预约演示
更新于:2025-09-09

Beijing Jishuitan Hospital
更新于:2025-09-09
概览
标签
皮肤和肌肉骨骼疾病
肿瘤
内分泌与代谢疾病
单克隆抗体
小分子化药
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 单克隆抗体 | 2 |
| 小分子化药 | 1 |
关联
3
项与 首都医科大学附属北京积水潭医院 相关的药物作用机制 p300-CBP转录因子抑制剂 |
在研适应症 |
非在研适应症 |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
CN116333150
专利挖掘靶点 |
作用机制- |
在研机构 |
原研机构 |
非在研适应症- |
最高研发阶段药物发现 |
首次获批国家/地区- |
首次获批日期- |
CN117264066
专利挖掘靶点 |
作用机制- |
在研机构 |
原研机构 |
非在研适应症- |
最高研发阶段药物发现 |
首次获批国家/地区- |
首次获批日期- |
223
项与 首都医科大学附属北京积水潭医院 相关的临床试验NCT07121023
A Multicenter, Randomized, Double-blind, Placebo-controlled Phase Ib Clinical Study to Evaluate the Safety, Efficacy and Kinetic Characteristics of TRD205 Tablets for Postoperative Analgesia After Unilateral Hallux Valgus Orthopedic Surgery
This is a multicenter, randomized, double-blind, placebo-controlled phase Ib study designed to evaluate the safety, efficacy and PK of TRD205 tablets administered orally .
开始日期2025-08-26 |
申办/合作机构  北京泰德制药股份有限公司 北京泰德制药股份有限公司 [+1] |
NCT07086417
A Prospective Controlled Study on Robot-Assisted Versus Conventional Arthroscopic Anterior Cruciate Ligament Reconstruction
This study aims to evaluate the clinical efficacy and biomechanical accuracy of a newly developed robot-assisted system for anterior cruciate ligament (ACL) reconstruction. ACL injuries are among the most common sports-related injuries in active individuals and athletes, and reconstruction surgery is considered the gold standard for treatment. However, failure rates remain substantial-up to 10.3% in the general population and as high as 40% among high-demand patients, such as young athletes. One of the key technical factors contributing to graft failure is improper placement of the femoral and tibial bone tunnels during surgery, which affects graft isometry, kinematics, and long-term function.
This prospective, randomized, controlled clinical trial will include 120 adult patients with confirmed ACL rupture. Subjects will be randomly assigned to two groups: an experimental group undergoing robot-assisted arthroscopic ACL reconstruction, and a control group receiving traditional arthroscopic ACL reconstruction. The investigational robot system is a custom-designed platform that integrates three primary modules: (1) preoperative planning, (2) intraoperative surgical assistance, and (3) biomechanical evaluation. The robot is designed to support precise tunnel positioning using CT/MRI fusion, dynamic motion tracking during surgery, and post-placement assessment of graft trajectory, potential impingement, and mechanical stability.
Preoperative planning includes 3D reconstruction from CT and MRI imaging to identify optimal anatomical positions for the femoral and tibial tunnels. During surgery, the robot allows the surgeon to perform the procedure in a semi-active "cooperative control" mode, combining robotic precision with the surgeon's decision-making and dexterity. Unlike existing robotic platforms, this system enables limited patient limb movement and real-time position tracking, making it uniquely suited for arthroscopic procedures.
The primary endpoint of the study is anterior tibial translation at 2 years postoperatively, measured using the KT-1000 arthrometer to compare surgical and contralateral knees. Secondary endpoints include subjective knee function scores (Lysholm, Kujala, and Tegner), CT and MRI evaluation of tunnel placement and graft integration, and physical examination results. Follow-up will occur preoperatively and at 2 years postoperatively via in-person visits.
Sample size was calculated based on differences in Lysholm scores from prior navigation-assisted studies. With an alpha level of 0.05, a power of 0.90, and a 20% expected loss to follow-up, 60 patients will be enrolled in each group. Data will be analyzed using SPSS v22.0. Parametric and non-parametric tests will be applied as appropriate, including ANOVA, Bonferroni post-hoc comparisons, and Cochran's Q for repeated categorical variables. A p-value <0.05 will be considered statistically significant.
This study is conducted by the Department of Sports Medicine at Beijing Jishuitan Hospital, a leading orthopedic and trauma care center in China, affiliated with Capital Medical University. The project is supported by the Capital Clinical Special Diagnosis and Treatment Technology Research and Translational Application Program and involves collaboration with Beijing Tiansing Bomed Medical Devices Co., Ltd., which is responsible for the robotic platform development and maintenance.
Ethical approval has been obtained from the Institutional Review Board of Beijing Jishuitan Hospital. All patients will provide written informed consent before enrollment. The study complies with the Declaration of Helsinki, Good Clinical Practice (GCP) guidelines, and local regulations.
Expected outcomes of this study include evidence of improved tunnel placement accuracy, enhanced functional recovery, and potentially reduced failure rates with robot-assisted ACL reconstruction. If successful, this research may contribute to broader adoption of robotic technology in arthroscopic sports medicine surgery and support future innovations in orthopedic surgical robotics.
This prospective, randomized, controlled clinical trial will include 120 adult patients with confirmed ACL rupture. Subjects will be randomly assigned to two groups: an experimental group undergoing robot-assisted arthroscopic ACL reconstruction, and a control group receiving traditional arthroscopic ACL reconstruction. The investigational robot system is a custom-designed platform that integrates three primary modules: (1) preoperative planning, (2) intraoperative surgical assistance, and (3) biomechanical evaluation. The robot is designed to support precise tunnel positioning using CT/MRI fusion, dynamic motion tracking during surgery, and post-placement assessment of graft trajectory, potential impingement, and mechanical stability.
Preoperative planning includes 3D reconstruction from CT and MRI imaging to identify optimal anatomical positions for the femoral and tibial tunnels. During surgery, the robot allows the surgeon to perform the procedure in a semi-active "cooperative control" mode, combining robotic precision with the surgeon's decision-making and dexterity. Unlike existing robotic platforms, this system enables limited patient limb movement and real-time position tracking, making it uniquely suited for arthroscopic procedures.
The primary endpoint of the study is anterior tibial translation at 2 years postoperatively, measured using the KT-1000 arthrometer to compare surgical and contralateral knees. Secondary endpoints include subjective knee function scores (Lysholm, Kujala, and Tegner), CT and MRI evaluation of tunnel placement and graft integration, and physical examination results. Follow-up will occur preoperatively and at 2 years postoperatively via in-person visits.
Sample size was calculated based on differences in Lysholm scores from prior navigation-assisted studies. With an alpha level of 0.05, a power of 0.90, and a 20% expected loss to follow-up, 60 patients will be enrolled in each group. Data will be analyzed using SPSS v22.0. Parametric and non-parametric tests will be applied as appropriate, including ANOVA, Bonferroni post-hoc comparisons, and Cochran's Q for repeated categorical variables. A p-value <0.05 will be considered statistically significant.
This study is conducted by the Department of Sports Medicine at Beijing Jishuitan Hospital, a leading orthopedic and trauma care center in China, affiliated with Capital Medical University. The project is supported by the Capital Clinical Special Diagnosis and Treatment Technology Research and Translational Application Program and involves collaboration with Beijing Tiansing Bomed Medical Devices Co., Ltd., which is responsible for the robotic platform development and maintenance.
Ethical approval has been obtained from the Institutional Review Board of Beijing Jishuitan Hospital. All patients will provide written informed consent before enrollment. The study complies with the Declaration of Helsinki, Good Clinical Practice (GCP) guidelines, and local regulations.
Expected outcomes of this study include evidence of improved tunnel placement accuracy, enhanced functional recovery, and potentially reduced failure rates with robot-assisted ACL reconstruction. If successful, this research may contribute to broader adoption of robotic technology in arthroscopic sports medicine surgery and support future innovations in orthopedic surgical robotics.
开始日期2025-08-01 |
申办/合作机构  首都医科大学附属北京积水潭医院 首都医科大学附属北京积水潭医院 [+1] |
NCT07149896
A Prospective Randomized Controlled Study Of Collagen Scaffold For The Repair Of Elbow Cartilage Injury
The goal of this clinical trial is to test whether adding a collagen protein scaffold can improve cartilage repair in elbow joint injuries, compared to standard surgery alone. The study will enroll 90 patients (aged 18-55) with elbow cartilage damage who haven't responded to conservative treatments.
The main questions it aims to answer are:
* Does the collagen scaffold help regenerate better-quality cartilage (measured by MRI scans at 3 and 6 months)?
* Do patients experience better pain relief and elbow function after this combined treatment?
Researchers will compare two groups:
* Experimental group : Receives microfracture surgery + collagen scaffold implant
* Active Comparator group : Receives microfracture surgery alone
Participants will:
* Undergo arthroscopic surgery (either procedure)
* Complete follow-up visits at 1 week, 1 month, 3 months, and 6 months
* Have MRI scans and functional assessments
* Report pain levels and daily activity limitations through questionnaires
The main questions it aims to answer are:
* Does the collagen scaffold help regenerate better-quality cartilage (measured by MRI scans at 3 and 6 months)?
* Do patients experience better pain relief and elbow function after this combined treatment?
Researchers will compare two groups:
* Experimental group : Receives microfracture surgery + collagen scaffold implant
* Active Comparator group : Receives microfracture surgery alone
Participants will:
* Undergo arthroscopic surgery (either procedure)
* Complete follow-up visits at 1 week, 1 month, 3 months, and 6 months
* Have MRI scans and functional assessments
* Report pain levels and daily activity limitations through questionnaires
开始日期2025-08-01 |
申办/合作机构 |
100 项与 首都医科大学附属北京积水潭医院 相关的临床结果
登录后查看更多信息
0 项与 首都医科大学附属北京积水潭医院 相关的专利(医药)
登录后查看更多信息
2,463
项与 首都医科大学附属北京积水潭医院 相关的文献(医药)2025-10-01·JOURNAL OF AFFECTIVE DISORDERS
Associations between executive functions and negative emotions: A moderated mediation analysis of emotion regulation and gender
Article
作者: Wang, Yujie ; Liu, Xiaoting ; Gao, Shuyan ; Zhu, Xiaoliang ; Zhao, Fujun ; Zhao, Xin
BACKGROUND:
Although the relationship between executive functions (EFs) and negative emotions has been well documented, the underlying mechanisms remain unclear. This study examines the potential mediating role of emotion regulation and the moderating effect of gender through a moderated mediation model.
METHODS:
Participants included 604 university students (Mage = 19.05 years, SD = 1.09; 372 females). Inhibitory control, working memory, and cognitive flexibility were assessed using the Go/No-Go task, the number running memory tasks (1750 ms and 750 ms), and the digit shifting task, respectively. Emotion regulation was measured using the Emotion Regulation Questionnaire. Anxiety and depression were assessed using the Chinese versions of the Beck Anxiety Inventory and Beck Depression Inventory. Data were analyzed using SPSS 29.0 and Mplus 8.3.
RESULTS:
Cognitive reappraisal mediated the relationship between working memory and anxiety and depression. Gender moderated the relationship between cognitive reappraisal and negative emotions. Specifically, cognitive reappraisal negatively predicted anxiety and depression only among female participants.
CONCLUSIONS:
Our findings contribute to the existing literature by clarifying the underlying mechanisms linking various subcomponents of EFs and negative emotions. The results underscore the pivotal role of emotion regulation, particularly cognitive reappraisal, and gender-specific pathways in understanding and improving university students' mental health.
2025-09-01·Bioactive Materials
Radiation cross-linked collagen scaffolds facilitate root coverage and keratinized gingival regeneration
Article
作者: Xu, Ling ; Li, Jian ; Chen, Xin ; Wang, Bozhao ; Wu, Caiming ; Li, Hongwei
The study aimed to develop a radiation cross-linked collagen scaffold (RCS) and assess its potential for root coverage and keratinized gingival regeneration, addressing the prevalent issue of gingival recession and limitations of traditional treatments. RCS was prepared through electron beam irradiation and cross-linking followed by freeze-drying. Its properties were evaluated, including Fourier transform infrared analysis, swelling behavior, microscopic observation, porosity measurement, compression modulus and structural stability. In a rat gingival recession model with 96 rats divided into four groups, the root coverage index and gingival health indices were measured, and histological analyses were conducted. The cross-linked network structure of RCS provided excellent mechanical properties and stability. In the rat model, RCS effectively promoted gingival regeneration, with the RCS group achieving a root coverage index of 87.7 ± 2.7 %, which was 54.13 %, 42.83 % and 8.41 % higher than that of the sham operation group, non-crosslinked group and chemical crosslinked group respectively. Histological analysis showed that RCS promoted anti-inflammatory macrophage polarization, enhanced collagen deposition and gingival lamina propria fiber density and increased angiogenesis. Additionally, RCS exhibited good biosafety, as blood indices and organ coefficients remained normal. In conclusion, RCS effectively promotes gingival regeneration and is a promising keratinized gingiva substitute for gingival recession, offering a new option for oral tissue repair.
2025-09-01·CLINICAL RHEUMATOLOGY
Synovial infiltrating immune cell heterogeneity associated with synovitis severity and systemic disease activity in rheumatoid arthritis: a cross-sectional study
Article
作者: Zhang, Liang ; Li, Ru ; Rao, Peishi ; Zhang, Baozhen ; Sun, Xiaolin ; Li, Zhanguo ; Jin, Shanzhao
BACKGROUND:
Rheumatoid arthritis (RA) is a systemic autoimmune disorder characterized by persistent synovitis. The synovial inflammation observed in RA exhibits significant heterogeneity among patients.
OBJECTIVE:
To examine the relationship between major immune cell infiltration profiles in synovium and both local synovitis and systemic manifestations of RA.
METHODS:
This cross-sectional study enrolled 22 RA patients (mean disease duration: 7.1 years). Synovial biopsies were assessed using the Krenn synovitis scoring system (no/low/high-grade) and immunohistochemical pathotyping (lympho-myeloid, diffuse-myeloid, pauci-immune). Clinical parameters, including the Disease Activity Score-28 (DAS28), erythrocyte sedimentation rate (ESR), C-reactive protein (CRP), and autoantibody status, were systematically analyzed.
RESULTS:
Immunohistochemical analysis identified three distinct synovial pathotypes, characterized by specific distributions of CD3⁺ T cells, CD68⁺ macrophages, CD20⁺ B cells, and CD138⁺ plasma cells. Synovitis severity, classified using the Krenn scoring system, demonstrated significant differences in synovial inflammation and stromal hyperplasia. High-grade synovitis exhibited significantly increased infiltration of CD20⁺ B cells (p = 0.025) and CD138⁺ plasma cells (p = 0.034) compared to non-inflamed tissues. Pathotype distribution analysis revealed a predominance of the pauci-immune subtype in the no-synovitis group (59.1%), whereas the lympho-myeloid subtype became progressively more prevalent with increasing synovitis severity (31.8% overall).ESR, CRP, and DAS28, were significantly correlated with synovial immune cell infiltration and synovitis severity, whereas no significant association was observed between synovitis and RA associated autoantibodies such as anti-cyclic citrullinated peptide (anti-CCP) or rheumatoid factor (RF).
CONCLUSION:
These findings underscore the complex interactions between synovial infiltrating immune cells, synovitis severity, and systemic disease activity, providing valuable insights for the development of personalized RA management strategies. Key Points •This study characterizes synovial immune infiltrates in relation to synovitis severity and systemic disease activity in RA •High-grade synovitis, as assessed by the Krenn scoring system, is associated with increased infiltration of immune cells, such as CD20⁺ B cells and CD138⁺ plasma cells, contributing to more severe inflammation and joint destruction. •The distribution of synovial pathotypes (lympho-myeloid, diffuse-myeloid, pauci-immune) correlates with synovitis severity, providing insights into potential molecular subtypes that could influence clinical phenotypes and treatment responses in RA.
40
项与 首都医科大学附属北京积水潭医院 相关的新闻(医药)2025-08-04
编者按:2025年欧洲肿瘤内科学会(ESMO)年会将于10月17日~21日在德国柏林召开。作为肿瘤学界的年度盛典,ESMO每年都会吸引上万名世界各地的肿瘤学大咖,为全球学者提供了良好的交流平台。目前公布的摘要标题中,入选的中国肉瘤及黑色素瘤研究共有19篇,本文将提供目前已公布的研究信息。
01
口头报告
摘要号:2684O
英文标题:ARTEMIS-002: A Phase 2 Study of HS-20093 in Patients with Relapsed or Refractory Sarcomas
中文标题:ARTEMIS-002 - 一项HS-20093治疗复发或难治性肉瘤患者的2期研究。
讲者:谢璐教授 北京大学人民医院
当地时间:Sun, 19.10.2025 16:30 - 18:00
地点:Dortmund Auditorium - Hall 7.1a
摘要号:2689MO
英文标题:A phase II trial of the combination of Chidamide and Toripalimab in patients with advanced sarcoma
中文标题:一项西达本胺联合特瑞普利单抗治疗晚期肉瘤患者的2期临床研究
讲者:张星教授 中山大学肿瘤防治中心
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
摘要号:2690MO
英文标题:Extended efficacy and safety from the Phase 3 MANEUVER trial of pimicotinib in patients with tenosynovial giant cell tumour (TGCT)
中文标题:Pimicotinib在腱鞘巨细胞瘤(TGCT)患者的3期MANEUVER研究中疗效和安全性延长数据
讲者:牛晓辉教授 首都医科大学附属北京积水潭医院
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
02
肉瘤壁报
摘要号:2695P
英文标题:International Multicenter Retrospective Study of Spindle cell/ Sclerosing Rhabdomyosarcoma (S-RMS), A PUSH Platform Study
中文标题:梭形细胞/硬化性横纹肌肉瘤(S-RMS)的国际、多中心回顾性研究 - 一项PUSH平台研究
讲者:陈伟武教授 中国台湾大学医学院附设医院
摘要号:2701P
英文标题:Estrogen receptor and AKT pathways as potential novel targets for MDM2-amplified well-differentiated and dedifferentiated liposarcoma (WD/DD LPS)
中文标题:以雌激素受体和AKT通路作为MDM2扩增型高分化和去分化脂肪肉瘤(WD/DD LPS)的潜在治疗新靶点
讲者:陈伟武教授 中国台湾大学医学院附设医院
摘要号:2703P
英文标题:Comparison of Open Thoractomy, Mini-open Thoracoscopically Assisted Thoracotomy, and Video-assisted Thoracoscopic Surgery in Pulmonary Metastatic Oseosarcoma: Result of 20-year Experience at Ruijin Hospital
中文标题:开胸手术、微创胸腔镜辅助开胸手术和电视辅助胸腔镜手术在肺转移性骨肉瘤中的比较:瑞金医院20年经验分析结果
讲者:Zhusheng Zhang 上海交通大学医学院附属瑞金医院
摘要号:2704P
英文标题:Circulating exosomal PD-L1 at initial diagnosis predicts outcome and survival of patients with osteosarcoma
中文标题:初始诊断时循环外泌体PD-L1可预测骨肉瘤患者的结局和生存
讲者:Jun Wang 中国北京
摘要号:2707P
英文标题:Machine learning-based multimodal model for predicting early relapse in osteosarcoma patients following neoadjuvant chemotherapy
中文标题:基于机器学习的多模态模型可预测骨肉瘤患者新辅助化疗后早期复发
讲者:傅宇成 上海交通大学医学院附属瑞金医院
摘要号:2708P
英文标题:Utidelone in Refractory Advanced or Metastatic Soft Tissue Sarcoma (UTISARC): Updated Analysis of Efficacy and Safety
中文标题:优替德隆治疗难治性晚期或转移性软组织肉瘤(UTISARC)的疗效和安全性更新分析
讲者:刘杰 四川大学华西医院
摘要号:2714P
英文标题:The prognostic value of pretreatment blood monocyte ratio in primary osteosarcoma: A multicenter study of 449 cases after neoadjuvant therapy
中文标题:治疗前血清单核细胞比率在原发性骨肉瘤中的预后价值:一项449例新辅助治疗后病例的多中心研究。
讲者:傅宇成 上海交通大学医学院附属瑞金医院
摘要号:2716P
英文标题:Efficacy and safety of Surufatinib in patients with Advanced soft tissue sarcoma After Failure of Anthracycline Chemotherapy and Prior Effective Antiangiogenic Therapy: A Single-Arm, Prospective, Exploratory Phase II Study
中文标题:索凡替尼治疗蒽环类化疗失败且既往抗血管生成治疗有效的晚期软组织肉瘤患者的疗效和安全性:一项单臂、前瞻性、探索性2期研究
讲者:张晓伟教授 复旦大学附属肿瘤医院
摘要号:2722P
英文标题:Efficacy and safety of eribulin combined with anlotinib in patients with advanced liposarcoma and leiomyosarcoma
中文标题:艾日布林联合安罗替尼治疗晚期脂肪肉瘤和平滑肌肉瘤患者的疗效和安全性
讲者:叶挺教授 华中科技大学同济医学院附属协和医院
摘要号:2724P
英文标题:Clinical, Genomic, and Transcriptomic Characterization of Cardiac Angiosarcoma
中文标题:心脏血管肉瘤的临床、基因组和转录组特征
讲者:Liu Yuan 中国北京
摘要号:2740P
英文标题:Clinicopathological Characteristics and Prognositic Analysis of Malignant Phyllodes Tumor of the Breast: A Retrospective Study
中文标题:乳腺恶性叶状肿瘤的临床病理特征与预后分析:一项回顾性研究
讲者:Yiran Zhou 中国北京
摘要号:2754ep
英文标题:Targeting SPP1 with ATRA Represents a Promising Therapeutic Strategy for Osteosarcoma
中文标题:以ATRA靶向SPP1代表了一种有前景的骨肉瘤治疗策略
讲者:Hao Fu 中国杭州
03
黑色素瘤壁报
摘要号:1626P
英文标题:DNV3 Plus Toripalimab and Chemotherapy in Advanced Melanoma: An update on an Open-Label Investigator-initiated Trial
中文标题:DNV3联合特瑞普利单抗及化疗治疗晚期黑色素瘤:一项研究者发起的单臂临床研究数据更新
讲者:林晶教授 福建省肿瘤医院
摘要号:1627P
英文标题:Toripalimab Plus Bevacizumab and Chemoradiotherapy as Treatment in Patients With Advanced Melanoma: The IRAC Phase 2 Nonrandomized Clinical Trial
中文标题:特瑞普利单抗联合贝伐珠单抗和放化疗治疗晚期黑色素瘤:2期非随机临床研究IRAC
讲者:邹征云教授 南京鼓楼医院
摘要号:1652P
英文标题:A Phase I Clinical Trial of Autologous Tumor-Infiltrating Lymphocytes FAST-TIL (HS-IT101) with low-dose IL-2 for the Treatment of Advanced Solid Tumors: Subgroup Analysis of Melanoma
中文标题:一项自体肿瘤浸润淋巴细胞FAST-TIL (HS-IT101) 联合低剂量IL-2治疗晚期实体瘤的1期临床研究:黑色素瘤亚组分析
讲者:姜愚教授 四川大学华西医院
摘要号:1677P
英文标题:Recombinant human adenovirus type 5 (H101) Combined with Anti-PD-1 Therapy in Advanced Melanoma Resistant to PD-1 Inhibitors: Mechanistic Insights from Spatial Tumor Microenvironment Analysis
中文标题:重组人5型腺病毒(H101)联合抗PD-1疗法用于PD-1抑制剂耐药的晚期黑色素瘤:基于肿瘤微环境空间分析的机制探索
讲者:林晶教授 福建省肿瘤医院
(来源:《肿瘤瞭望》编辑部)
声 明
凡署名原创的文章版权属《肿瘤瞭望》所有,欢迎分享、转载。本文仅供医疗卫生专业人士了解最新医药资讯参考使用,不代表本平台观点。该等信息不能以任何方式取代专业的医疗指导,也不应被视为诊疗建议,如果该信息被用于资讯以外的目的,本站及作者不承担相关责任。
临床结果临床2期
2025-08-04
编者按:2025年欧洲肿瘤内科学会(ESMO)年会将于10月17日~21日在德国柏林召开。作为肿瘤学界的年度盛典,ESMO每年都会吸引上万名世界各地的肿瘤学大咖,为全球学者提供了良好的交流平台。目前,ESMO官网已公布部分口头报告标题,肿瘤瞭望特别整理了肉瘤与黑色素瘤领域目前公布的口头报告内容,分享给广大读者;此外,入选LBA的研究摘要标题则将在9月20日公布,届时也将第一时间进行分享。
01
中国专家口头报告
摘要号:2684O
英文标题:ARTEMIS-002: A Phase 2 Study of HS-20093 in Patients with Relapsed or Refractory Sarcomas
中文标题:ARTEMIS-002 - 一项HS-20093治疗复发或难治性肉瘤患者的2期研究。
讲者:谢璐教授 北京大学人民医院
当地时间:Sun, 19.10.2025 16:30 - 18:00
地点:Dortmund Auditorium - Hall 7.1a
摘要号:2689MO
英文标题:A phase II trial of the combination of Chidamide and Toripalimab in patients with advanced sarcoma
中文标题:一项西达本胺联合特瑞普利单抗治疗晚期肉瘤患者的2期临床研究
讲者:张星教授 中山大学肿瘤防治中心
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
摘要号:2690MO
英文标题:Extended efficacy and safety from the Phase 3 MANEUVER trial of pimicotinib in patients with tenosynovial giant cell tumour (TGCT)
中文标题:Pimicotinib在腱鞘巨细胞瘤(TGCT)患者的3期MANEUVER研究中疗效和安全性延长数据
讲者:牛晓辉教授 首都医科大学附属北京积水潭医院
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
02
肉瘤
摘要号:2685O
英文标题:Phase II of sunitinib plus nivolumab in advanced alveolar soft-part sarcoma results from the GEIS, ISG and UCL IMMUNOSARC II study
中文标题:舒尼替尼联合纳武利尤单抗治疗晚期肺泡状软组织肉瘤的2期研究结果,来自GEIS、ISG和UCL IMMUNOSARC II研究
讲者:Nadia Hindi Muñiz (Madrid, Spain)
当地时间:Sun, 19.10.2025 16:30 - 18:00
地点:Dortmund Auditorium - Hall 7.1a
摘要号:2686O
英文标题:EFTISARC-NEO: A phase II study of neoadjuvant eftilagimod alpha, pembrolizumab and radiotherapy in patients with resectable soft tissue sarcoma
中文标题:EFTISARC-NEO:一项eftilagimod alpha、帕博利珠单抗和放疗新辅助治疗可切除软组织肉瘤患者的2期研究
讲者:Katarzyna Kozak (Warsaw, Poland)
当地时间:Sun, 19.10.2025 16:30 - 18:00
地点:Dortmund Auditorium - Hall 7.1a
摘要号:2687MO
英文标题:CONgRAtS: a randomized phase II study of nivolumab±relatlimab in patients with TLS-positive soft-tissue sarcomas
中文标题:CONgRAtS:一项纳武利尤单抗±relatlimab治疗TLS阳性软组织肉瘤患者的随机2期研究
讲者:Florent Peyraud (Bordeaux, France)
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
摘要号:2688MO
英文标题:A phase 1/2 trial of FOG-001, a first-in-class direct β-catenin:TCF4 inhibitor: safety and preliminary antitumor activity in patients with desmoid tumors
中文标题:一项FOG-001(一种首创直接β-连环蛋白:TCF4抑制剂)的1/2期临床研究:在硬纤维瘤患者中的安全性和初步抗肿瘤活性
讲者:Gregory M. Cote (Boston, United States of America)
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
摘要号:2691MO
英文标题:Twenty year survival of advanced gastrointestinal stromal tumours treated with imatinib, Exploratory long-term follow-up of the BFR14 trial
中文标题:伊马替尼治疗晚期胃肠道间质瘤的二十年生存数据:BFR14试验探索性长期随访
讲者:Quentin Devin (Lyon, France)
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
摘要号:2692MO
英文标题:Efficacy and safety of Metronomic Cyclophosphamide (MC) VS Doxorubicin (DOXO) as first line treatment in older patients (PTS) with advanced soft tissue sarcomas (STS) - The METROPHOLYS trial.
中文标题:节拍环磷酰胺对比多柔比星作为晚期软组织肉瘤(STS)老年患者一线治疗的疗效和安全性 - METROPHOLYS研究
讲者:Antonella Brunello (Padova, Italy)
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
摘要号:2693MO
英文标题:COTESARC - A multicentre, open-label, Phase I-II evaluating the combination of MEK and PDL-1 inhibitors in patients with advanced soft tissue sarcoma (STS)
中文标题:COTESARC研究 - 一项评估MEK和PD-L1抑制剂联合治疗晚期软组织肉瘤(STS)患者的多中心、开放标签1-2期临床试验。
讲者:Armelle Dufresne (Lyon, France, CEDEX 3)
当地时间:Fri, 17.10.2025 16:00 - 17:30
地点:Essen Auditorium - Hall 7.2a
03
黑色素瘤
摘要号:1600O
英文标题:Efficacy and safety of IMA203, a PRAME-directed T-cell receptor (TCR) T-cell therapy, in patients with previously treated advanced or metastatic uveal melanoma from a Ph 1 trial
中文标题:靶向PRIME的T细胞受体(TCR)T细胞疗法IMA203,治疗既往经治晚期或转移性葡萄膜黑色素瘤患者的1期临床试验疗效和安全性结果
讲者:Sapna P. Patel (Aurora, United States of America, TX)
当地时间:Mon, 20.10.2025 08:30 - 10:00
地点:Dortmund Auditorium - Hall 7.1a
摘要号:1601O
英文标题:3-year survival with neoadjuvant-adjuvant pembrolizumab from SWOG S1801
中文标题:来自SWOG1801研究的帕博利珠单抗新辅助-辅助治疗的3年生存数据
讲者:Sapna P. Patel (Aurora, United States of America, TX)
当地时间:Mon, 20.10.2025 08:30 - 10:00
地点:Dortmund Auditorium - Hall 7.1a
摘要号:1602O
英文标题:Enucleation prevention and vision preservation in primary uveal melanoma (UM): preliminary results from a phase 2 study of neoadjuvant darovasertib
中文标题:原发性葡萄膜黑色素瘤的预防眼球摘除和视力保留:来自darovasertib新辅助治疗2期研究的初步结果
讲者:Marcus O. Butler (Toronto, Canada, Ontario)
当地时间:Mon, 20.10.2025 08:30 - 10:00
地点:Dortmund Auditorium - Hall 7.1a
摘要号:1604MO
英文标题:Randomization to Adjuvant Nivolumab or Ipilimumab + Nivolumab Based on Pathological Response to A Single Dose of Neoadjuvant Nivolumab in Stage III melanoma
中文标题:基于单剂纳武利尤单抗新辅助治疗的病理应答,随机使用纳武利尤单抗或伊匹木单抗+纳武利尤单抗辅助治疗III期黑色素瘤患者的研究
讲者:John Miura (Philadelphia, United States of America)
当地时间:Sat, 18.10.2025 14:45 - 16:15
地点:Koblenz Auditorium - Hall 5.2
摘要号:1605MO
英文标题:Primary results from a randomized phase 2 trial of BNT111 in combination with cemiplimab with calibrator monotherapy arms in anti-PD-(L)1 relapsed/refractory melanoma
中文标题:BNT111联合cemiplimab(且设有校准单药治疗组)治疗抗PD-(L)1经治/难治性黑色素瘤患者的随机2期试验初步结果
讲者:Paolo A. Ascierto (Naples, Italy)
当地时间:Sat, 18.10.2025 14:45 - 16:15
地点:Koblenz Auditorium - Hall 5.2
(来源:《肿瘤瞭望》编辑部)
声 明
凡署名原创的文章版权属《肿瘤瞭望》所有,欢迎分享、转载。本文仅供医疗卫生专业人士了解最新医药资讯参考使用,不代表本平台观点。该等信息不能以任何方式取代专业的医疗指导,也不应被视为诊疗建议,如果该信息被用于资讯以外的目的,本站及作者不承担相关责任。
临床2期临床结果临床3期临床终止
2025-07-29
点击“蓝字”关注我们编者按2型糖尿病(T2DM)相关慢性肾脏病(CKD)常伴随血流动力学改变、代谢紊乱及炎症纤维化,共同导致不良心肾结局。在管理中,相较于调控血流动力学和强化控糖,抗炎抗纤维化是相对短板。随着非甾体类盐皮质激素受体拮抗剂(nsMRA,如非奈利酮)等创新药物的突破,T2DM相关CKD的早期干预取得重要进展。因此,早期筛查、联合干预及主动风险管理以改善患者长期心肾预后,已成为临床迫切需求。在近期于京沪两地成功举办的“肾知泌解——2025糖尿病合并肾病临床管理推进项目”巡讲中,国内外内分泌及肾病领域权威专家汇聚一堂,共同探讨T2DM相关CKD管理的前沿进展。会议特邀国际专家美国代谢研究所Yehuda Handelsman教授,从全球视角剖析T2DM相关CKD的疾病负担与诊疗挑战,并深入解读nsMRA非奈利酮在心肾保护中的关键作用及循证证据。本篇将重点呈现全球视角下的T2DM相关CKD疾病认知与管理策略革新。全球视角:nsMRA用于糖尿病合并CKD治疗——改善心肾结局的关键支柱Yehuda Handelsman教授在报告中强调,糖尿病已成为全球重大健康威胁,2019年影响4.63亿成人,预计2045年将突破7亿[1]。其严重后果不仅在于高血糖本身,更在于并发症——约40%的T2DM患者会进展为糖尿病肾病(DKD),而DKD占所有CKD病例的一半;更严峻的是,90%的DKD患者最终死于心力衰竭或动脉粥样硬化性心血管事件,仅少数(10%)进展至肾衰竭[2]。白蛋白尿是预测心肾疾病结局的关键早期指标,尿白蛋白肌酐比值(UACR)升高和估算肾小球滤过率(eGFR)降低均显著增加心血管死亡和肾脏事件风险,且风险随CKD严重程度递增[3,4],凸显早期干预的迫切性。01靶向心肾损伤重要机制:非奈利酮抗炎抗纤维化,带来治疗新突破尽管采用标准治疗,T2DM合并CKD患者仍面临CKD进展高风险[5-7],驱动因素包括血流动力学异常、代谢紊乱以及炎症纤维化[8-10]。前两者已有多种疗法,但针对炎症纤维化的治疗长期缺失。盐皮质激素受体(MR)在肾脏、心脏及血管组织等广泛分布,其过度激活通过稳态失调及炎症纤维化进程导致心肾损伤。非奈利酮作为一种新型选择性nsMRA,精准靶向MR过度激活,发挥抗炎抗纤维化作用,提供心肾双重保护(图1),现已成为T2DM相关CKD患者的重要治疗选择。图1. 非奈利酮:一种新型选择性nsMRA,阻断MR过度激活02里程碑式研究奠定心肾保护基石:非奈利酮的循证之路FIGARO-DKD[11,12]和FIDELIO-DKD[13,14]两项里程碑研究共同证实了非奈利酮在T2DM合并CKD患者中的心肾保护作用。其汇总分析FIDELITY[15]提供了该领域规模最大(>13 000例患者)、覆盖人群最广(涵盖不同CKD分期)的Ⅲ期临床证据,夯实了其卓越疗效(图2)和安全性:显著降低肾脏复合结局风险达23%;显著降低心血管复合结局风险14%,且独立于基线肾功能或SGLT2i/GLP-1RA使用;对血压影响轻微,不影响血糖,高钾血症发生率虽有增加但导致停药罕见。<< 滑动查看下一张图片 >>图2. FIDELITY汇总分析:非奈利酮显著降低肾心结局风险纳入FIDELIO-DKD、FIGARO-DKD和FINEARTS-HF的进一步汇总分析FINE-HEART研究[16]表明,非奈利酮可为广泛心血管-肾脏-代谢(CKM)综合征患者带来多重获益(图3),包括降低心血管事件、肾脏复合结局风险以及全因死亡风险,且安全性和耐受性良好,确立了其在CKM综合征管理中的重要地位。图3. FINE-HEART预设疗效终点03突破性联合策略:CONFIDENCE研究引领早期强化治疗新格局近期公布的CONFIDENCE研究[17]标志着T2DM合并CKD治疗策略的重大革新,证实同步起始非奈利酮与SGLT2i联合治疗具有疗效叠加效应:早期快速、显著降低UACR:联合治疗组14天内UACR降幅即超过30%,180天时降幅达52%,显著优于任一单药治疗(图4);安全性良好且可控:导致停药的高钾血症少见。研究观察到起始联合后出现的初始eGFR下降可预测,且在停药后基本可逆。图4. 同步起始联合非奈利酮与SGLT2i治疗疗效叠加,优于任一单药治疗CONFIDENCE研究为临床实践提供了关键证据:对于T2DM合并CKD患者,在疾病早期即同步启动非奈利酮与SGLT2i联合治疗,能够实现UACR的快速、显著和协同性降低,最大化心肾保护潜力,且安全性可控。这确立了早期、主动的“双支柱”联合策略作为优化T2DM合并CKD患者预后的重要选择。04未来展望:非奈利酮持续拓展心肾保护疆域非奈利酮的探索仍在深入,多项进行中的研究(如FINEOVATE、MOONRAKER、THUNDERBALL、FINE-ONE、FIND-CKD、FIONA)涵盖各类肾脏疾病合并心衰、心衰、CKD、CKD合并1型糖尿病(T1DM)、非糖尿病性CKD以及CKD儿童和青少年等广泛人群,有望重塑更广泛人群的心肾保护格局。北京场专家讨论:非奈利酮临床实践与研究方向在讨论环节中,首都医科大学附属北京积水潭医院霍丽丽教授结合临床实践指出,当前非奈利酮20 mg标准剂量应用比例仍有提升空间,10 mg剂量在真实世界中已展现良好疗效;既往多采用序贯治疗模式,但随着循证证据积累,起始联合SGLT2i等治疗策略具有可行性。首都医科大学宣武医院孙宇教授则展望未来研究方向,强调FINE-ONE研究对探索非奈利酮在T1DM合并CKD患者中的治疗价值具有重要意义,有望填补该领域治疗空白。两位专家一致认为,非奈利酮的临床应用需结合剂量优化、联合治疗及适应症拓展,以最大化患者心肾获益。结语本次巡讲通过国际权威专家的视角,清晰描绘了T2DM相关CKD带来的沉重疾病负担及早期干预的紧迫性。非奈利酮作为创新nsMRA,直击炎症纤维化核心机制,其强大的心肾保护作用已被FIGARO-DKD、FIDELIO-DKD、FIDELITY及FINE-HEART等里程碑式研究以及CONFIDENCE研究展现的早期联合优势所证实,为全球患者提供了改善长期预后的关键武器。专家讨论进一步解析了非奈利酮的优化应用策略,以及对治疗模式转变和未来探索方向的见解。下篇报道将聚焦中国经验,深入探讨如何在“见微知危”理念下实现糖尿病相关CKD的早期筛查与强化管理,敬请期待!会场掠影参考文献(上下滑动可查看)1. International Diabetes Federation. IDF Diabetes Atlas. 9th edn.2. Tuttle KR, et al. Clin J Am Soc Nephrol. 2022; 17(7): 1092-1103.3. Ninomiya T, et al. J Am Soc Nephrol. 2009; 20(8): 1813-1821.4. KDIGO Diabetes Work Group. Kidney Int. 2020; 98(suppl): S1-S115.5. Brenner BM, et al. N Engl J Med. 2001; 345(12): 861-869.6. Wheeler DC, et al. Lancet Diabetes Endocrinol. 2021; 9(1): 22-31.7. Wheeler DC, et al. Nephrol Dial Transplant. 2020; 35(10): 1700-1711.8. Alicic RZ, et al. Clin J Am Soc Nephrol. 2017; 12(12): 2032-2045.9. Mora-Fernández C, et al. J Physiol. 2014; 592(18): 3997-4012.10. Bauersachs J, et al. Hypertension. 2015; 65(2): 257-263.11. Pitt B, et al. N Engl J Med. 2021; 385(24): 2252-2263.12. Ruilope LM, et al. Nephrol Dial Transplant. 2023; 38(2): 372-383.13. Bakris GL, et al. N Engl J Med. 2020; 383(23): 2219-2229.14. Filippatos G, et al. Circulation. 2021; 143(6): 540-552.15. Agarwal R, et al. Eur Heart J. 2022; 43(6): 474-484.16. Vaduganathan M, et al. Nat Med. 2024; 30(12): 3758-3764.17. Agarwal R, et al. N Engl J Med. 2025 Jun 5. doi: 10.1056/NEJMoa2410659.声明:本文仅供医疗卫生专业人士了解最新医药资讯参考使用,不代表本平台观点。该等信息不能以任何方式取代专业的医疗指导,也不应被视为诊疗建议,如果该信息被用于资讯以外的目的,本站及作者不承担相关责任。最新《国际糖尿病》读者专属微信交流群建好了,快快加入吧!扫描左边《国际糖尿病》小助手二维码(微信号:guojitnb),回复“国际糖尿病读者”,ta会尽快拉您入群滴!(来源:《国际糖尿病》编辑部)版权声明版权属《国际糖尿病》所有。欢迎个人转发分享。其他任何媒体、网站未经授权,禁止转载。
临床终止临床研究诊断试剂
100 项与 首都医科大学附属北京积水潭医院 相关的药物交易
登录后查看更多信息
100 项与 首都医科大学附属北京积水潭医院 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2025年09月19日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
2
1
临床前
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
A-485 ( p300-CBP transcription factors ) | 绝经期后骨质疏松 更多 | 临床前 |
CN117264066 ( ENO1 )专利挖掘 | 营养和代谢疾病 更多 | 药物发现 |
CN116333150 ( ENO1 )专利挖掘 | 肿瘤 更多 | 药物发现 |
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
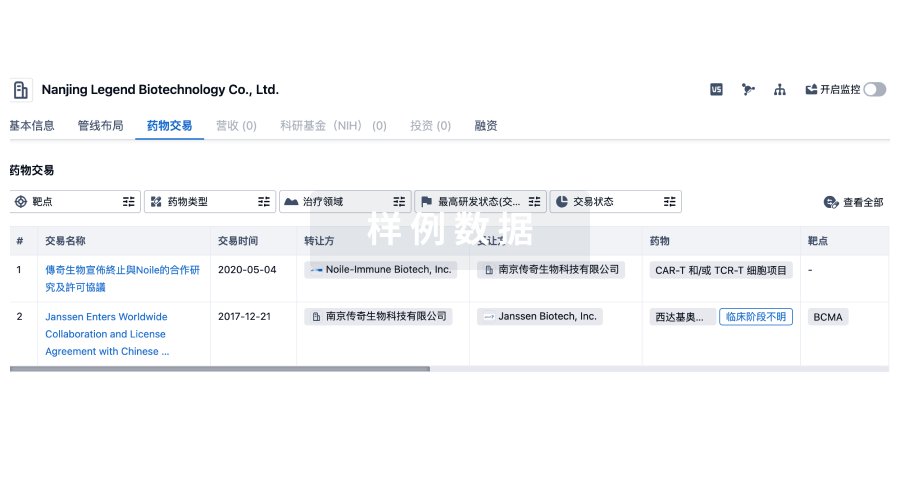
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
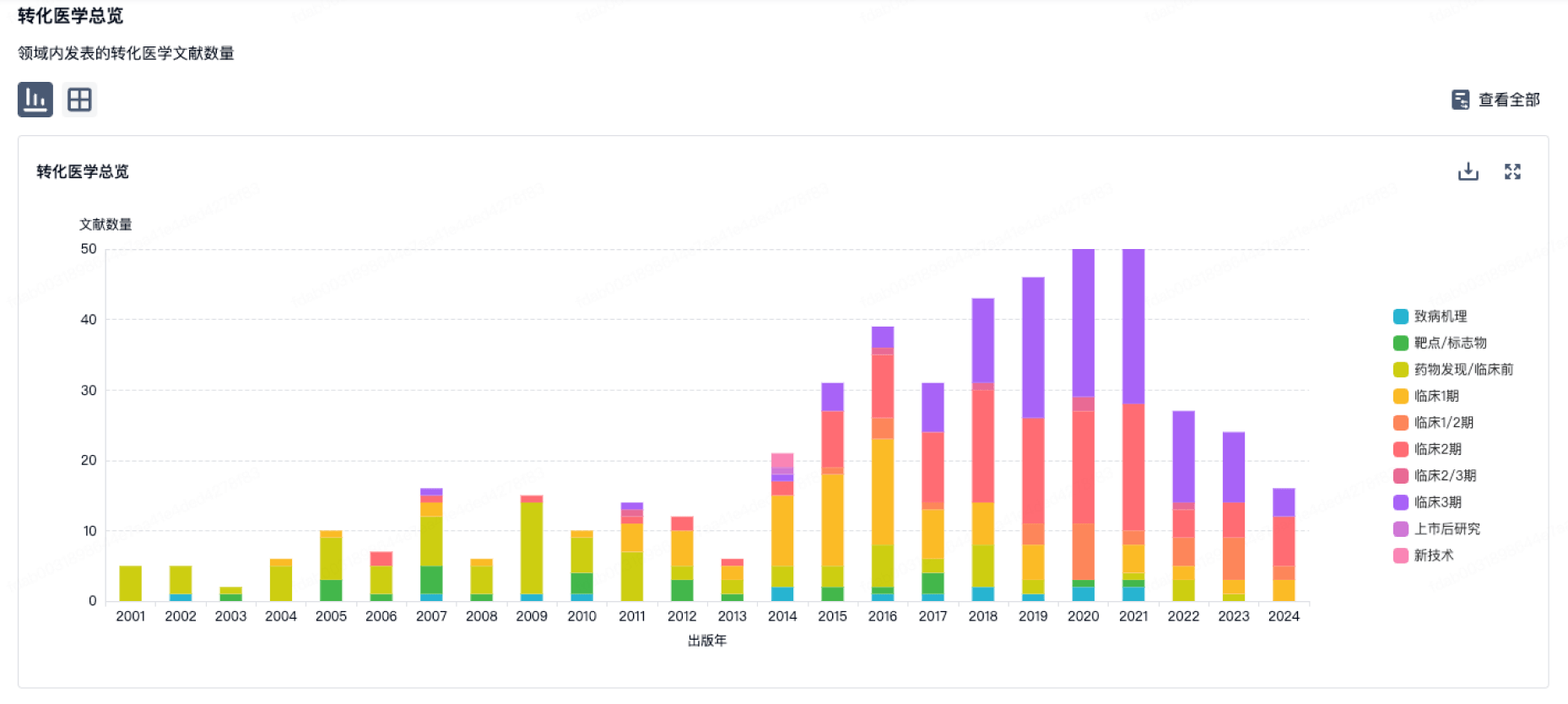
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
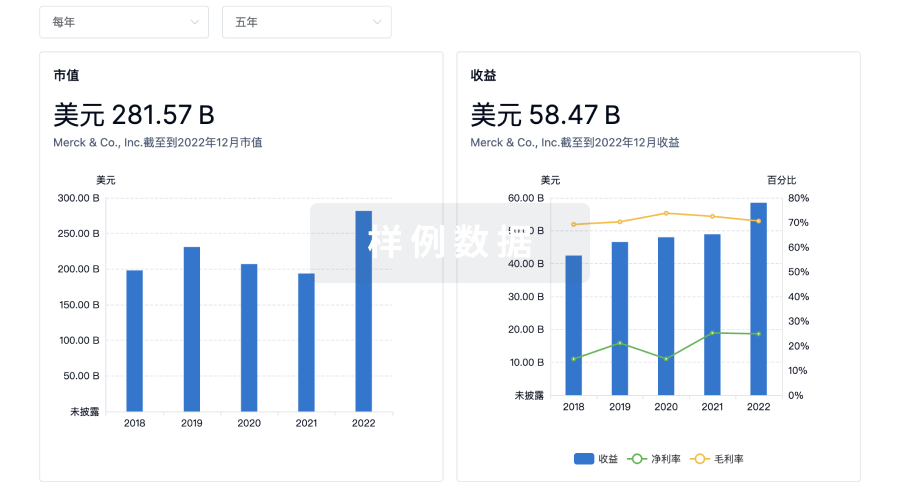
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
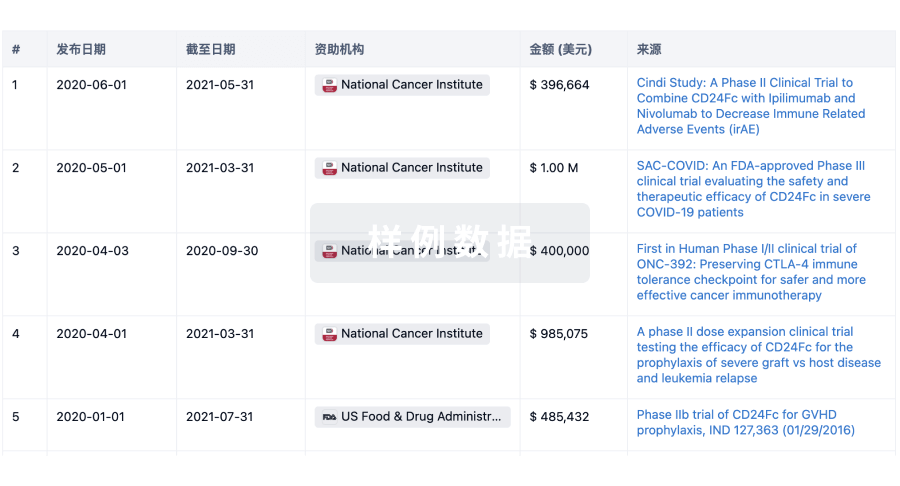
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
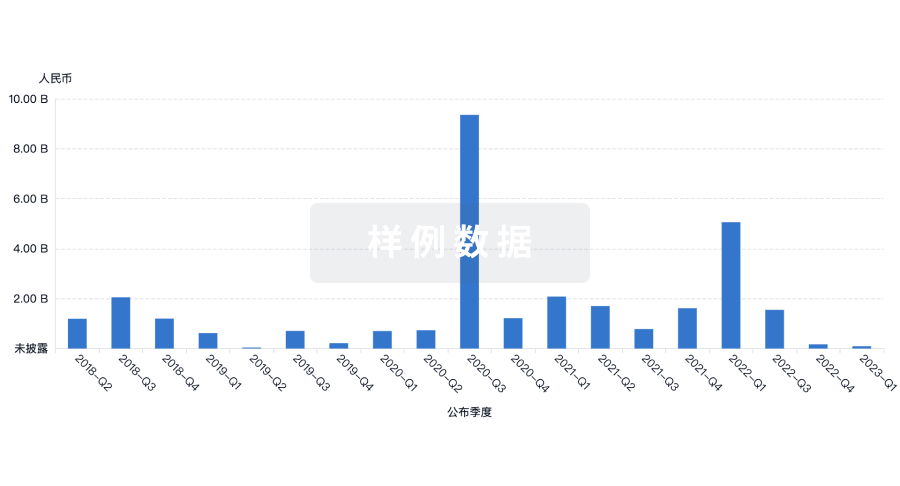
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
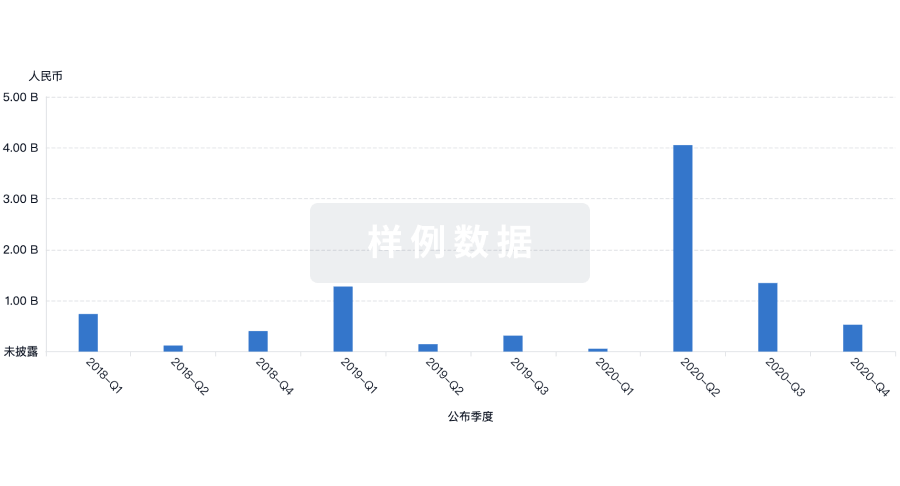
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
