预约演示
更新于:2025-05-07
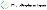
MicroBiopharm Japan Co., Ltd.
更新于:2025-05-07
概览
标签
泌尿生殖系统疾病
皮肤和肌肉骨骼疾病
肿瘤
小分子化药
关联
2
项与 MicroBiopharm Japan Co., Ltd. 相关的药物作用机制 DNA抑制剂 [+1] |
非在研适应症- |
最高研发阶段批准上市 |
首次获批国家/地区 日本 |
首次获批日期1988-03-29 |
作用机制 DNA抑制剂 [+1] |
非在研适应症- |
最高研发阶段批准上市 |
首次获批国家/地区 日本 |
首次获批日期1981-06-04 |
100 项与 MicroBiopharm Japan Co., Ltd. 相关的临床结果
登录后查看更多信息
0 项与 MicroBiopharm Japan Co., Ltd. 相关的专利(医药)
登录后查看更多信息
11
项与 MicroBiopharm Japan Co., Ltd. 相关的文献(医药)2023-01-01·Bioorganic & Medicinal Chemistry Letters
ez-ADiCon: A novel glyco-remodeling based strategy that enables preparation of homogenous antibody-drug conjugates via one-step enzymatic transglycosylation with payload-preloaded bi-antennary glycan complexes
Article
作者: Kanamori, Satoshi ; Chon, Hyongi ; Narumi, Hideki ; Nagahara, Takashi ; Ohara, Keiichiro ; Suzuki, Tomohiko ; Hibino, Kazuhiro
2021-12-01·Parasitology International
Exploring natural microbial resources for the discovery of anti-malarial compounds
Article
作者: Yamashita, Michio ; Nozaki, Tomoyoshi ; Matsumoto, Atsuko ; Wibowo, Agung Eru ; Siska, Eka ; Puspitasari, Dian Japany ; Watanabe, Azuma ; Sakura, Takaya ; Hidayati, Dyah Noor ; Kurnia, Kesi ; Dewi, Diana ; Oktaviani, Avi Nurul ; Inaoka, Daniel Ken ; Waluyo, Danang ; Nonaka, Kenichi ; Kristiningrum ; Mahsunah, Anis Herliyati ; Rahmawati, Yulia ; Putri, Tiara Zovi ; Dobashi, Kazuyuki ; Miyazaki, Yukiko ; Pramisandi, Amila ; Nugroho, Nuki Bambang ; Hartuti, Endah Dwi ; Bernawati, Putri ; Adipratiwi, Nadia ; Mori, Mihoko ; Chrisnayanti, Evita ; Tokiwa, Toshiyuki ; Melinda ; Nurkanto, Arif ; Afrianti, Kiki Rizkia ; Ariyani, Titin ; Nurlaila ; Suryani ; Prabandari, Erwahyuni Endang ; Shiomi, Kazuro
2020-12-01·Bioscience, Biotechnology, and Biochemistry4区 · 工程技术
Crystal structure of a glycoside hydrolase family 68 β-fructosyltransferase from Beijerinckia indica subsp. indica in complex with fructose
4区 · 工程技术
Article
作者: Fujii, Tadashi ; Hirano, Katsuaki ; Nagaya, Mika ; Tamura, Keisuke ; Tonozuka, Takashi ; Kawai, Reika ; Tochio, Takumi ; Nishikawa, Atsushi ; Kitamura, Junichi
100 项与 MicroBiopharm Japan Co., Ltd. 相关的药物交易
登录后查看更多信息
100 项与 MicroBiopharm Japan Co., Ltd. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2025年07月18日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
批准上市
2
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
盐酸阿柔比星 ( DNA x Top II ) | 肺癌 更多 | 批准上市 |
盐酸吡柔比星 ( DNA x Top II ) | 子宫癌 更多 | 批准上市 |
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
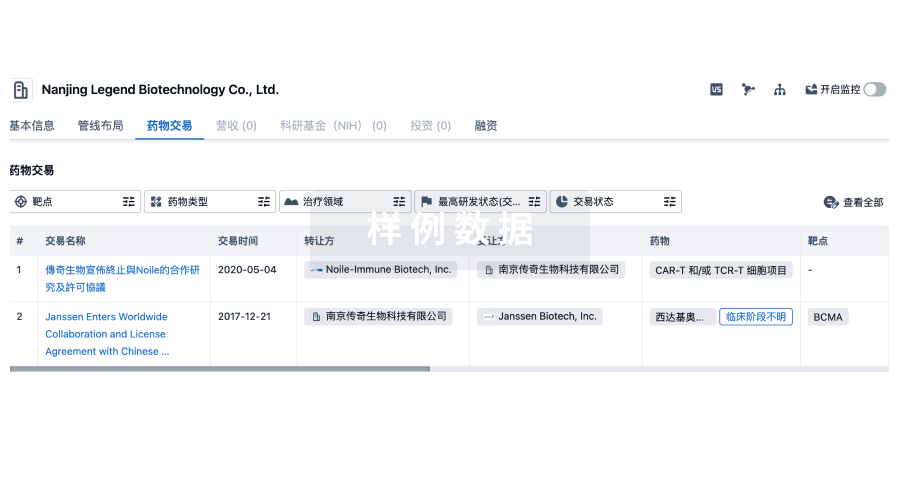
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
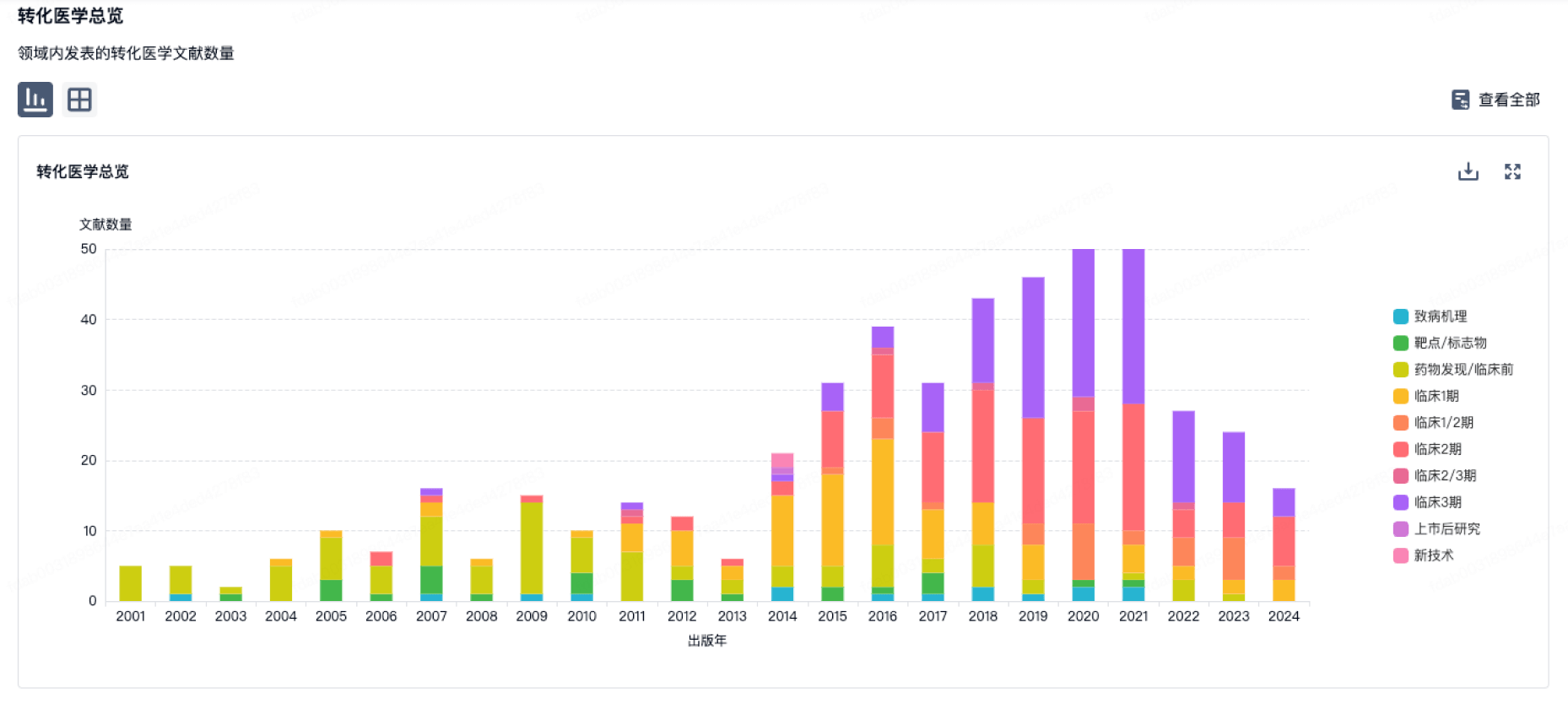
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
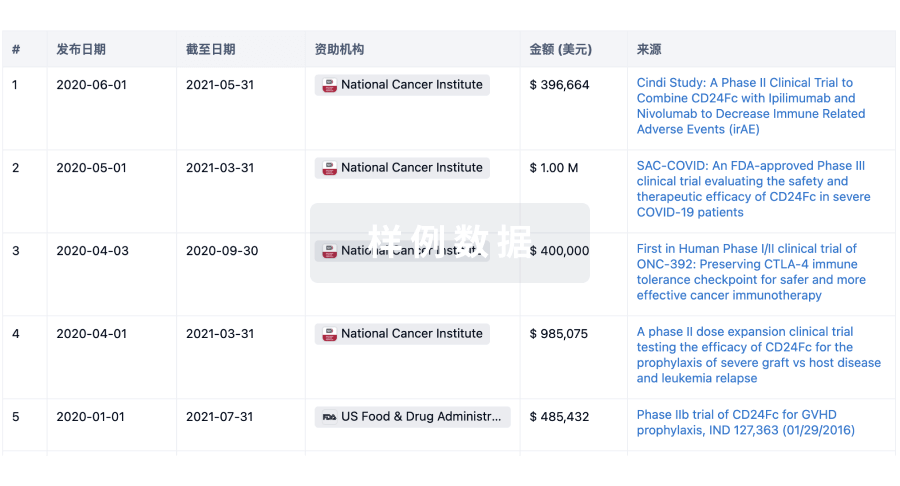
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
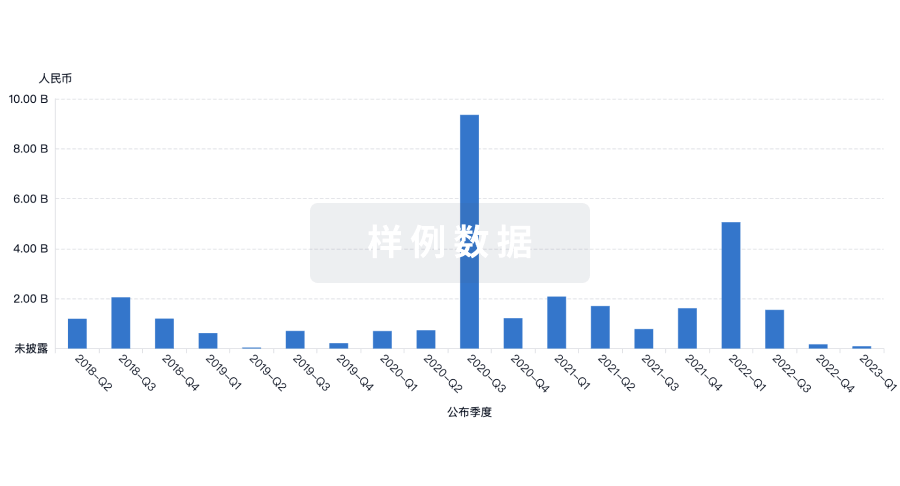
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
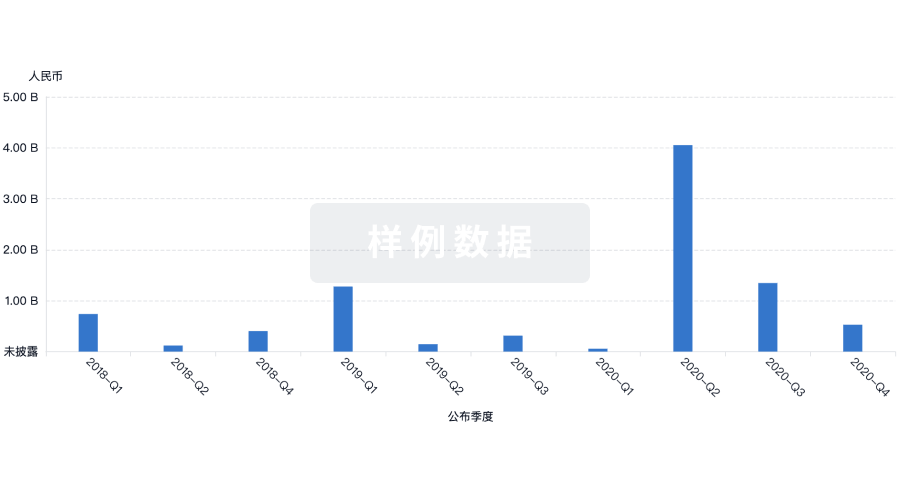
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用

