更新于:2024-09-19

Harmonic Discovery, Inc.
更新于:2024-09-19
概览
标签
血液及淋巴系统疾病
肿瘤
小分子化药
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 小分子化药 | 1 |
| 排名前五的靶点 | 数量 |
|---|---|
| FLT3(酪氨酸蛋白激酶受体FLT3) | 1 |
关联
1
项与 Harmonic Discovery, Inc. 相关的药物靶点 |
作用机制 FLT3调节剂 |
在研机构 |
原研机构 |
在研适应症 |
非在研适应症- |
最高研发阶段临床前 |
首次获批国家/地区- |
首次获批日期- |
100 项与 Harmonic Discovery, Inc. 相关的临床结果
登录后查看更多信息
0 项与 Harmonic Discovery, Inc. 相关的专利(医药)
登录后查看更多信息
4
项与 Harmonic Discovery, Inc. 相关的文献(医药)2024-02-01·Current Opinion in Structural Biology
AI for targeted polypharmacology: The next frontier in drug discovery
Review
作者: Cichońska, Anna ; Rahman, Rayees ; Ravikumar, Balaguru
2023-01-01·Bioinformatics advances
Investigating the conformational landscape of AlphaFold2-predicted protein kinase structures.
Article
作者: Trozzi, Francesco ; Ravikumar, Balaguru ; Rahman, Rayees ; Al-Masri, Carmen ; Patek, Marcel ; Cichonska, Anna ; Tran, Oanh ; Lin, Shu-Hang ; Sahni, Navriti
DockDB: A database management system for large-scale docking studies
作者: Patek, Marcel ; Sahni, Navriti ; Ravikumar, Balaguru ; Rahman, Rayees ; Cichonska, Anna ; Trozzi, Francesco
3
项与 Harmonic Discovery, Inc. 相关的新闻(医药)2024-04-01
— Companies will co-develop inhibitors of a dark kinase target (NEK1) identified for cancer dependency by Turbine's Simulated Cell™ platform —
LONDON and BUDAPEST, Hungary and NEW YORK, April 1, 2024 /PRNewswire/ -- Turbine, a company that is building the world's first predictive simulation of patient biology, and Harmonic Discovery (Harmonic), a therapeutics company building an integrated computational and experimental platform for kinase drug discovery and targeted polypharmacology, have entered into a collaboration to co-develop novel cancer therapies that will inhibit a dark kinase identified for cancer dependency using Turbine's Simulated Cell™ platform.
"We believe that the dark kinase NEK1 holds great potential as a drug target for developing cancer therapies addressing important resistance challenges in the clinic," said Daniel Veres, M.D., Ph.D., Chief Scientific Officer and Co-Founder of Turbine. "Coupling Harmonic's deep expertise in multi-specific kinase-targeted drug design with our unique ability to guide drug discovery by running simulated experiments with causal interpretation in a vast biological search space will help realize a more innovative and efficient process than traditional approaches. We are excited for the opportunity to demonstrate the power of our platforms to collectively boost R&D success in the clinic."
"Our fully integrated kinase drug discovery platform aims to challenge current drug discovery paradigms by building a machine learning-first infrastructure that integrates multiple aspects of kinase drug discovery," said Rayees Rahman, Ph.D., CEO and Co-Founder of Harmonic Discovery. "Turbine's exciting biological simulation technology is highly complementary to our approach. We are confident that integrating both companies' innovative platforms will yield a radically more effective approach to selecting and advancing kinase-targeted therapies that can substantially improve patient outcomes in oncology."
The collaboration provides a framework in which biological and mechanistic insights from Turbine's simulations will directly inform Harmonic's generative chemistry models – with the goal of bringing forward new, differentiated, and effective kinase inhibitor therapies for the patients who need them most. Harmonic will assume responsibilities for computational chemistry and medicinal chemistry activities, while Turbine will be responsible for in silico simulations and wet-lab validation of target biology, including the identification of both synergistic kinase co-targets to NEK1 and patient populations most likely to benefit from therapies. Financial terms of the collaboration are not disclosed.
About Turbine
Countless resources and time get expended globally to research and advance novel therapies that end in clinical failure and no patient benefit. Imagine a world where it is possible to predict any potential drug's effect on translatable biological models – including those that may not even be available for lab-based testing – while accurately representing patient biology. Turbine is revolutionizing the status quo by working to ensure that we always run the right wet lab experiments.
With machine learning that understands the logic by which human cells make decisions, Turbine has built the world's first predictive simulation of patient biology. The Simulated Cell™ platform models the protein signaling that decides cell fate and facilitates in silico experiments at scales that are impossible in the physical world, to empower the biopharma industry by guiding experiments that identify and confirm disease driving mechanisms.
Turbine's platform has been validated with leading pharma companies such as AstraZeneca, Ono Pharmaceutical, Cancer Research Horizons, and Bayer. In addition, Turbine has received funding from industry leaders such as MSD (Merck & Co, Inc. Rahway NJ, USA) Global Health Innovation Fund, Accel, and MassMutual Ventures, among others.
For more information, visit or follow Turbine on LinkedIn.
About Harmonic Discovery
Harmonic Discovery (the "Company") is a precision pharmacology company applying its generative chemistry platform to advance next-generation kinase inhibitors. The Company's platform is trained on multiple layers of proprietary data and is integrated with domain expertise to design safer and more efficacious kinase therapeutics.
The Company's lead programs are focused on malignant hematology indications, with exploratory discovery efforts in additional therapeutic areas including immuno-oncology, immunology and inflammation. The Company has raised $8 million in Seed Financing to date from Innovation Endeavors, Fifty Years, Y Combinator and other select investment firms. The Company was founded in 2021 and is headquartered at Johnson & Johnson's JLABS ("JLABS@NYC") in New York City, New York.
To learn more, visit
SOURCE Harmonic Discovery Inc.
2024-01-30
Compound has demonstrated in vivo results indicating potentially best-in-class profile in FLT3 mutated AML
Pre-clinical development activities are underway, with IND enabling studies expected to commence in late 2024
NEW YORK, Jan. 30, 2024 /PRNewswire/ -- Harmonic Discovery (the "Company"), a precision pharmacology company focusing on developing next generation kinase inhibitors, today announced an exclusive licensing agreement of a novel small-molecule drug candidate for the treatment of FLT3 mutated acute myeloid leukemia (AML) from BioVentures (University of Arkansas for Medical Sciences) and UC San Francisco (UCSF). The compound, named HD-10019, is currently in pre-clinical development and is expected to commence IND-enabling studies in late 2024.
HD-10019 was jointly developed in the laboratories of professors Hong-Yu Li, PhD, previously at the University of Arkansas for Medical Sciences, and Neil Shah, MD, PhD, of UCSF. HD-10019 is a type II kinase inhibitor that was rationally designed to overcome tyrosine kinase domain resistance mutations in the FLT3 kinase and improve myelosuppression versus standard of care agents for FLT3 mutated AML. Additionally, this molecule is anticipated to avoid cardiac liabilities seen in other kinase inhibitors.
"Since we provided the first evidence that FLT3-ITD is a valid therapeutic target in AML in 2012, three FLT3 tyrosine kinase inhibitors have been approved in the USA and throughout much of the world. While these agents represent advances in the management of this poor-risk subset of AML patients, their impact on clinical outcomes has been modest as a consequence of their vulnerability to secondary drug-resistant kinase domain mutations in FLT3-ITD and their insufficient selectivity toward FLT3, which is hypothesized to be a prime reason for the myelosuppression observed with these agents," said Dr. Neil Shah, whose laboratory performed pre-clinical studies of the compound. "To this end, we have identified and characterized HD-10019. The clinical development of such an agent may facilitate the ability to achieve effective and durable FLT3 inhibition when used in conjunction with chemotherapy and as maintenance therapy thereafter."
Harmonic Discovery's precision pharmacology platform has further corroborated the potential of HD-10019 as a best-in-class drug based on its selectivity profile. The Company intends to leverage its platform to further advance the development of this program. "We are excited to work with BioVentures and UCSF and take a step forward in delivering a novel treatment to FLT3 mutated AML patients," said Rayees Rahman, PhD, CEO of Harmonic Discovery.
Wilson Sonsini Goodrich & Rosati served as legal advisor to Harmonic Discovery on matters relating to the Licensing Agreement and related corporate and patent strategy matters.
About Harmonic Discovery Inc.
Harmonic Discovery (the "Company") is a precision pharmacology company applying its generative AI platform to advance next-generation kinase inhibitors. The Company's platform is trained on multiple layers of proprietary data and is integrated with domain expertise to design safer and more efficacious kinase therapeutics.
The Company's lead programs are focused on malignant hematology indications, with exploratory discovery efforts in additional therapeutic areas including immuno-oncology, immunology and inflammation. The Company has raised $8 million in Seed Financing to date from Innovation Endeavors, Fifty Years, Y Combinator and other select investment firms. The Company was founded in 2021 and is headquartered at Johnson & Johnson's JLABS ("JLABS@NYC") in New York City, New York.
To learn more, visit
CONTACT: pr@harmonicdiscovery.com
SOURCE Harmonic Discovery Inc.
引进/卖出
2022-10-19
·创鉴汇
▎药明康德内容团队编辑医药行业有一条著名的“双十定律”,即需要超过10年时间、10亿美元的成本,才有可能成功研发出一款新药。即使如此,大约只有10%的新药能被批准进入临床期。由此可见新药研发工作的周期之长、风险之大。但现如今,AI制药为新药研发创造了一种新的可能。窥其本质,以数据为基础的AI制药是通过机器自主学习数据、收集归纳数据,总结出不拘泥于已有经验的药物研发规律,进而优化从靶点发现到临床试验的药物研发各个环节。基于现有的成果,AI制药展现出了大幅提升药物研发效率与成功率的潜力,让医药行业看到了打破“双十定律”的曙光。过去1个月,AI制药赛道融资事件接连不断,多家公司的AI技术获资本青睐。其中,比尔•盖茨寄予厚望的Nimbus Therapeutics再获1.25亿美元助力,运用基于机器学习的预测建模方法挑战不可成药靶点,研发小分子创新药;TCR细胞疗法新锐ImmunoScape受安进(Amgen)看好,其平台可识别出现频率仅有0.001%的抗原特异性CD8阳性T细胞;今年2月已融资5000万元的百葵锐生物又完成了数千万元融资,打造AI合成生物学平台,设计开发高效“细胞工厂”,提升合成效率……以下还有更多值得一看的融资资讯。01# AI应用于小分子药物研发 # 比尔•盖茨持续看好!Nimbus再获1.25亿美元助力,运用基于机器学习的预测建模方法研发小分子创新药9月12日,Nimbus Therapeutics宣布完成1.25亿美元融资,参与本轮融资的包括新投资机构Bain Capital Life Sciences和SV Health Investors,其他老投资者如比尔·盖茨、Pfizer Ventures、RA Capital Management、Atlas Venture等也参与其中。Nimbus Therapeutics成立于2009年,是一家基于结构的药物发现和突破性药物设计开发公司,专注于研发新型小分子药物,旨在挑战与多种人类疾病有关、经过充分验证但不可成药的靶点。该公司结合了前沿计算技术和基于机器学习的预测建模方法,搭建基于蛋白质结构的药物发现技术平台,设计靶向蛋白质的强效、选择性小分子创新药物。该平台能够更有效地将高分辨率的蛋白质结构数据用于靶向潜在靶点,并调节其功能活性。本轮融资所获资金将用于支持该公司在研TYK2抑制剂NDI-034858在中重度斑块状银屑病、活动性银屑病关节炎中的2b期临床试验,以及治疗银屑病的3期临床试验。值得一提的是,这不是Nimbus Therapeutics第一次获得资本青睐。早在2011年,成立仅3年的Nimbus就获得了Atlas Venture、SR One和礼来风险投资共同牵头的A轮融资,彼时比尔•盖茨也参与其中。了解更多Nimbus信息,请点击>>Harmonic Discovery完成800万美元种子轮融资,搭建AI激酶药物研发平台 9月8日,激酶药物研发公司Harmonic Discovery宣布完成800万美元种子轮融资,本轮融资由Innovation Endeavors领投,Y Combinator等共同参与,所获资金将用于推进激酶药物的研发。该公司正在搭建一个以机器学习为重点的激酶药物发现平台,该平台所基于的AI模型在组成激酶药物的原子和键的拓扑结构、激酶结构的三维内外、激酶驱动的疾病动态细胞环境中进行学习,并整合了系统生物学、化学信息学、药物化学等多学科知识,旨在针对癌症和自身免疫性疾病等开发新型疗法。了解更多Harmonic信息,请点击>>02# AI应用于大分子药物研发 # 发现0.001%抗原特异性CD8 T细胞!TCR细胞疗法新锐完成1400万美元融资9月20日,ImmunoScape宣布完成1400万美元融资,所获资金将用于加速疗法开发,以及推动候选疗法进入临床。值得注意的是,安进亦参与这次的融资。ImmunoScape专注于开发下一代T细胞受体(TCR)细胞疗法,其深度免疫组(Deep Immunomics)平台能够以极高的灵敏度对病患的样本进行大型的T细胞探勘。通过结合专有的条形码科技,能够识别那些与治疗相关的T细胞以及其所表达的TCR。根据公司官方网站,该平台可以经常识别出现频率仅有0.001%的抗原特异性CD8阳性T细胞。此外, ImmunoScape专有的TargetScape靶标筛选平台,利用多肽-主要组织相容性复合体(MHC)编码系统与大型抗体组进行高通量筛选,以人工智能方式识别出数百个可能具有治疗益处的候选T细胞。这些筛选出具抗原特异性的候选T细胞会再经过单细胞测序,对其TCR序列、表型、基因表达量进行分析。所识别的创新TCR靶标会通过进一步的开发,在体外与体内模型中,检视其被运用在癌症治疗的疗效、安全性与特异性。了解更多ImmunoScape信息,请点击>>开源机器学习平台驱动大分子药物研发,百奥几何完成千万美元天使轮融资 9月21日,百奥几何宣布完成千万美元天使轮融资,本轮投资方为高榕资本。目前,百奥几何团队联合英伟达(NVIDIA)、英特尔(intel)、IBM等公司联合发布了针对大分子药物研发的开源机器学习平台TorchProtein,致力于通过AI加速药物研发的进程。该平台开源了深度学习对大分子建模的一个通用框架、基于蛋白质三维几何结构的一个预训练大模型,以及专门用于评价深度学习对蛋白质建模效果的标准数据集,其开创性算法能够帮助研究人员更好地提取蛋白质特征。在进行大量的预训练后,该算法后期将显著降低对有标签的数据的依赖,进一步降低研发成本。据悉,目前百奥几何已经完成了对人工智能大分子药物设计平台的建设,并且,为了完成干湿实验闭环,百奥几何也在逐步打造高通量大分子药物湿实验验证平台。了解更多百奥几何信息,请点击>>设计开发高效“细胞工厂”,数千万元助力百葵锐生物搭建AI合成生物学平台9月7日,合成生物学企业百葵锐生物宣布完成数千万元Pre-A+轮融资,本轮融资由天津万联道一资本和星陀资本共同参与。2月,百葵锐生物已完成5000万元Pre-A轮融资。百葵锐成立于2019年,围绕蛋白精准设计和蛋白分子机器技术建立全态链合成生物学平台,专注于多领域的高效生物合成。该公司打造蛋白精准设计与蛋白分子机器技术平台,结合生物大数据机器学习和理性设计,开发功能强大的蛋白元件。并以此为基础,对蛋白分子进行定向进化,进一步提高蛋白性能。该公司运用智能自动化高通量筛选系统,结合高效基因编辑技术,构建具有高度协同效应的蛋白机器,从而能够快速打通产品在细胞中的代谢合成通路,设计开发出高效“细胞工厂”,提升合成效率。了解更多百葵锐生物信息,请点击>>开发AI算法基因治疗模拟平台,WhiteLab Genomics完成1000万美元A轮融资9月13日,WhiteLab Genomics宣布完成1000万美元A轮融资,本轮融资由Debiopharm Innovation Fund与Omnes Capital共同参与,所获资金将助力该公司运用AI算法加速基因组疗法研发。WhiteLab Genomics是一家基因疗法、细胞疗法研发公司,旨在利用数据及AI算法开发用于基因治疗的模拟平台,加快创新药物研发进程,彻底改变细胞基因疗法的发展。该公司的技术平台有助于快速开发目标载体和有效载荷,节省患者获得新基因组疗法所需的宝贵时间。目前,其解决方案已被运用于包括RNA、DNA和细胞疗法在内的新药开发计划之中。了解更多WhiteLab信息,请点击>>03# AI应用于临床前研究 # 疾病生物学AI模型化,数千万元助力爱信智耀探索新靶点研发创新药 9月19日,爱信智耀(Ailomics Therapeutics)宣布完成数千万元天使轮融资,由启明创投领投,三一创新投资和华桐资本跟投,所获资金将用于该公司的核心技术平台搭建以及研发管线推进。基于对疾病生物学的深度理解,爱信智耀致力于针对具有重大临床需求的免疫性疾病和肿瘤进行创新药物研发。随着此次融资的完成,该公司将搭建新一代计算生物、实验生物和抗体研发的综合技术平台。通过深度学习等AI技术系统地分析病人的多组学大数据(包括单细胞和空间组学),发现导致疾病进展的关键细胞类型和细胞内外信号通路,选择有潜力解决重大临床需求的新靶点用于研发潜在“first-in-class”药物。爱信智耀创始人兼CEO张清博士表示,疾病生物学错综复杂,爱信智耀将利用快速发展的测序技术和数据科学,结合中国大量且高度集中的临床样本,对疾病生物学进行数字化和AI模型化。为了更加提高新药的临床转化成功率,公司将把新药科研“人源化”,建立包括人体器官芯片在内的人源疾病模型平台。了解更多爱信智耀信息,请点击>>AI病理学模型加速临床前及转化研究,Aignostics完成1400万欧元A轮融资9月16日,Aignostics宣布完成超额认购1400万欧元A轮融资,本轮融资由Wellington Partners领投,勃林格殷格翰风险基金(BIVF)、High-Tech Gründerfonds(HTGF)等跟投,所获资金将用于推进该公司的AI病理学模型开发。Aignostics深耕AI于病理学领域的应用,能够结合专有技术构建定制的AI模型。其AI模型不同于当前的已有的解决方案,它涵盖了苏木精-伊红染色(H&E)、免疫组织化学(IHC)和多重免疫荧光(mIF)在内的关键组织染色技术,能够提高对疾病生物学、新型生物标志物和药物反应特征的理解,从而加速临床前和转化研究。Aignostics专有的“可解释AI”(Explainable AI)技术能够揭示AI预测和决策所基于的特征,改革了传统的AI黑盒模型。此外,Aignostics还开发了一种独特的算法,能够不依赖于病理学家的手动注释,自动训练AI。了解更多Aignostics信息,请点击>>利用机器学习匹配分子与潜在适应症,Arpeggio完成1700万美元A轮融资9月7日,生物技术公司Arpeggio Biosciences宣布完成1700万美元A轮融资,本轮融资由Builders VC领投,腾讯等跟投,所得资金将支持该公司药物管线以及其转录监测技术的持续开发。Arpeggio的技术可直接读出转录,并可以监测超过10万个剪切体在患病和健康细胞中的活性,这些细胞可能暴露于小分子、反义寡核苷酸或生物制品等多样化的刺激之下。利用机器学习,Arpeggio重建了与分子相关的生物网络,并将分子与其有望治疗的疾病进行匹配。了解更多Arpeggio信息,请点击>>临床数据提取算法加速试验患者匹配,Dyania Health完成530万美元种子轮融资 9月19日,Dyania Health宣布完成530万美元种子轮融资,本轮融资由Innospark Ventures领投,所获资金将用于加快该公司临床数据算法研发等。Dyania Health成立于2019年,其创始人致力于解决临床试验中一个重要但未解决的问题,即根据病历人工预筛选患者参与临床研究既繁琐又耗时。就大多数临床试验而言,根据严苛的临床试验标准匹配患者的过程通常需要18至24个月的时间。Dyania Health的临床数据提取算法能够从电子病历(EMR)中识别非结构化和结构化患者数据,并推断患者是否符合临床试验准入标准,从而实现自动化病历预筛选,提高临床试验患者匹配效率。了解更多Dyania信息,请点击>>了解更多全球AI制药领域新动态请点击下图访问我们的小程序免责声明:药明康德内容团队专注介绍全球生物医药健康研究进展。本文仅作信息交流之目的,文中观点不代表药明康德立场,亦不代表药明康德支持或反对文中观点。本文也不是治疗方案推荐。如需获得治疗方案指导,请前往正规医院就诊。版权说明:本文由药明康德内容团队根据公开资料整理编辑,欢迎个人转发至朋友圈,谢绝媒体或机构未经授权以任何形式转发/复制至其他平台。转发授权请在「创鉴汇」微信公众号留言联系我们。读者们请星标⭐创鉴汇,第一时间收到我们的推送新型疗法专题PROTAC疗法 | 细胞疗法 | 基因疗法 | 溶瘤病毒 | mRNA | 反义寡核苷酸 | RNAi | 寡核苷酸疗法 | 抗体偶联药物 | 人工智能Medtech专题生物材料 | 液体活检 | 二尖瓣技术 | 手术机器人 | 神经介入 | POCT | 脑机接口 | 人工心脏 |投资机构专题高瓴创投 | 礼来亚洲基金 | 红杉中国 | 诺华 | RA Capital Management | 启明创投 | 奥博资本 | Alexandria Venture Investments | Flagship pioneering | 腾讯 国家园区专题以色列 | 日本 | 英国 | 张江 点击“在看”,分享创鉴汇健康新动态
小分子药物创新药细胞疗法抗体基因疗法
100 项与 Harmonic Discovery, Inc. 相关的药物交易
登录后查看更多信息
100 项与 Harmonic Discovery, Inc. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2024年10月02日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
临床前
1
登录后查看更多信息
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
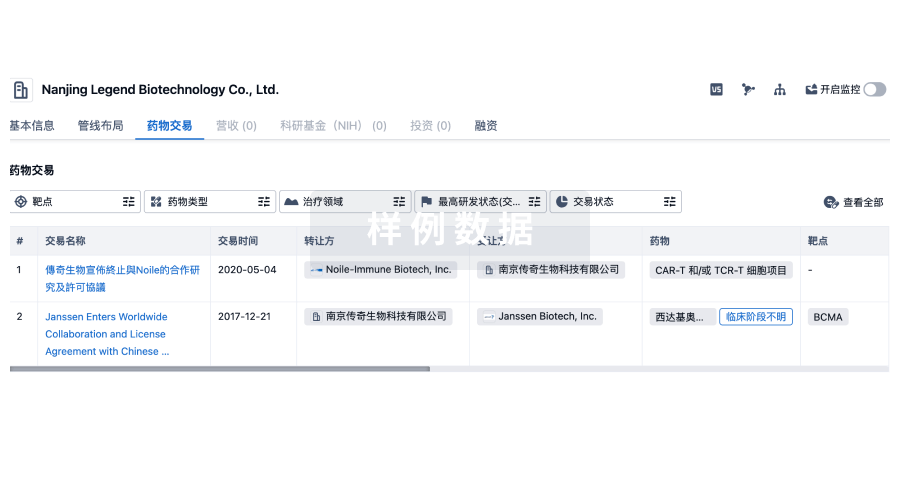
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
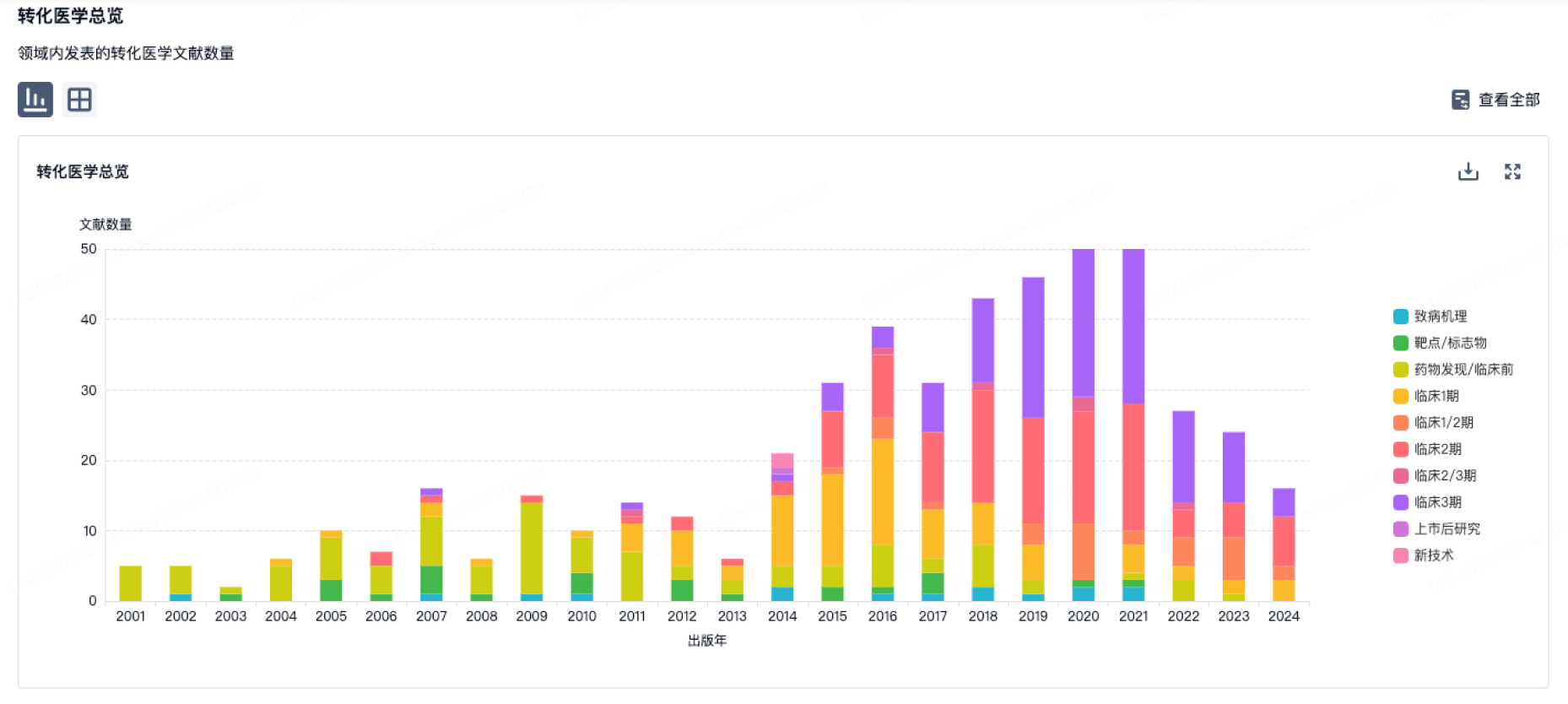
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
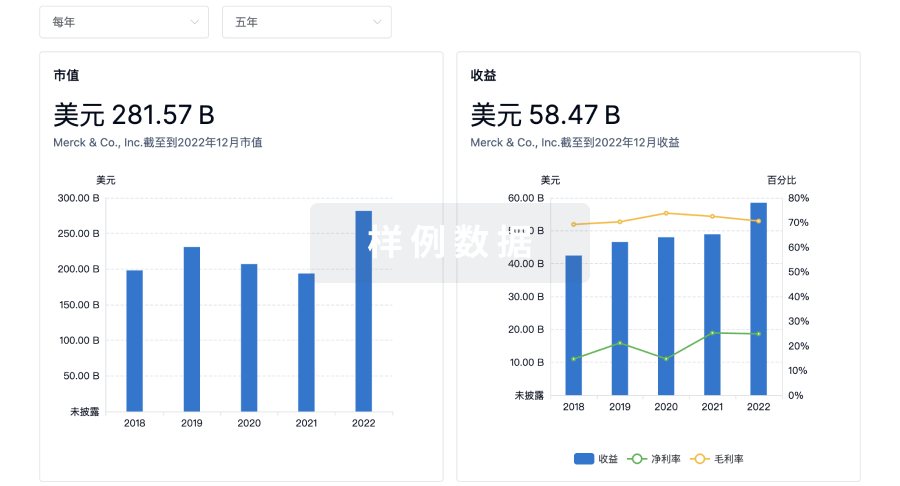
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
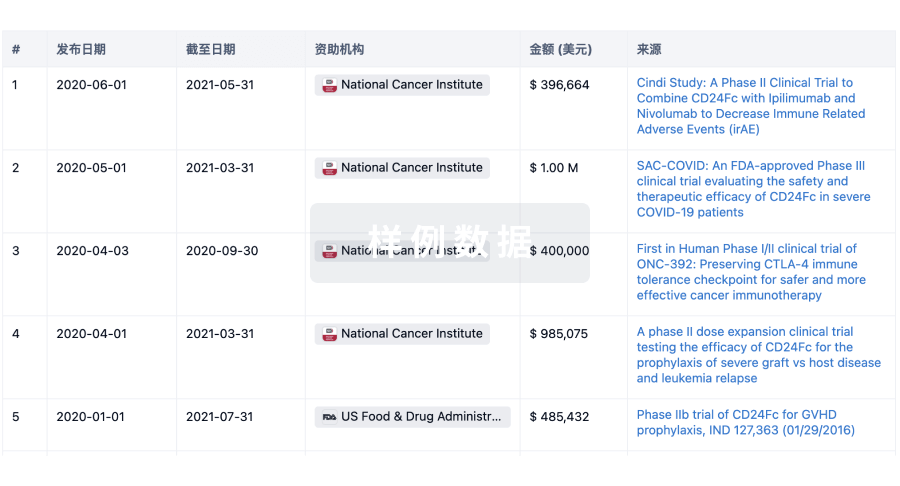
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
标准版
¥16800
元/账号/年
新药情报库 | 省钱又好用!
立即使用
来和芽仔聊天吧
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用