预约演示
更新于:2025-01-23
PDL1 x GAS6
更新于:2025-01-23
关联
1
项与 PDL1 x GAS6 相关的药物WO2024145291
专利挖掘作用机制- |
在研机构 |
原研机构 |
在研适应症 |
非在研适应症- |
最高研发阶段药物发现 |
首次获批国家/地区- |
首次获批日期1800-01-20 |
100 项与 PDL1 x GAS6 相关的临床结果
登录后查看更多信息
100 项与 PDL1 x GAS6 相关的转化医学
登录后查看更多信息
0 项与 PDL1 x GAS6 相关的专利(医药)
登录后查看更多信息
12
项与 PDL1 x GAS6 相关的文献(医药)2023-01-01·Frontiers in oncology
Clinical and genetic characteristics in pancreatic cancer from Chinese patients revealed by whole exome sequencing.
Article
作者: Tang, Yichen ; Peng, Xuehui ; Huang, Xiaobing ; You, Nan ; He, Yonggang ; Zheng, Lu ; Li, Yuming ; Li, Jing ; Liu, Chuang ; Wu, Jing ; Huang, Wen ; Li, Ling
2022-01-01·International Review of Cell and Molecular Biology
Mertk: An emerging target in cancer biology and immuno-oncology
Article
作者: Davra, Viralkumar ; Wang, Ziren ; Calianese, David ; Lahey, Kevin C ; Desind, Samuel ; Zaman, Ahnaf ; Hill, Amanda ; Gadiyar, Varsha ; Birge, Raymond B ; Patel, Radhey
2021-04-01·Cell Death & Differentiation1区 · 生物学
KPNB1-mediated nuclear translocation of PD-L1 promotes non-small cell lung cancer cell proliferation via the Gas6/MerTK signaling pathway
1区 · 生物学
Article
作者: Huang, Jian-An ; Liu, Zeyi ; Cai, Tingting ; Zhang, Yang ; Zeng, Yuanyuan ; Zhang, Weijie ; Zhang, Ruochen ; Zhu, Jianjie ; Fu, Yulong ; Liu, Ting ; Du, Wenwen
23
项与 PDL1 x GAS6 相关的新闻(医药)2024-03-23
·生物探索
引言免疫检查点抑制剂,特别是PD-1/PD-L1抑制剂,已被批准用于不可切除的肝细胞癌(HCC)。然而,高耐药率仍然限制了它们的疗效,这突出表明迫切需要了解潜在的机制并制定克服耐药性的策略。2024年3月20日,复旦大学董琼珠、钦伦秀及陆录共同通讯在Cell Reports Medicine 在线发表题为“Disruption of MerTK increases the efficacy of checkpoint inhibitor by enhancing ferroptosis and immune response in hepatocellular carcinoma”的研究论文,该研究证明了高MER原癌基因酪氨酸激酶(MerTK)表达的HCC在两种同基因小鼠模型和接受抗PD-1/PD-L1治疗的患者中表现出抗PD-1/PD-L1耐药。在机制上,MerTK通过ERK/SP1途径通过SLC7A11的上调来限制铁死亡,并通过募集髓源性抑制细胞(MDSCs)促进免疫抑制性肿瘤微环境(TME)的发展,从而使HCC对抗PD-1/PD-L1产生耐药性。Sitravatinib是MerTK的一种抑制剂,通过促进肿瘤铁死亡和减少MDSC向TME的浸润,使耐药HCC对抗PD-L1治疗增敏。肝细胞癌(HCC)仍然是一个全球性的健康挑战,其发病率在全球范围内不断增加临床上,大多数HCC患者被诊断为晚期,治疗选择有限。最近,免疫疗法,如免疫检查点抑制剂(ICIs)抗CTLA4、抗PD-1/PD-L1、它们的联合或与其他药物的联合,在癌症患者中取得了无与伦比的成功。此外,nivolumab、pembrolizumab和atezolizumab等ICIs目前已被美国食品和药物管理局(FDA)批准用于HCC的全身治疗。然而,由于抗PD-1/PD-L1治疗经常出现耐药,反应率仍远不能令人满意,目前仅在约15% - 20%的HCC患者中观察到总体获益率。因此,迫切需要确定最佳患者选择的生物标志物,并确定克服这种耐药性的有效靶点。MER原癌基因酪氨酸激酶(MerTK)是酪氨酸受体激酶TAM家族(TYRO3、AXL和MerTK)的一员,在包括HCC在内的多种恶性肿瘤中高度表达6 (the Human Protein Atlas: proteinatlas.org)。同源蛋白生长抑制特异性蛋白6 (Gas6)和蛋白S (Pros1)都是MerTK活化的天然配体,可诱导MerTK的同二聚化和自磷酸化,激活MEK/ERK、PI3K/AKT、JAK/STAT等下游通路,从而达到肿瘤细胞增殖、抗凋亡、抑制炎症和肿瘤生长的目的。最近的研究表明,MerTK作为一种代谢调节剂,通过整合有氧糖酵解和氧化磷酸化,在调节HCC生长中起着关键作用。此外,MerTK的受体/配体活性已在多种免疫细胞类型中得到证实,并增强抗肿瘤免疫,这对癌症免疫治疗具有重要意义。模式图(Credit: Cell Reports Medicine)铁死亡作为一种代谢调节的细胞死亡形式,在癌症生物学中起着重要的作用。近年来,越来越多的证据表明,对铁死亡的抑制与对常规癌症治疗的耐药性有关,并且靶向铁死亡可能为癌症提供治疗潜力。最近,研究表明,在免疫治疗过程中,对铁死亡的调节可以重塑肿瘤生态位,从而导致T细胞介导的肿瘤根除。在HCC中,铁死亡已被证明在全身治疗(如靶向治疗和化疗)中具有耐药性。该研究发现MerTK是抵抗PD-1/PD-L1阻断的主要机制,通过增加髓源性抑制细胞(MDSC)的浸润,限制肿瘤细胞铁死亡并引起免疫抑制性肿瘤微环境(TME)。MerTK阻断可能有效对抗这种耐药,并使HCC对抗PD-L1治疗敏感。此外,MerTK可作为sitravatinib,联合抗PD-L1联合治疗筛查HCC患者的生物标志物。总之,该研究发现MerTK可以作为患者分层的预测性生物标志物,并且是克服HCC中抗PD-1/PD-L1耐药的有希望的靶点。原文链接https://doi-org.libproxy1.nus.edu.sg/10.1016/j.xcrm.2024.101420责编|探索君排版|探索君文章来源|“iNature”End往期精选围观一文读透细胞死亡(Cell Death) | 24年Cell重磅综述(长文收藏版)热文Nature | 破除传统:为何我们需要重新思考肿瘤的命名方式热文Nature | 2024年值得关注的七项技术热文Nature | 自身免疫性疾病能被治愈吗?科学家们终于看到了希望热文CRISPR技术进化史 | 24年Cell综述
免疫疗法
2024-02-24
·生物探索
引言近年来,以免疫检查点抑制剂(ICIs)为代表的肿瘤免疫治疗被认为是癌症治疗领域最成功的方法之一,但病人获益有限,甚至病人获益之后仍可出现耐药问题。2024年2月20日,复旦大学董琼珠、钦伦秀及陆录共同通讯在Cell Reports Medicine 在线发表题为“Disruption of MerTK increases the efficacy of checkpoint inhibitor by enhancing ferroptosis and immune response in hepatocellular carcinoma”的研究论文,该研究表明靶向MerTK可通过增强肝细胞癌的铁死亡和免疫应答来提高免疫检查点抑制剂的疗效。在这项研究中,研究人员证明了MER原癌基因酪氨酸激酶(MerTK)在两种ICI耐药的肝癌小鼠模型和接受抗PD-1/PDL1治疗的肝癌患者中高表达,MerTK是导致肝癌ICIs耐药的“元凶”之一。机制上,MerTK通过ERK/SP1通路上调SLC7A11的表达抑制肝癌细胞铁死亡,并通过募集髓源性抑制细胞(MDSCs)促进肝癌免疫抑制性肿瘤微环境(TME)的形成,从而使肝癌对抗PD-1/PD-L1产生耐药性。Sitravatinib是MerTK的一种抑制剂,通过促进肿瘤细胞铁死亡和减少MDSC向TME的浸润,使耐药HCC对抗PD-L1治疗增敏。肝细胞癌(HCC)仍然是一个全球性的健康挑战,其发病率在全球范围内不断增加。临床上,大多数HCC患者被诊断为晚期,治疗选择有限。最近,免疫疗法,如ICIs抗CTLA4、抗PD-1/PD-L1、它们的联合或与其他药物的联合,在癌症患者中取得了无与伦比的成功。此外,nivolumab、pembrolizumab和atezolizumab等ICIs目前已被FDA批准用于HCC的全身治疗。然而,由于抗PD-1/PD-L1治疗经常出现耐药,反应率仍远不能令人满意,目前仅在约15% - 20%的HCC患者中观察到总体获益率。因此,迫切需要确定最佳患者选择的生物标志物,并确定克服这种耐药性的有效靶点。MerTK是酪氨酸受体激酶TAM家族(TYRO3、AXL和MerTK)的一员,在包括HCC在内的多种恶性肿瘤中高度表达(the Human Protein Atlas: proteinatlas.org)。同源蛋白生长抑制特异性蛋白6 (Gas6)和蛋白S (Pros1)都是MerTK活化的天然配体,可诱导MerTK的同二聚化和自磷酸化,激活MEK/ERK、PI3K/AKT、JAK/STAT等下游通路,从而促进肿瘤细胞增殖、抗凋亡、抑制炎症和肿瘤恶性进程。最近的研究表明,MerTK作为一种代谢调节剂,通过整合有氧糖酵解和氧化磷酸化,在调节HCC生长中起着关键作用。此外,MerTK的受体/配体活性已在多种免疫细胞类型中得到证实,对癌症的免疫治疗具有重要意义。模式图(Credit: Cell Reports Medicine)铁死亡作为一种代谢调节的细胞死亡形式,在癌症生物学中起着重要作用。近年来,越来越多的证据表明,抑制铁死亡与传统癌症治疗的耐药性密切相关,靶向铁死亡可能为癌症治疗提供潜力。最近,研究表明,在免疫治疗过程中,调节铁死亡可以重塑肿瘤免疫微环境,从而导致T细胞介导的肿瘤消除。在HCC中,铁死亡已被证明与肿瘤治疗(如靶向治疗和化疗)耐药相关。该研究表明,MerTK上调SLC7A11表达抑制肿瘤细胞铁死亡,同时通过在HCC微环境中募集MDSCs促进肿瘤TME,从而导致抗PD-L1治疗耐药。靶向MerTK是提高PD-L1抗体治疗效果的有效策略。在PD-L1耐药肝癌中,MerTK抑制剂Sitravatinib和ICIs联合使用可有效增加肿瘤细胞铁死亡,减少肝癌微环境中MDSCs募集,促进CD8+ T细胞活化。该研究表明MerTK可作为ICB疗效预测标志物,并且是克服HCC中抗PD-1/PD-L1耐药的潜在靶点。原文链接https://doi-org.libproxy1.nus.edu.sg/10.1016/j.xcrm.2024.101415责编|探索君排版|探索君文章来源|“iNature”End往期精选围观一文读透细胞死亡(Cell Death) | 24年Cell重磅综述(长文收藏版)热文Nature | 破除传统:为何我们需要重新思考肿瘤的命名方式热文Nature | 2024年值得关注的七项技术热文Nature | 自身免疫性疾病能被治愈吗?科学家们终于看到了希望热文Nature | 癌症与神经系统的致命舞蹈
免疫疗法
2024-02-23
·今日头条
本文为转化医学网原创,转载请注明出处
作者:Rainbow
导读:
肝细胞癌(HCC)仍然是全球性健康挑战,其发病率在全球范围内不断增加。
2月20日,复旦大学董琼珠、钦伦秀、陆路共同通讯在期刊《Cell Reports Medicine》上在线发表题为“Disruption of MerTK increases the efficacy of checkpoint inhibitor by enhancing ferroptosis and immune response in hepatocellular carcinoma”的研究论文,
本研究表明MerTK上调SLC7A11表达以抑制肿瘤细胞的铁死亡,在HCC微环境中通过招募MDSCs来促进促肿瘤TME,并因此导致抗PD-L1治疗的耐药性。靶向MerTK是增加PD-L1抗体治疗疗效的有效策略。MerTK抑制剂Sitravatinib与ICIs的联合可以有效增加PD-L1耐药TME中的肿瘤细胞铁死亡,减少MDSC募集,并促进CD8+T细胞的活化。
https://www-cell-com.libproxy1.nus.edu.sg/cell-reports-medicine/fulltext/S2666-3791(24)00038-7
研究背景
01
肝细胞癌(HCC)仍然是全球性健康挑战,其发病率在全球范围内不断增加。临床上,大多数HCC患者被诊断为晚期病症,治疗选择有限。最近,免疫治疗,如免疫检查点抑制剂(ICIs)抗CTLA4、抗PD-1/PD-L1及其联合使用,或与其他药物联合使用,在治疗癌症患者方面取得了巨大的成功。此外,nivolumab, pembrolizumab和atezolizumab等ICIs目前已获得美国食品药品监督管理局(FDA)批准,用于HCC的全身治疗。然而,由于抗PD-1/PD-L1治疗常见的耐药性,患者的反应率仍然远不能令人满意,目前大约只有15%–20%的HCC患者能够获得整体收益率。
因此,迫切需要确定最佳患者选择的生物标志物,并确定有效的靶点以克服这种耐药性。
MER原癌基因酪氨酸激酶(MerTK)是TAM家族(TYRO3、AXL和MerTK)受体酪氨酸激酶的成员,高度表达于包括HCC在内的多种恶性肿瘤中。
同源蛋白生长抑制特异性蛋白6(Gas6)和蛋白S(Pros1)都是MerTK激活的天然配体,它们诱导MerTK的同源二聚化和自磷酸化,并激活下游途径,如MEK/ERK、PI3K/AKT和JAK/STAT,促进肿瘤细胞增殖、抗凋亡、抑制炎症和肿瘤生长。最近的研究表明,作为代谢调节因子,MerTK在调控HCC生长中发挥关键作用,通过整合有氧糖酵解和氧化磷酸化。此外,MerTK的受体/配体活性已在多种免疫细胞类型中得到证实,并增强抗肿瘤免疫,这对癌症免疫治疗具有重要意义。
作为一种代谢调节的细胞死亡形式,铁死亡(Ferroptosis)在癌症生物学中扮演着重要角色。近年来,越来越多证据表明,抑制铁死亡与传统癌症治疗的耐药性相关,而靶向铁死亡可能为癌症治疗提供潜在机会。最近的研究表明,铁死亡的调控可以重塑肿瘤微环境,在免疫治疗期间导致T细胞介导的肿瘤根除。
在肝细胞癌(HCC)中,铁死亡已被证实在靶向治疗和化疗等系统疗法中赋予药物耐药性。
研究发现
02
在这项研究中,研究人员证明了具有高 MER 原癌基因酪氨酸激酶 (MerTK) 表达的 HCC 在两种同基因小鼠模型和接受抗 PD-1/PD-L1 治疗的患者中表现出抗 PD-1/PD-L1 耐药性。从机制上讲,MerTK 通过 ERK/SP1 通路上调 SLC7A11 来限制铁死亡,并通过募集髓源性抑制细胞 (MDSC) 促进免疫抑制性肿瘤微环境(TME)的发展,使 HCC 对抗 PD-1/PD-L1 产生抗性。Sitravatinib 是一种 MerTK 抑制剂,通过促进肿瘤铁死亡和减少 MDSC 浸润到TME中,使耐药HCC对抗 PD-L1 治疗敏感。
研究发现 MerTK 可以作为患者分层的预测性生物标志物,并作为克服HCC抗PD-1/PD-L1 耐药性的有希望的靶点。
研究结果
03
综上所述,
本研究表明MerTK上调SLC7A11表达以抑制肿瘤细胞的铁死亡,在HCC微环境中通过招募MDSCs来促进促肿瘤TME,并因此导致抗PD-L1治疗的耐药性。靶向MerTK是增加PD-L1抗体治疗疗效的有效策略。MerTK抑制剂Sitravatinib与ICIs的联合可以有效增加PD-L1耐药TME中的肿瘤细胞铁死亡,减少MDSC募集,并促进CD8+T细胞的活化。
参考资料:
https://www-cell-com.libproxy1.nus.edu.sg/cell-reports-medicine/fulltext/S2666-3791(24)00038-7
注:本文旨在介绍医学研究进展,不能作为治疗方案参考。如需获得健康指导,请至正规医院就诊。
热门·直播/活动
🕓 上海|3月01日-02日
▶第三届长三角单细胞组学技术应用论坛暨空间组学前沿论坛将在上海召开,诚邀您的参与!
点击对应文字 查看详情
免疫疗法
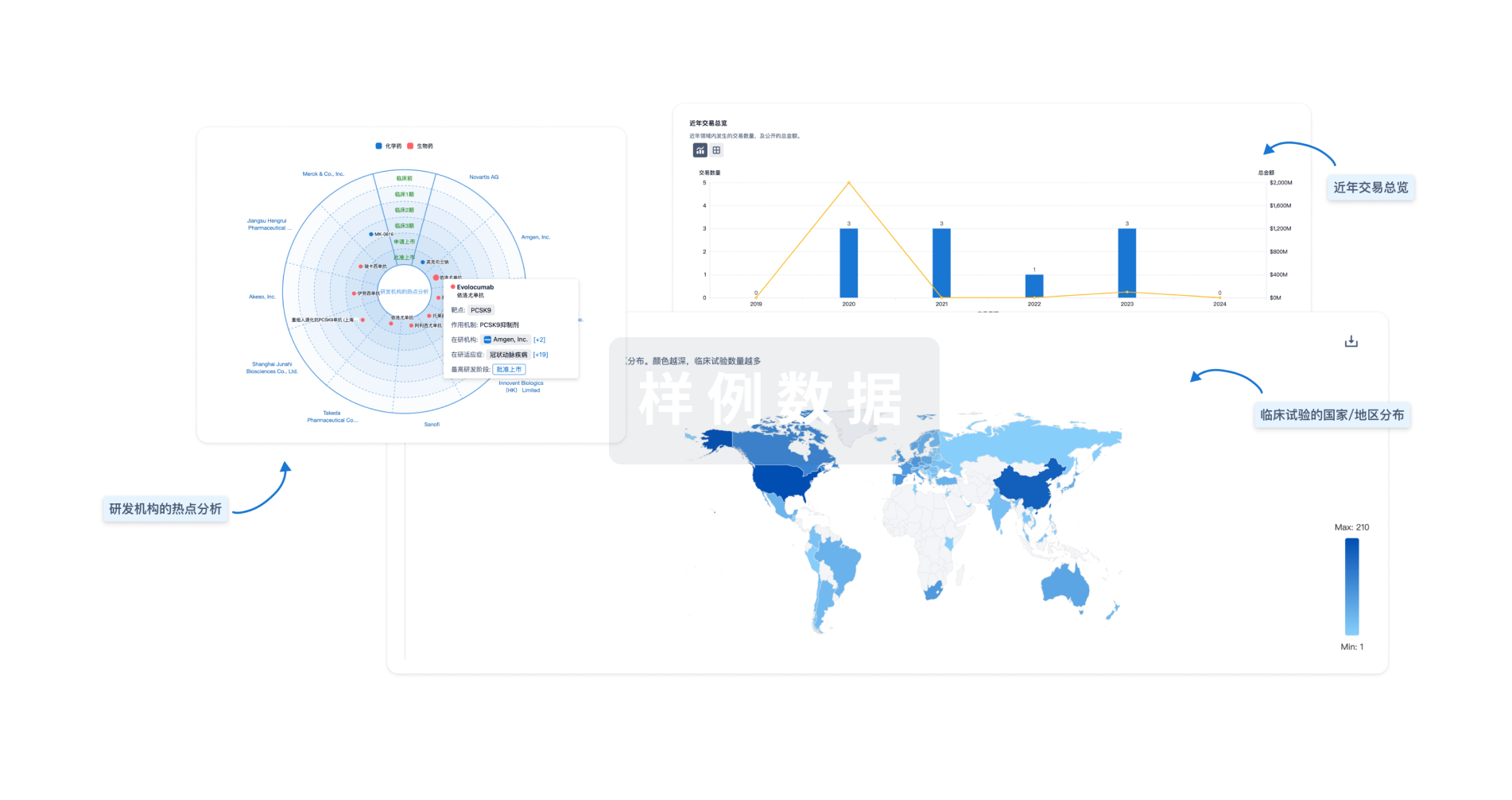
分析
对领域进行一次全面的分析。
登录
或
来和芽仔聊天吧
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
