞受体工程化 T 细胞 (TCR-T) 疗法 - SCG101 的初步数据公布,引起巨大轰动!
SCG101 显示出显著的抗肿瘤活性,一位患者接受单剂量 SCG101 单药治疗后肿瘤缩小 74.5%(mRECIST)。数据截止报告显示,肿瘤反应可持续 6.9 个月以上。
更值得一提的是,这位患者治疗后的活检显示肝脏中 HBsAg+ 肝细胞 100% 被根除!
2024年的欧洲肝病研究协会(EASL)大会上,SCG101公布了首次人体的临床试验结果,非常振奋人心!
在刚刚过去的2025年欧洲肝病研究协会(EASL)大会上g公布了最新的临床数据:
SCG101在晚期HBV 相关肝细胞癌 (HCC) 患者中展现出良好的抗病毒和抗肿瘤双重活性。在 6 例晚期HBV-HCC患者中,患者接受 5.0×10^7 ~ 1.0×10^8 TCR+ T细胞/kg 的 SCG101 治疗,客观缓解率 (ORR) 为
33%
。所有获得部分缓解( PR) 的患者均维持缓解超过 6 个月,其中 1名幸运的患者靶病灶完全缓解并维持无进展生存期
27 个月
。
这项试验纳入的 17 名符合条件的乙肝相关肝细胞癌患者,这些患者都是既往做过各类治疗失败的晚期难治患者。令人惊喜的是,在接受单次 SCG101 输注后,所有患者的血清 HBsAg 水平均呈现迅猛下降趋势。尤为瞩目的是,
94%
的患者在短短 28 天内,HBsAg 水平从 1.0 log₁₀急剧降至 4.6 log₁₀,并且在长达一年的时间内,始终维持在 100 IU/mL 以下的低水平。此外,
23.5%
的患者甚至实现了 HBsAg 完全消失,这一成果在 HBV 相关疾病治疗中具有重大意义。
在抗肿瘤活性方面,SCG101 同样表现出色。高达
47%
的患者在接受输注后,出现了可测量的肿瘤消退迹象。截至数据统计截止日期,患者的中位总生存期(OS)尚未达到,意味着该疗法有望为患者带来更长久的生存可能,极大地改善了患者的预后情况。
北京协和医院肝外科主任杜顺达医学博士在相关新闻稿中指出:“
SCG101 展现出的抗病毒和抗肿瘤双重功效,极具前景,特别是对于那些经历过大量前期治疗的患者而言。持续的 HBsAg 清除以及肿瘤缓解现象表明,SCG101 很可能成为 HBV 相关肝细胞癌患者全新的免疫治疗选择,有望填补该领域长期以来尚未满足的重大临床需求。
”
SCG 细胞治疗公司首席执行官 Christy Ma 在新闻稿中补充道:“这些积极的数据,标志着 SCG101 研发进程中迈出了关键一步,也充分验证了我们利用精准 T 细胞疗法治疗慢性乙型肝炎(HBV)感染以及 HBV 相关肝癌的策略的正确性。SCG101 作为首款在 HBV 相关肝细胞癌(HCC)患者中,同时证实能够实现病毒学清除和肿瘤消退的 T 细胞受体(TCR)T 细胞疗法,这些数据让我们备受鼓舞。我们满怀期待,将通过进一步的临床开发,推动 SCG101 尽快应用于临床,为有需求的患者带来这种潜在的治愈性疗法。”
宣战肝癌的终极武器!全新TCR-T免疫疗法震撼登场
人体的免疫防御机制主要依靠体内的白细胞军团,作为白细胞的一种,T细胞有着不可替代的作用,它们是人体里面的特种兵,一旦发现癌细胞,T细胞首先主动出击,杀灭敌人,因此,它也被称为“杀手T细胞”。
大部分肿瘤患者体内没有足够能识别和杀伤肿瘤细胞的T细胞,对于这些患者,医生可以采用一种称为工程T细胞受体(TCR)治疗的方法。
TCR-T技术,即T细胞受体(TCR)嵌合型T细胞技术,也叫亲和力增强的TCR”技术,需要从患者身上获取T细胞,为T细胞配备新的T细胞受体,使其能够靶向特定的癌症抗原的疗法,这种新型的疗法可以允许医生为每个患者的肿瘤和不同类型的T细胞选择最合适的靶点进行工程改造,使治疗个体化,并为患者提供更大的缓解希望。
这款疗法之所以备受期待,上市后引起全球轰动,开启实体瘤治疗新纪元,主要是因为以下三点:
1.扩增数量高达数十至上百亿的“活的”细胞疗法。
与CAR-T疗法不同,TCR-T是通过收集患者自己的 T 细胞,并对其进行改造使其具有改良的 TCR 而制成的。这些 TCR 具有增强的识别肿瘤特异性抗原蛋白的能力,这种蛋白质在肝癌,滑膜肉瘤等一些癌细胞表面高表达。一旦经过改造并扩增到数十至上百亿级别,具有增强 TCR 的 T 细胞就会通过一次性治疗放回您的体内,精准识别并疯狂攻击表达特定的癌细胞。
2.肿瘤缩小甚至消失,有望治疗多种实体瘤
除了滑膜肉瘤,这款疗法在包括恶性程度极高的
滑膜肉瘤,卵巢癌,头颈癌,胃癌,粘液样/圆形细胞脂肪肉瘤,非小细胞肺癌,尿路上皮癌,食道癌和黑色素瘤等各类
临床上可以说是没有任何标准治疗方案的极晚期患者中显示出了卓越的潜力,众多接受了新型的TCR-T疗法的患者,产生了强烈的响应。
3.“一针”即可清除肿瘤。
与 CAR-T 和TIL一样,TCR-T 疗法是一种一次性治疗。
步骤1 :血液和肿瘤组织检测,确认HLA类型和MAGE-A4阳性
步骤2 :T细胞采集,这些细胞将用于制造 TECELRA。
步骤3 :治疗前化疗(4 天),您将接受淋巴细胞清除术。
步骤4 :TECELRA 给药(1 个输液袋最长 1 小时)
步骤5 :输注后监测(最长 4 周)
CAR T细胞疗法
TCR-T细胞疗法
结构体
– 由细胞内和细胞外结构域组成的工程受体
– 天然或最低限度工程化的 TCR
目标
– 仅针对细胞表面抗原
– 针对 MHC 类分子呈现的细胞表面和细胞内抗原
优点
– 细胞靶标不受 MHC 复合体限制。
– 血液癌症的良好治疗效果
– 实现更多样化的靶标,因为它可以靶向细胞内抗原
– 允许与 MHC 进行高度敏感的结合相互作用实体瘤治疗中的令人鼓舞的结果
缺点
– 对实体瘤影响不大
– 副作用如细胞因子释放综合征和神经毒性
– 受 MHC 相容性限制
– 细胞因子释放综合征和神经毒性等副作用
FDA 状态
– 批准治疗多种癌症
– 预计上市时间(2024年8月)
CAR-T和TCR-T疗法的区别
国内患者如何接受TCR-T疗法?
除了SCG101,目前国内有包括香雪生物,百吉生物、来恩生物、华夏英泰、可瑞生物、天科雅、普瑞金、深圳宾德、泛恩生物、立凌生物等17家生物公司布局TCR-T赛道,其中50多个TCR-T在研项目已进入或者获批临床。其中,香雪生命科学的TAEST16001注射液是国内进度最快的产品之一,已进入II期临床阶段。此外,进度较快的还有华夏英泰的YT-E001,正在开展治疗Epstein-Barr病毒(EBV)相关鼻咽癌的 II期临床。
好消息是,目前多款针对肝癌,宫颈癌,胰腺癌,结直肠癌等各类实体瘤的临床试验正在进行中,很多患者已通过全球肿瘤医生网成功入组。
部分入组条件
18-75岁,男女不限,无严重基础疾病;
仅患一种恶性实体肿瘤,肝癌、头颈部鳞癌、宫颈癌、胰腺癌等实体瘤;
经标准治疗失败或缺乏有效治疗方法;
各项检查符合要求且至少1年的预期寿命。
想寻求TCR-T疗法及其他国内外新抗癌治疗帮助,且经济条件允许的情况下,可以先将病历提交至全球肿瘤医生网医学部进行初步评估,一旦审核通过,有机会获得”天价“疗法免费治疗的机会。
随着全球首款TCR-T细胞疗法的成功上市和国内企业的不断努力,全球范围内TCR-T细胞疗法领域有望迎来爆发式增长。预计在未来几年内,将有更多TCR-T细胞疗法产品获批上市,为患者提供更多样化、更精准的治疗选择。
越来越多的医生将有能力让合适的患者在正确的时间获得个体化”量体裁衣“式的治疗方案!相信这些效果好的抗癌疗法都会加快在我国上市和纳入医保的步伐,让更多的百姓获益,我们共同期待!随着更多治疗方案的发展,晚期肿瘤患者将迎来更多治疗选择和希望,我们期待人类攻克癌症的那一天早日到来。
细胞疗法免疫疗法
2025-05-21
·今日头条
2025年5月,国家药品监督管理局签发《临床试验批准通知书》(编号:2025LP01317),经审查,
TAEST1901注射液符合《中华人民共和国药品管理法》及相关规定的药品注册要求,同意开展治疗基因型为HLA-A*02:01且肿瘤抗原AFP表达阳性的晚期胃癌临床试验。
作为香雪生命科学研发管线的第二个产品,TAEST1901注射液通过慢病毒转导自体T细胞,使其表达AFP抗原特异性的TCR(T细胞受体),靶向HLA-A*02:01与AFP抗原肽组成的复合物。
值得关注的是,该产品此前(2022年4月)已获得国家药监局新药临床注册批准,适用于HLA-A*02:01基因型、AFP抗原阳性的晚期肝癌或其他晚期肿瘤的治疗,目前针对首个适应症晚期肝癌的1期临床试验正在推进中。期待这款靶向实体瘤的TCR-T疗法在临床研究中取得积极进展,为肿瘤患者提供更多治疗选择!
▲截图源自“NMPA”
国产TCR-T疗法全面开花!8大创新药获临床许可,横扫肝癌、宫颈癌、胰腺癌、结直肠癌等
除上述香雪制药近期获批临床的TAEST1901注射液外,国内还有多款TCR-T疗法已获得中国国家药品监督管理局新药IND临床试验默示许可,包括但不限于:
1、TAEST16001注射液
由香雪生命科学技术(广东)研发,2024年6月其IND申请获批准,用于
HLA-A*02:01阳性并且表达NY-ESO-1抗原的晚期软组织肉瘤
的治疗。同年7月30日被纳入突破性治疗品种名单,是
中国首个获得IND批件并开展临床研究的TCR-T细胞产品
。
2、SCG101注射液
SCG101注射液是星汉德生物研发的新一代人乳头瘤病毒(HPV)特异性T细胞受体工程化T细胞治疗产品。2022-2023年先后获得中国国家药监局、美国食品药品监督管理局(FDA)、新加坡药监局的临床试验批准,最初用于治疗
乙型肝炎病毒(HBV)相关的肝细胞癌
;2023年11月,其适应症进一步拓展至
HBV相关的肝内胆管癌
。
在国际细胞与基因治疗大会(ISCT)上,曾公布过“SCG101TCR-T治疗HBV相关肝细胞癌”的突破性案例。一位HBV相关肝细胞癌患者在接受单剂SCG101治疗后,不仅达到
部分缓解(PR),肿瘤体积更缩小74.5%
,且
体内乙肝感染被100%清除
。治疗第28天,
肿瘤靶病灶较基线缩小66%
;治疗第4个月,
肿瘤进一步缩小至74.5%
,
另有一处病灶更是完全消失
(详见下图)。截至数据统计时,患者无进展生存期已超6.9个月,且持续维持缓解状态。
▲图源“SCG”,版权归原作者所有,如无意中侵犯了知识产权,请联系我们删除
3、SCG1422
星汉德生物研发的新一代人乳头瘤病毒(HPV)特异性T细胞受体工程化T细胞治疗产品,2024年6月30日获美国FDA批准新药临床试验(IND)申请,用于HPV相关实体瘤治疗。2025年1月2日,中国NMPA也批准该产品的新药临床试验申请,适应症为
HPV感染相关恶性肿瘤
,包括
宫颈癌、口咽癌、头颈癌、阴道癌、外阴癌和阴茎癌
等。
目前SCG142TCR-T疗法已启动临床招募,现面向HPV相关恶性肿瘤患者提供免费治疗机会。阅读原文了解详情:【TCR-T细胞疗法免费招募】SCG142TCR-T疗法终于启动临床啦!现正招募HPV相关恶性肿瘤患者!
4、DCTY1102注射液
由北京鼎成肽源生物技术有限公司开发。2024年8月8日获得中国NMPA临床试验默示许可,是
国际上首款靶向HLA-A*11:01基因型、KRASG12D突变的TCR-T细胞治疗产品
,用于
晚期胰腺癌、结直肠癌
等恶性实体肿瘤患者的治疗。
5、GZL-016注射液
来恩生物研发的一款mRNATCR-T细胞疗法。2024年11月12日获中国NMPA的临床默示许可,用于
乙型肝炎病毒相关肝细胞癌
的治疗。
6、CRTE7A2-01 TCR-T细胞注射液
由北京可瑞生物研发的一款TCR-T疗法。2023年11月21日获国家药监局药品评审中心的新药临床试验(IND)默示许可,用于治疗
HPV16阳性HLA-A*02:01阳性晚期实体肿瘤(头颈部肿瘤、宫颈癌、肛门癌等)
。
7、IX001 TCR-T注射液
由镔铁生物研发的一款TCR-T疗法。2024年10月9日获得中国NMPA的临床试验默示许可,用于基因型为HLA-A*11:01、肿瘤抗原
KRASG12V表达为阳性的晚期胰腺癌
的治疗。
8、TC-N201注射液
由天科雅生物研发的一款TCR-T疗法,其新药IND已获得中国NMPA批准,是
国内首个治疗宫颈癌的TCR-T临床试验
,同时也是
全球首个批准的加载抗PD-1抗体的HPVTCR-T临床试验
,目前正在开展治疗实体瘤的1期临床研究。阅读原文了解详情:【TCR-T疗法免费招募】TC-N201注射液终于启动临床啦!现正急招滑膜肉瘤等NY-ESO-1阳性实体瘤患者!
TCR-T联合DC疫苗疗效惊艳,转移性黑色素瘤患者肿瘤消退率近70%!
TCR-T细胞疗法除单药治疗外,还可与其他疗法(如DC疫苗)联用。《临床癌症研究》一项临床试验(NCT00910650)探讨了MART-1 TCR转基因淋巴细胞过继转移联合MART-1肽脉冲树突状细胞(DC)疫苗,在HLA-A2.1阳性转移性黑色素瘤患者中的疗效。
研究共纳入14例HLA-A*0201阳性且MART-1阳性的转移性黑色素瘤患者,平均年龄为50岁,其中9例为伴有内脏和/或骨转移的M1c期,4例为肺转移的M1b期,1例为仅皮肤、淋巴结和皮下转移的M1a期。
结果显示:
69%(9例)的患者出现肿瘤消退迹象
。在10例可评估疗效的患者中,8例在治疗第30天的PET扫描(详见下图)或体检中显示初始抗肿瘤活性证据。
▲图源“Clin Cancer Res”,版权归原作者所有,如无意中侵犯了知识产权,请联系我们删除
综上,采用短程体外操作的DC疫苗联合TCR工程T细胞的双细胞疗法具有可行性,且展现出明确的抗肿瘤活性。
非病毒TCR-T疗法成“精准抗癌导弹”,肺癌患者12周即达部分缓解,病灶显著缩小甚至消失!
在第六届国际癌症免疫治疗会议(CICON)上,公布了一项“首次运用基于非病毒Sleeping Beauty系统制备的TCR-T细胞疗法成功治疗实体瘤”的突破性案例。
该患者是一位34岁KRAS G12D突变的肺腺癌女性,初诊后接受左下肺叶切除术,随后进行4个周期的顺铂联合长春瑞滨术后辅助化疗,仍出现肺部疾病复发。此后,患者尝试卡铂+培美曲塞+帕博利珠单抗联合治疗,病情缓解后改为培美曲塞+帕博利珠单抗维持治疗,最终病情再度恶化。在此情况下,患者加入临床试验,接受KRAS突变特异性TCR-T细胞疗法——该疗法采用非病毒Sleeping Beauty系统制备TCR-T细胞,具有TCR表达量高、纯度高的优势。
治疗结果显示:患者在接受淋巴细胞清除化疗及TCR-T细胞输注后,
右下叶病灶彻底消失
(详见下图1),
右上肺病灶明显缩小
(详见下图2),
右肺门淋巴结肿大缓解,
且
右上肺原本因难以测量未评估的病灶也出现缩小
(详见下图3)。治疗第12周时,患者达到
部分缓解(PR)
状态。
▲图源“alaunos”,版权归原作者所有,如无意中侵犯了知识产权,请联系我们删除
MASCT疗法创造奇迹,助转移性宫颈癌患者无癌生存超9年
全球知名期刊《Nature Communications》报道了一则“转移性宫颈鳞癌患者在接受MASCT疗法治疗后,无癌生存超9年”的突破性案例。
该患者于2011年确诊为HPV阳性转移性宫颈鳞状细胞癌,尽管接受了根治性切除术及辅助化放疗,病情仍持续进展。初次诊断33个月后,右侧骶髂关节出现转移病灶,因标准治疗失败,患者需寻求其他疗法。幸运的是,后续检查发现其携带肿瘤抗原特异性TCR(10F04),得以接受多抗原刺激细胞疗法(MASCT)——一种自体T细胞与树突状细胞(DC)结合的免疫治疗方案。
自2014年起,患者多次接受MASCT治疗,病展现出强烈的免疫应答及长期生存获益。治疗结果显示:该患者
转移性骨肿瘤得到有效控制
(详见下图),且
无癌生存期已超过9年
。
▲截图源自“nature communications”
小编寄语
TCR-T疗法创新性地开创了“抗癌+抗病毒”双效合一的治疗范式。近年来,科研团队正加速探索其在病毒相关性实体瘤领域的应用,尤其是乙肝病毒引发的肝细胞癌,以及人乳头瘤病毒(HPV)相关的宫颈癌等重点癌种。值得振奋的是,随着2024年首款TCR-T细胞治疗产品成功获批上市,标志着这一前沿技术正式迈入临床应用阶段!我们期待未来有更多突破性抗癌疗法获批,助力实体瘤患者实现长期带瘤生存的希望。
参考资料
[1]Chodon T,et al.Adoptive transfer of MART-1 T-cell receptor transgenic lymphocytes and dendritic cell vaccination in patients with metastatic melanoma. Clin Cancer Res. 2014 May 1;20(9):2457-65.
https://pmc-ncbi-nlm-nih-gov.libproxy1.nus.edu.sg/articles/PMC4070853/
[2]Long J,et al.HLA-class II restricted TCR targeting human papillomavirus type 18 E7 induces solid tumor remission in mice[J]. Nature Communications,2024, 15(1): 1-13.
https://www-nature-com.libproxy1.nus.edu.sg/articles/s41467-024-46558-4
[3]https://alaunos.com/wpcontent/uploads/2022/09/30SEP2022-CICON-vF-1.pdf
本文为全球肿瘤医生网原创,未经授权严禁转载
临床申请细胞疗法临床1期免疫疗法
2025-04-29
·今日头条
黑色素瘤是一种免疫原性高、突变负荷大且存在肿瘤间与肿瘤内异质性的癌症。尽管靶向治疗与检查点抑制剂的问世,一度为晚期黑色素瘤的治疗带来曙光,但居高不下的复发率,仍然让无数患者深陷困境。病情一旦进展,患者不仅要面对预后不佳的残酷现实,更陷入治疗手段匮乏的绝境。然而,正是黑色素瘤这种复杂而独特的免疫特性,成为了打破困局的关键突破口!基于 T 细胞的前沿疗法,如免疫检查点抑制剂、肿瘤浸润淋巴细胞疗法(TIL),在这片 “免疫战场” 上找到了治疗的可能和制胜的契机。
《美国癌症研究杂志》公布的重磅数据,无疑为抗癌征程注入一剂“强心针”!在针对 PD-1 抑制剂治疗后进展的晚期皮肤黑色素瘤患者中,非选择性自体 TIL 疗法创造了令人惊叹的战绩:疾病控制率高达 86%!这一数字不仅刷新了人们对黑色素瘤治疗的认知,更昭示着抗癌医学的重大突破 —— 曾经棘手的黑色素瘤难题,正被 TIL 疗法撕开一道希望的裂口,照亮无数患者重获新生的道路!
一、非选择性TIL疗法让晚期黑色素瘤患者持续缓解超7年,疾病控制率达86%
在黑色素瘤治疗领域,非选择性自体肿瘤浸润淋巴细胞(TIL)疗法以高度个性化优势成为创新典范。该疗法通过提取患者自身肿瘤反应性T细胞群,利用其中多样化的T细胞受体(TCR),精准锁定并攻击患者特异性肿瘤相关抗原及新抗原,为晚期皮肤黑色素瘤患者带来新希望。一项发表于《美国癌症研究杂志》的回顾性同情用药临床研究证实,非选择性TIL产品不仅可从消化处理后的肿瘤组织高效制备,更能为经检查点抑制剂或靶向治疗后病情进展的患者,开辟显著临床获益的新路径。
该研究纳入21例晚期皮肤黑色素瘤患者,中位年龄为45岁,均处于M1c或M1d期,超90%曾接受检查点抑制剂治疗,12例为PD-1抑制剂治疗后进展患者。所有患者顺利完成淋巴细胞清除化疗,其中4例使用冷冻保存的肿瘤消化液制备TIL。截至2019年12月31日,中位随访时间长达52.2个月(4.6-98.8个月),治疗数据堪称惊艳。
结果显示:所有患者的
客观缓解率(ORR)达67%,完全缓解率(CR)为19%,疾病控制率(DCR)高达86%
。所有接受治疗患者的
中位总生存期(OS)为21.3个月
(95%CI,6.8-无法估计),KM估计的
24个月总生存期(OS)率为48%
(95%CI,24-68)。
24%的患者实现TIL输注后超30个月的持续缓解,所有CR患者均维持无病生存。
▼所有接受治疗的患者(N=21)以及先前接受PD-1抑制剂治疗的患者亚组(n=12)的总生存率
▲图源“Am J Cancer Res”,版权归原作者所有,如无意中侵犯了知识产权,请联系我们删除
此外,既往接受PD-1抑制剂治疗的患者亚组中,
ORR为58%
(95%CI,28-85),其中
CR率为8%
(95%CI,0-38),
DCR为75%
(95%CI,43-95)。基线存在脑转移的7例患者,
ORR达71%
(29%CR);使用冻存肿瘤制备TIL的4例患者,
ORR达75%
(25%CR),展现出疗法强大的适应性。
值得关注的是,其中1例16岁的BRAF突变患者堪称奇迹:面对巨大病灶(病灶直径总和为103mm)、纵隔转移及脑转移,且对三种疗法耐药的绝境下,经TIL联合IL-2治疗后,
仅6周便显著减轻疾病负担,3个月实现部分缓解,60个月达成影像学完全缓解
(CR,详见下图)。截至数据截止,该
患者已持续CR超7年
(85个月),无需任何后续抗癌治疗,用生命的奇迹诠释了TIL疗法的无限潜力!
▲图源“Am J Cancer Res”,版权归原作者所有,如无意中侵犯了知识产权,请联系我们删除
二、中国TILs产品百家争鸣
除了上文提到的非选择性TIL细胞疗法外,我国紧随美国其后,多家生物公司相继布局TIL产品的研发,并在前期的临床研究中初见成效,目前多款产品已获批正式开展临床研究,这款救命的疗法对中国患者而言,终于不再是遥不可及的梦。下面全球肿瘤医生网小编汇总了其中的几款佼佼者,以供癌友们参考。
1、GC101:君赛生物
GC101是全球首个无需淋巴清除及IL-2输注的天然TIL细胞治疗产品,由君赛生物研发。2022年4月24日,获国家药品监督管理局(NMPA)的临床试验默示许可(受理号:CXSL2200070)。
GC101 TIL已展现出良好的疗效及安全性,入组患者中,有3例达到部分缓解(PR),2例获得完全缓解(CR),且CR持续时间分别>6个月、>8个月。
2、GC203:君赛生物
GC203 TIL细胞注射液是由君赛生物研发的另一款TIL细胞产品,它是全球首款基于非病毒载体而研发的TIL新药品种!2024年2月6日,国家药品监督管理局(NMPA)批准受理了该产品的临床新药试验(IND)申请。GC203延续了GC101的优势,即无需清淋治疗及IL-2治疗,患者可在普通病房完成治疗,预计将节省10万余元的配套临床费用。
GC203在治疗妇科肿瘤(如宫颈癌、子宫内膜癌、卵巢癌)方面,展现出了良好的疗效。GC203治疗后,19例可评估疗效的晚期癌症患者,肿瘤均出现明显缩小,客观缓解率(ORR)达到42.1%,疾病控制率(DCR)更是高达84.2%!
3、GT101:沙砾生物
2022年4月22日,由沙砾生物研发的GT101注射液,获国家药品监督管理局(NMPA)的临床试验默示许可(受理号:CXSL2200061),它是我国首个获批临床的TIL细胞产品!
GT101注射液在治疗黑色素瘤、非小细胞肺癌、宫颈癌等多款实体瘤领域,展现出了良好的效果。
4、GT201:沙砾生物
沙砾生物研发的第二款TIL产品-GT201,于2023年7月11日,获国家药品监督管理局(NMPA)的临床试验默示许可,这也是我国首款进入注册临床试验的基因编辑型TIL产品,适用于复发或转移性实体瘤。
5、C-TIL051:西比曼生物
西比曼生物研发的一款新型TIL产品——“C-TIL051”,新药IND申请于2022年10月,获美国FDA批准,用于PD-1抗体难治或复发性晚期非小细胞肺癌(NSCLC)的治疗。除了NSCLC外,该产品还适用于泛癌种实体瘤(如黑色素瘤、宫颈癌、头颈部鳞状细胞癌等)。
6、BST02:百吉生物
百吉生物研发的一款重磅TILs产品-BST02,Ⅰ/Ⅱ期临床试验申请于2023年10月26日,获美国食品药品监督管理局(FDA)批准,用于治疗各类肝癌。值得一提的是,BST02是全球首个进入临床阶段的治疗肝癌的TIL细胞产品。
7、ZLT-001:智瓴生物
智瓴生物自主研发的ZLT-001注射液,于2023年1月28日获NMPA的临床试验默示许可(受理号:CXSL2200552),它是华南地区首个获批临床的TIL产品,适用于晚期复发或转移性宫颈癌。
8、HV-101:天科雅生物
天科雅生物医药和杭州厚无生物医药公司联合研发的一款新型TIL产品,即“HV-101注射液”,于2023年1月29日获国家药品监督管理局(NMPA)的临床试验默示许可(受理号为:CXSL2200574),用于治疗晚期复发或转移性实体瘤。
9、HS-IT101:华赛伯曼
华赛伯曼公司研发的一款自体天然加强TIL产品——“HS-IT101注射液”,新药IND申请于2023年11月29日,获国家药品监督管理局(NMPA)批准(受理号:CXSL2300599),用于治疗晚期实体瘤。
10、LM103:蓝马医疗
“LM103注射液”是由苏州蓝马医疗技术有限公司研发的一款TIL产品,也是国内首个获临床许可,使用滋养细胞(Feeder)工艺的TIL产品。2023年7月13日,获NMPA临床试验默示许可,用于晚期实体瘤的治疗。
三、小编寄语
肿瘤浸润淋巴细胞(TIL)疗法自1988年首次应用于临床以来,已走过30余载的发展历程。相较于其他免疫细胞疗法,TIL疗法独具将患者自身肿瘤组织“变废为宝”的神奇功效,堪称实体瘤治疗领域当之无愧的抗癌黑科技。它既适用于早期癌症患者,可有效预防肿瘤的复发与转移;又能作为晚期患者的挽救性治疗手段,为癌症患者带来一线生机。
四、参考资料
[1]Pillai M,et.al.Clinical feasibility and treatment outcomes with nonselected autologous tumor-infiltrating lymphocyte therapy in patients with advanced cutaneous melanoma. Am J Cancer Res. 2022 Aug 15;12(8):3967-3984.
https://pmc-ncbi-nlm-nih-gov.libproxy1.nus.edu.sg/articles/PMC9441996/
本文为全球肿瘤医生网原创,未经授权严禁转载
细胞疗法免疫疗法临床结果
100 项与 北京天科雅生物科技有限公司 相关的药物交易
登录后查看更多信息
100 项与 北京天科雅生物科技有限公司 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2025年10月19日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
临床1期
3
2
临床2期
其他
6
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
TC-L200S | 爱泼斯坦-巴尔病毒相关鼻咽癌 更多 | 临床1/2期 |
EBV-Specific Anti-PD1 TCR-T Cells(TCRCure Biopharma) ( PD-1 ) | 头颈部鳞状细胞癌 更多 | 临床1/2期 |
TC-N201 | 晚期恶性实体瘤 更多 | 临床1期 |
H-101(TCRCure Biopharma) ( HLA-DR ) | 弥漫内生性脑桥神经胶质瘤 更多 | 临床1期 |
IL13Rα2 CAR-T(TCR CURE Biopharma) ( IL-13Rα2 ) | 胶质瘤 更多 | 临床1期 |
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
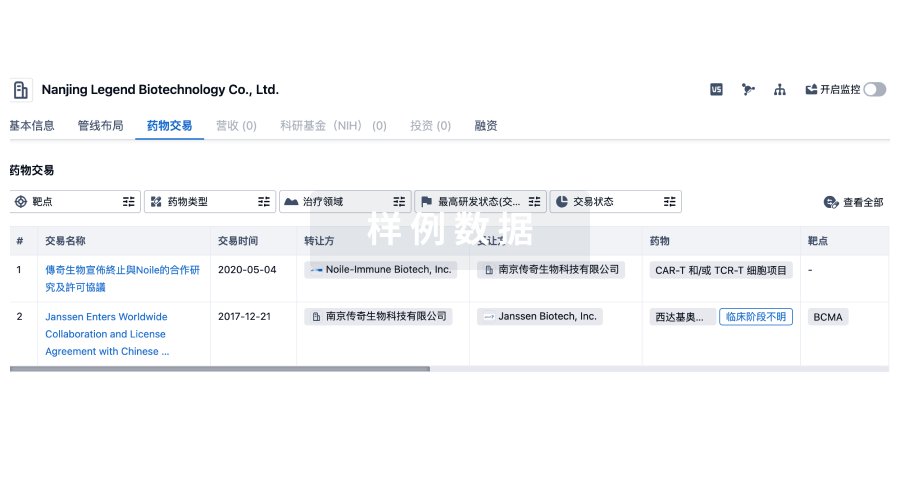
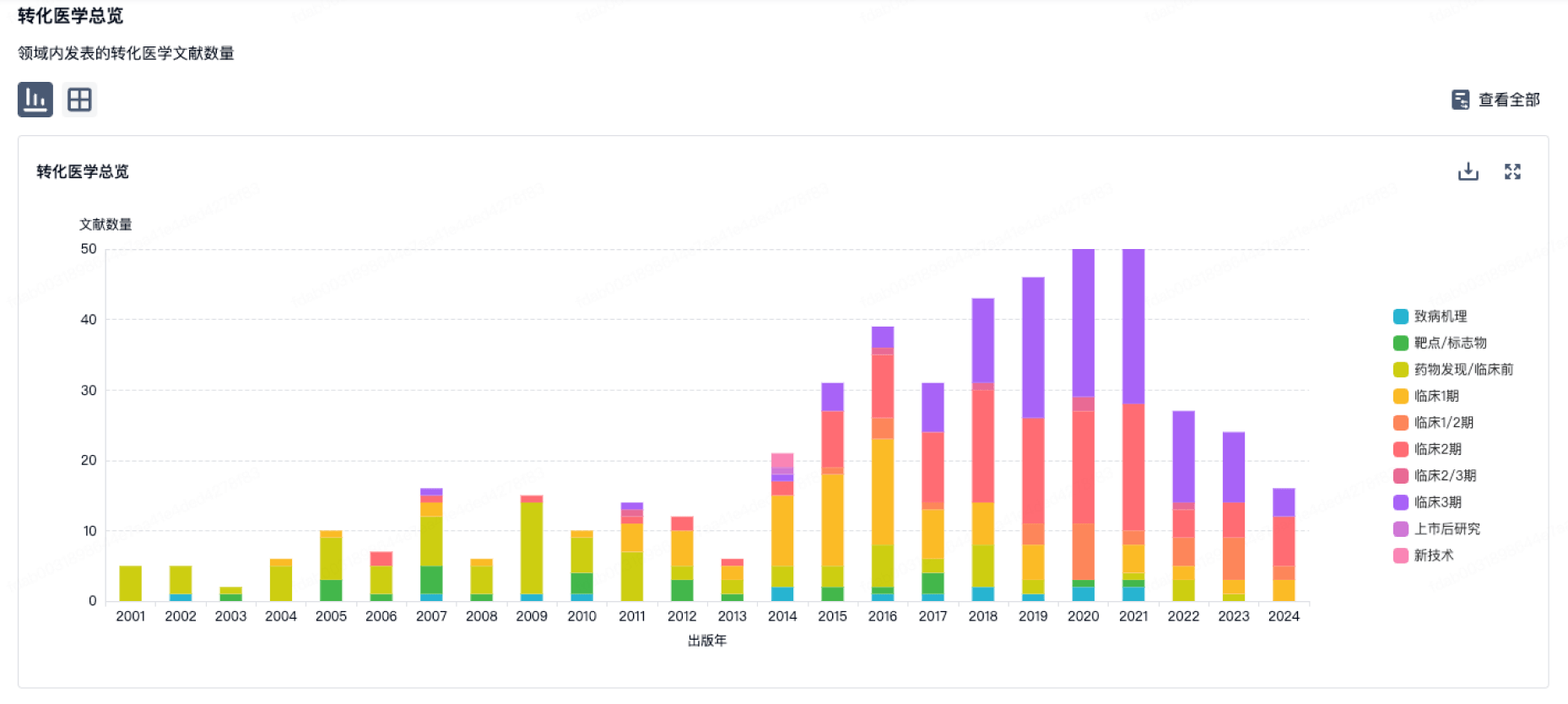
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
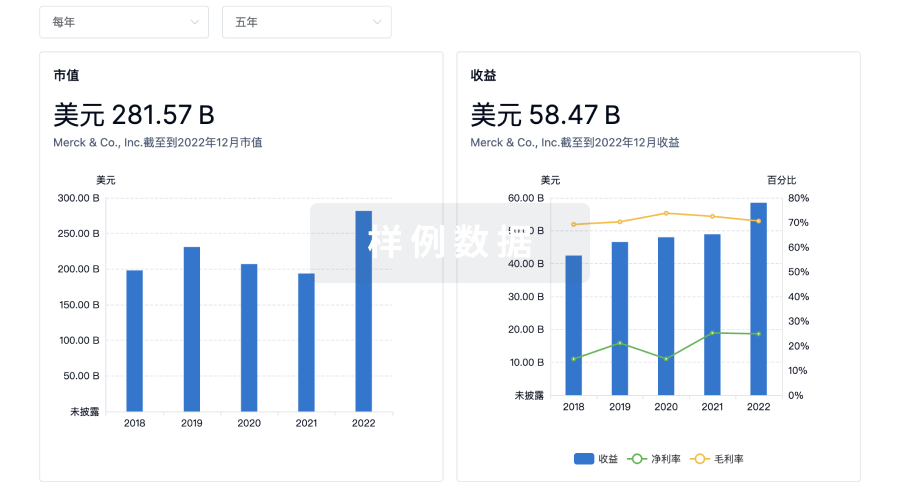
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
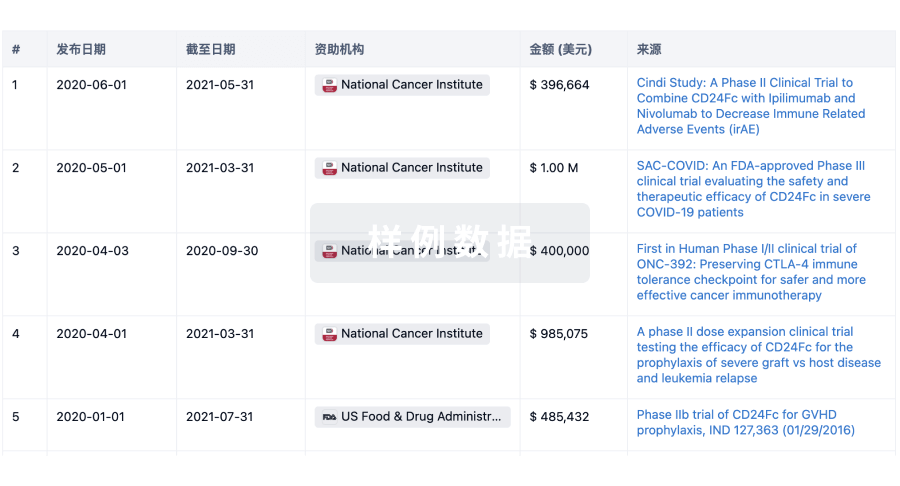
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
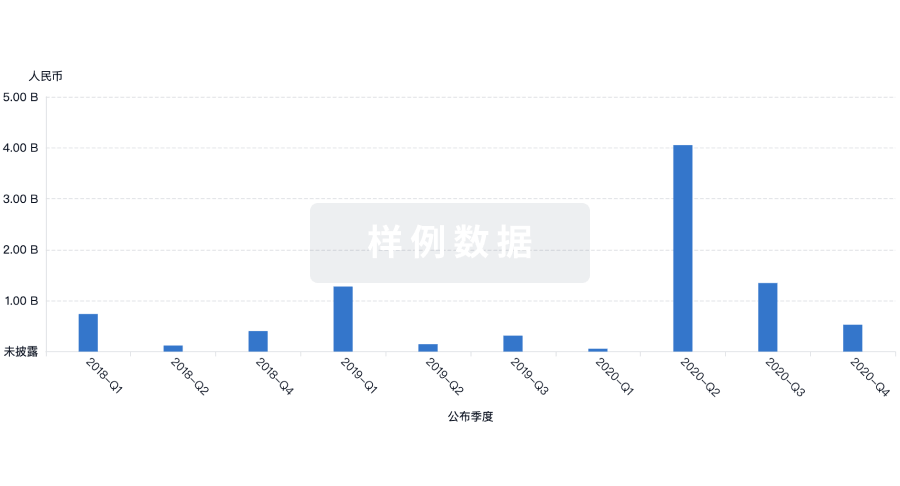
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
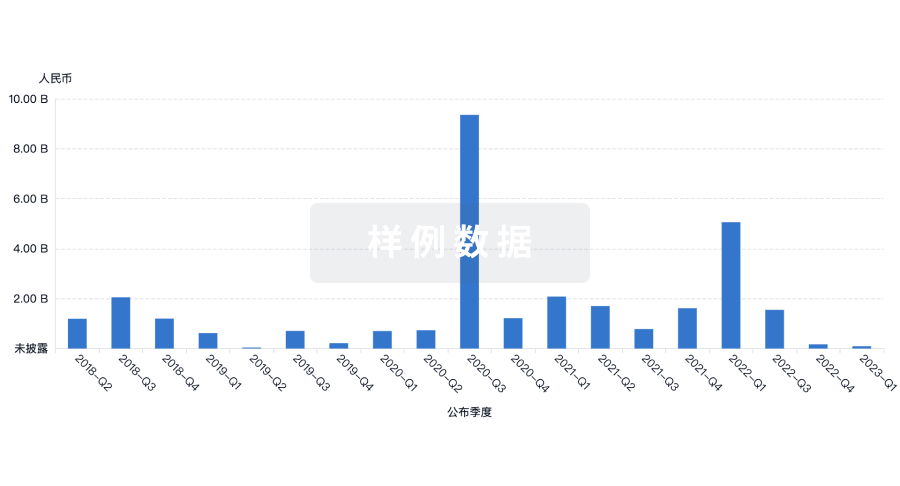
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
