更新于:2024-09-19

BioLeap, Inc.
更新于:2024-09-19
概览
关联
1
项与 BioLeap, Inc. 相关的药物作用机制 LDLR拮抗剂 [+1] |
在研机构- |
原研机构 |
在研适应症- |
最高研发阶段无进展 |
首次获批国家/地区- |
首次获批日期- |
100 项与 BioLeap, Inc. 相关的临床结果
登录后查看更多信息
0 项与 BioLeap, Inc. 相关的专利(医药)
登录后查看更多信息
3
项与 BioLeap, Inc. 相关的文献(医药)2017-08-01·Bioorganic & Medicinal Chemistry3区 · 医学
Design, synthesis and biological evaluation of renin inhibitors guided by simulated annealing of chemical potential simulations
3区 · 医学
Article
作者: Kulp, John L ; Smith, Emilie D ; Dickson, John K ; Barta, Thomas E ; Cloudsdale, Ian S ; Guarnieri, Frank ; Grella, Brian S
2011-07-20·Journal of the American Chemical Society1区 · 化学
Diverse Fragment Clustering and Water Exclusion Identify Protein Hot Spots
1区 · 化学
Article
作者: Pompliano, David L. ; Kulp, John L. ; Guarnieri, Frank
Contribution to affinity from quantum resonance: A fragment-based approach to RecA and PCSK9
作者: Kulp, John L. III ; Guarnieri, Frank K. ; Kulp, John L. Jr.
100 项与 BioLeap, Inc. 相关的药物交易
登录后查看更多信息
100 项与 BioLeap, Inc. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2024年10月06日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
其他
1
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
PCSK9/LDLR inhibitors (BioLeap) ( LDLR x PCSK9 ) | 高胆固醇血症 更多 | 无进展 |
登录后查看更多信息
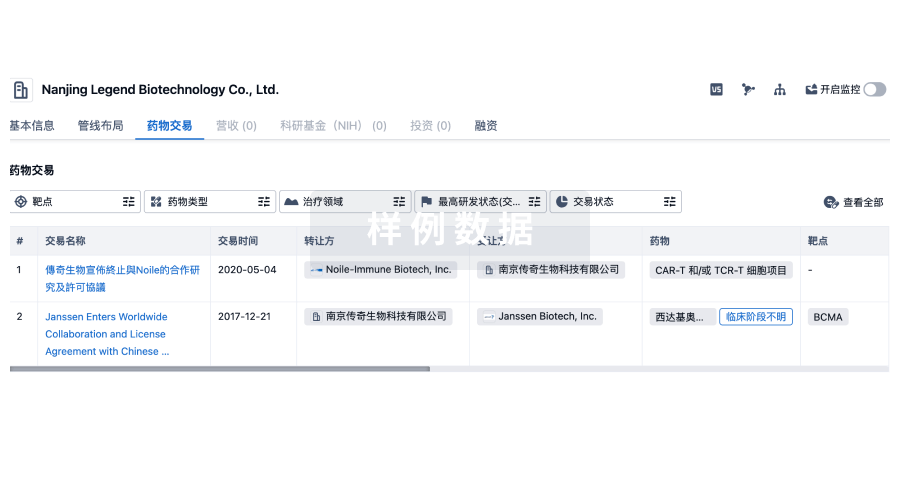
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
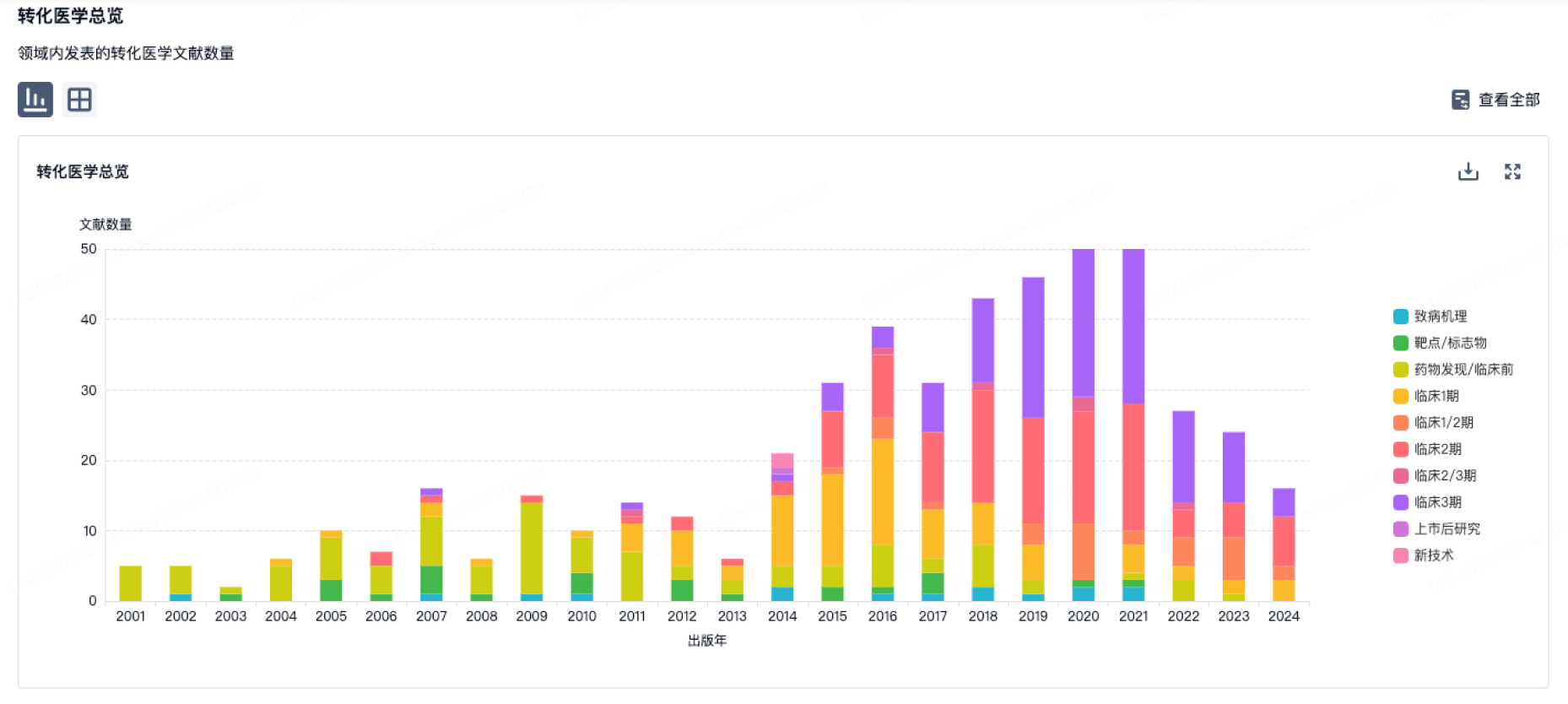
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
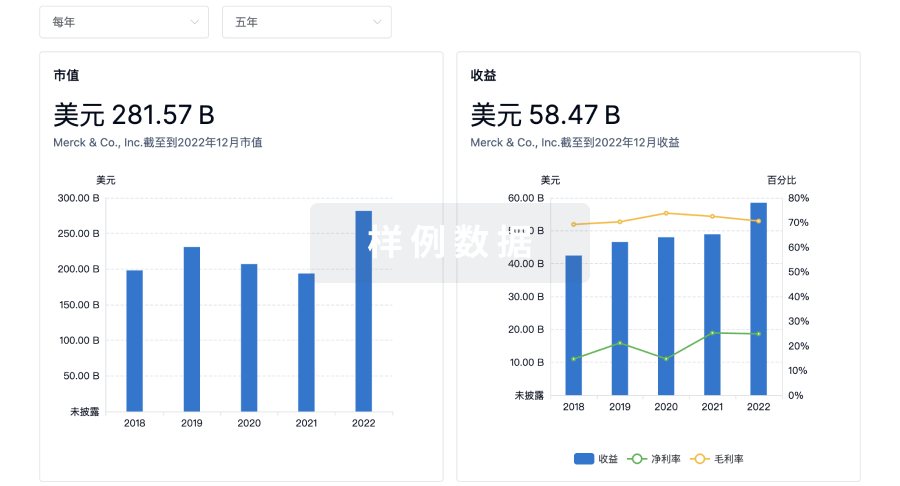
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
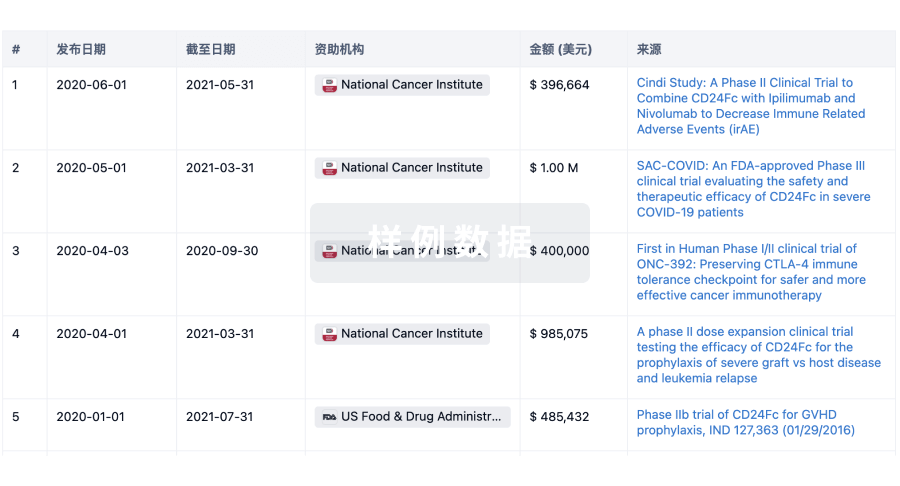
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
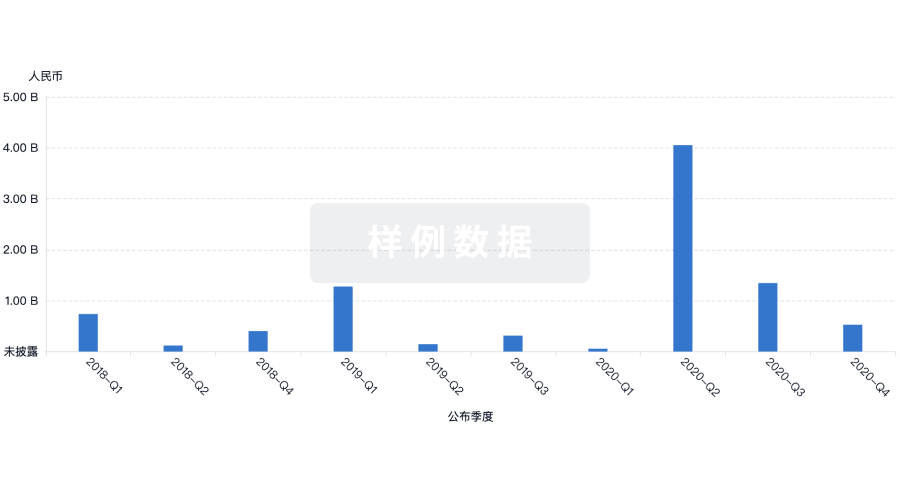
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
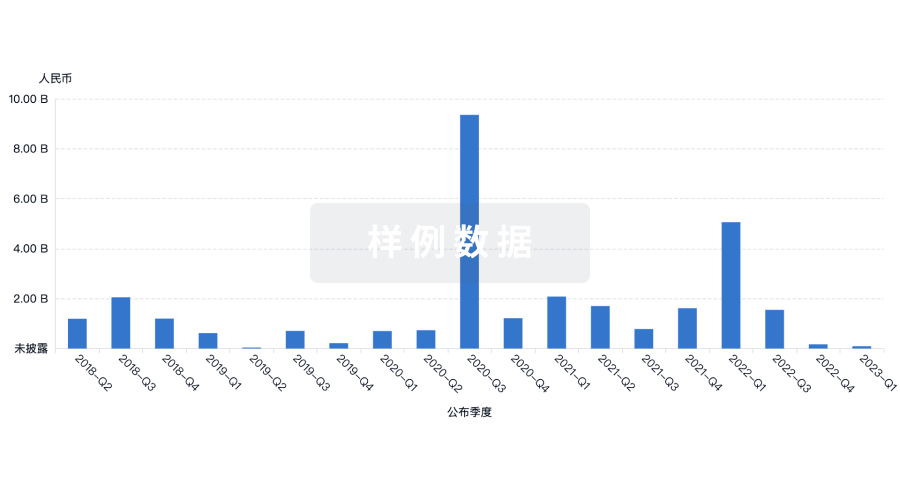
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
标准版
¥16800
元/账号/年
新药情报库 | 省钱又好用!
立即使用
来和芽仔聊天吧
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用