预约演示
更新于:2025-09-09

Caribou Biosciences, Inc.
更新于:2025-09-09
概览
标签
免疫系统疾病
肿瘤
血液及淋巴系统疾病
通用型CAR-T
CAR-T
小分子化药
疾病领域得分
一眼洞穿机构专注的疾病领域
暂无数据
技术平台
公司药物应用最多的技术
暂无数据
靶点
公司最常开发的靶点
暂无数据
| 排名前五的药物类型 | 数量 |
|---|---|
| 通用型CAR-T | 2 |
| CAR-T | 2 |
| 小分子化药 | 1 |
关联
5
项与 Caribou Biosciences, Inc. 相关的药物作用机制 B2M抑制剂 [+4] |
非在研适应症- |
最高研发阶段临床1期 |
首次获批国家/地区- |
首次获批日期- |
靶点 |
作用机制 CD19抑制剂 |
非在研适应症- |
最高研发阶段临床1期 |
首次获批国家/地区- |
首次获批日期- |
WO2024228989
专利挖掘靶点 |
作用机制- |
非在研适应症- |
最高研发阶段药物发现 |
首次获批国家/地区- |
首次获批日期- |
5
项与 Caribou Biosciences, Inc. 相关的临床试验NCT06752876
A Phase 1, Multicenter, Open-Label Study of CB-010, a CRISPR-Edited Allogeneic Anti-CD19 CAR-T Cell Therapy, in Patients With Refractory Systemic Lupus Erythematosus (GALLOP)
This is a Phase 1 study to evaluate the safety and efficacy of a single infusion of CB-010 in patients with refractory Systemic Lupus Erythematosus (SLE) with cohorts for lupus nephritis (LN) and extrarenal lupus (ERL).
开始日期2025-06-01 |
申办/合作机构 |
NCT06128044
A Phase 1, Multicenter, Open-Label Study of CB-012, a CRISPR-Edited Allogeneic Anti-CLL-1 CAR-T Cell Therapy in Patients With Relapsed/Refractory Acute Myeloid Leukemia
CB-012 is an allogeneic chimeric antigen receptor (CAR-T) cell therapy that targets C-type lectin-like molecule-1 (CLL-1). This is a Phase 1 study to evaluate the safety, preliminary efficacy, and pharmacokinetics, of CB-012 (the study treatment) in adults with acute myeloid leukemia (AML) that has come back after prior treatment (relapsed) or did not respond or is no longer responding to other treatment (refractory). Participants must have received at least 1 but not more than 3 prior lines of treatment for AML .
开始日期2024-02-08 |
申办/合作机构 |
NCT05722418
A Phase 1, Multicenter, Open-Label Study of CB-011, a CRISPR-Edited Allogeneic Anti-BCMA CAR-T Cell Therapy in Patients With Relapsed/Refractory Multiple Myeloma (CaMMouflage Trial)
This is a Phase 1 study to evaluate the safety of CB-011 (the study treatment), an allogeneic chimeric antigen receptor (CAR-T) cell therapy that targets the B cell maturation antigen (BCMA), to determine the best dose of CB-011, and to assess the effectiveness of CB-011 in treating multiple myeloma that has come back (relapsed) or that is no longer responding to other treatment (refractory).
开始日期2023-02-06 |
申办/合作机构 |
100 项与 Caribou Biosciences, Inc. 相关的临床结果
登录后查看更多信息
0 项与 Caribou Biosciences, Inc. 相关的专利(医药)
登录后查看更多信息
11
项与 Caribou Biosciences, Inc. 相关的文献(医药)2024-04-02·Cancer immunology research
High-Specificity CRISPR-Mediated Genome Engineering in Anti-BCMA Allogeneic CAR T Cells Suppresses Allograft Rejection in Preclinical Models
Article
作者: An, Zili ; Guo, Raymond ; Skoble, Justin ; Scherer, Jessica ; Fowler, Tristan W. ; Nyer, David B. ; Kohrs, Bryan ; Bryan, Mara ; Toh, Mckenzi S. ; Larroca Vicena, Vanina ; Schilling, Benjamin ; Degagné, Émilie ; Kufeldt, Heinrich J. ; Irby, Matthew J. ; Banh, Lynda ; Kwong, George ; Owen, Arthur L.G. ; Thompson, Matthew ; Fuller, Christopher K. ; Reyes, Gustavo A. ; Shaw, McKay ; Mutha, Devin ; Smith, Stephen C. ; McSweeney, Kyle ; Garner, Elizabeth ; Churchward, Glen ; Davis, Ryan T. ; Kanner, Steven B. ; Stanaway, Morena ; Gradia, Scott ; Donohoue, Paul D. ; Roy, Suparna ; Edwards, Leslie ; Ruan, Finey
Abstract:
Allogeneic chimeric antigen receptor (CAR) T cell therapies hold the potential to overcome many of the challenges associated with patient-derived (autologous) CAR T cells. Key considerations in the development of allogeneic CAR T cell therapies include prevention of graft-vs-host disease (GvHD) and suppression of allograft rejection. Here, we describe preclinical data supporting the ongoing first-in-human clinical study, the CaMMouflage trial (NCT05722418), evaluating CB-011 in patients with relapsed/refractory multiple myeloma. CB-011 is a hypoimmunogenic, allogeneic anti–B-cell maturation antigen (BCMA) CAR T cell therapy candidate. CB-011 cells feature 4 genomic alterations and were engineered from healthy donor–derived T cells using a Cas12a CRISPR hybrid RNA–DNA (chRDNA) genome-editing technology platform. To address allograft rejection, CAR T cells were engineered to prevent endogenous HLA class I complex expression and overexpress a single-chain polyprotein complex composed of beta-2 microglobulin (B2M) tethered to HLA-E. In addition, T-cell receptor (TCR) expression was disrupted at the TCR alpha constant locus in combination with the site-specific insertion of a humanized BCMA-specific CAR. CB-011 cells exhibited robust plasmablast cytotoxicity in vitro in a mixed lymphocyte reaction in cell cocultures derived from patients with multiple myeloma. In addition, CB-011 cells demonstrated suppressed recognition by and cytotoxicity from HLA-mismatched T cells. CB-011 cells were protected from natural killer cell–mediated cytotoxicity in vitro and in vivo due to endogenous promoter-driven expression of B2M–HLA-E. Potent antitumor efficacy, when combined with an immune-cloaking armoring strategy to dampen allograft rejection, offers optimized therapeutic potential in multiple myeloma.See related Spotlight by Caimi and Melenhorst, p. 385
2023-07-01·Cytotherapy
Allogeneic chimeric antigen receptor-T cells with CRISPR-disrupted programmed death-1 checkpoint exhibit enhanced functional fitness.
Article
作者: Fowler, Tristan W. ; Williams, Carolyn ; van Overbeek, Megan ; Bryan, Mara ; Lau, Elaine ; Skoble, Justin ; Kwong, George ; Kohrs, Bryan ; Garner, Elizabeth ; Smith, Stephen C. ; Clarke, Starlynn C. ; Fuller, Chris K. ; McCawley, Shannon ; Sun, Bee-Chun ; Davis, Ryan T. ; Irby, Matthew ; Gradia, Scott ; Kanner, Steven B. ; Donohoue, Paul D. ; Lilley, Graham W. J. ; Storlie, Meghan ; Banh, Lynda ; Edwards, Leslie
BACKGROUND AIMS:
Therapeutic disruption of immune checkpoints has significantly advanced the armamentarium of approaches for treating cancer. The prominent role of the programmed death-1 (PD-1)/programmed death ligand-1 axis for downregulating T cell function offers a tractable strategy for enhancing the disease-modifying impact of CAR-T cell therapy.
METHODS:
To address checkpoint interference, primary human T cells were genome edited with a next-generation CRISPR-based platform (Cas9 chRDNA) by knockout of the PDCD1 gene encoding the PD-1 receptor. Site-specific insertion of a chimeric antigen receptor specific for CD19 into the T cell receptor alpha constant locus was implemented to drive cytotoxic activity.
RESULTS:
These allogeneic CAR-T cells (CB-010) promoted longer survival of mice in a well-established orthotopic tumor xenograft model of a B cell malignancy compared with identically engineered CAR-T cells without a PDCD1 knockout. The persistence kinetics of CB-010 cells in hematologic tissues versus CAR-T cells without PDCD1 disruption were similar, suggesting the robust initial debulking of established tumor xenografts was due to enhanced functional fitness. By single-cell RNA-Seq analyses, CB-010 cells, when compared with identically engineered CAR-T cells without a PDCD1 knockout, exhibited fewer Treg cells, lower exhaustion phenotypes and reduced dysfunction signatures and had higher activation, glycolytic and oxidative phosphorylation signatures. Further, an enhancement of mitochondrial metabolic fitness was observed, including increased respiratory capacity, a hallmark of less differentiated T cells.
CONCLUSIONS:
Genomic PD-1 checkpoint disruption in the context of allogeneic CAR-T cell therapy may provide a compelling option for treating B lymphoid malignancies.
2023-01-01·Methods in molecular biology (Clifton, N.J.)
Clinical Applications of Flow Cytometry in Cancer Immunotherapies: From Diagnosis to Treatments
Article
作者: Mishra, Hemant K
The scope of flow cytometry is rapidly expanding in the diagnosis of various cancers, and it is being used routinely as an aid in classifying leukemias and lymphomas. There are several applications of flow cytometry to enumerate tumorigenic anomalies in patients. The unusual distribution of cells in various locations, their DNA content, cell proliferation rate, dysregulated expression of several surface receptors, and expression of tumor antigens are some examples that can be characterized by using different flow cytometry-based techniques. For instance, the differential diagnosis between chronic lymphocytic leukemia (CLL) and various other mature B-cell neoplasms can be made by immunophenotyping in combination with absolute counting of numerous cellular subsets or by enumerating their percent distributions. Flow cytometry has several advantages over conventional techniques which include the ability to acquire a multiparametric data in a relatively shorter time and facilitate the comparative analysis of specific cellular subsets in an efficient manner.In addition to diagnosis, there are several other applications of flow cytometry in the management of various cancers which include treatment monitoring or even selecting a personalized precision-based immunotherapy in synch with advanced genetic tests to increase the chances of favorable prognosis and complete remission. The detection of chimeric antigen receptors (CARs) on various engineered effector cells can also be determined along with their specificity in engaging the targets. Furthermore, the assessment of numerous immunological parameters, their effector functions and potencies including the proliferation dynamics, cytokine secretion profiles, and activation efficiencies can also be measured before starting immunotherapies in patients.This chapter is a brief overview of flow cytometry applications in the diagnosis and treatment strategies of various cancers.
206
项与 Caribou Biosciences, Inc. 相关的新闻(医药)2025-07-21
- "Graduation" space designed to help growing life science companies remain in the ecosystem and thrive
BERKELEY, Calif., July 21, 2025 /PRNewswire/ -- Bakar Labs, the University of California, Berkeley's flagship incubator for life science, energy, and materials startups, announced today the launch of a new building on campus that will provide critical infrastructure for life science startups scaling beyond the earliest stages.
Continue Reading
A rendering of the Innovative Genomics Institute–Bakar Labs building, which will be constructed at the corner of Oxford St. and University Ave. on the north edge of the Berkeley Innovation Zone. Image: DGA + Weiss/Manfredi
Launched in partnership with the Innovative Genomics Institute (IGI), the IGI-Bakar Labs building will house expansion space for biotech companies transitioning out of early incubators as they exceed 20-30 employees. This space will help retain high-potential startups in the Berkeley ecosystem by offering state-of-the-art labs and offices, flexibility, community support, and proximity to campus resources so companies can continue to grow. Construction is projected to be complete in late 2028.
"It's important to keep startups in Berkeley because they're not just creating jobs for skilled graduates, they're feeding a cycle of innovation," said David Schaffer, a UC Berkeley professor of chemical and biomolecular engineering and director of QB3 and Bakar Labs. "When leaders stay connected to the ecosystem, they return to Bakar Labs as mentors, advisors and role models. That continuity strengthens the entire community."
The IGI-Bakar Labs building fills a long-standing gap between incubator and independent space, with demand for "next-stage" lab space poised to outpace supply in the Bay Area.
The IGI, founded by Nobel laureate and UC Berkeley professor Jennifer Doudna, develops genome editing solutions for unmet needs in health and agriculture. Doudna herself has co-founded many companies that are commercializing CRISPR genome editing; her first, Caribou Biosciences, launched in the QB3 Garage incubator at Berkeley. Two other Doudna Lab spinoffs, Azalea Therapeutics and CatenaBio, are current Bakar Labs tenants. The close proximity of IGI research labs with active, growing companies in the new building could lead to mutually beneficial partnerships in developing impactful genomic technologies.
"Bakar Labs gave us the foundation to move fast and stay focused during our earliest days," said Sophia Lugo, CEO & co-founder of Radar Therapeutics. "The community, resources, and collaborative energy were unlike anything else in the Bay Area. As we've grown, it's meant everything to have a path forward that lets us scale without losing that connection. This new expansion space is exactly what companies like ours need to keep building, without leaving behind the ecosystem that helped us get here."
"Bakar Labs provides what every biotech startup needs—an environment that accelerates both scientific rigor and company formation. At Ray Therapeutics, this ecosystem helped us refine our strategy, build a strong team, and move quickly toward the clinic. It's more than a lab space, it's an ecosystem for life science companies," said Peter Francis, MD, PhD, CSO and CMO at Ray Therapeutics.
Plans for the IGI-Bakar Labs building were approved by the Regents of the University of California on Thursday, July 17. Floors 1-3 will hold IGI offices and labs, and floors 4-6 will be occupied by Bakar Labs, which will also enjoy a rooftop conference space with views overlooking the Berkeley campus. With 72,000 square feet designed to support up to 14 growth-stage companies, the expansion will accommodate scaling life science companies while maintaining the spirit of ingenuity and collaboration that defines the Bakar Labs community.
The new facility builds on Bakar Labs' success since launching its first location on the south side of campus, in a landmark building originally home to the Berkeley Art Museum and renovated into a biotech incubator that has helped nurture more than 45 seed-stage startups. Collectively, since 2021 Bakar-incubated companies have raised more than $700 million in capital and created hundreds of jobs across the region.
The IGI-Bakar Labs community will also benefit from BEVC, a venture firm embedded in the expanding Bakar ecosystem that partners with entrepreneurs to start and finance startup companies.
"We created the spaces where breakthrough ideas are born, and now we're building the spaces where they grow," said Schaffer. "This new facility represents the next phase of our mission to support startups that have validated their technology and are ready to scale, while staying connected to the community that shaped them. Berkeley's innovation network is serving as a launchpad for the future of biotech innovation."
Schaffer, a serial entrepreneur himself, has co-founded seven biotech spinouts from his UC Berkeley lab, including 4D Molecular Therapeutics (NASDAQ FDMT), Ignite Immunotherapies (acquired by Pfizer), and Rewrite (acquired by Intellia).
The original Bakar Labs location is known for its striking architecture, which fosters creativity, collaboration, and a strong sense of identity. The IGI-Bakar Labs building will carry forward that design philosophy, offering a visually distinctive and inspiring space that reflects the bold innovation happening inside its walls.
Tenants at the new Bakar Labs site will benefit from access to scientific and networking events at IGI and Bakar Labs; interactions with fellow tenants at IGI-Bakar and three other Bakar Labs incubators; exposure to Bakar Labs industry affiliates for potential business relationships and partnerships; and a high-quality hiring pool of 35,000 UC Berkeley students.
About Bakar Labs
Bakar Labs is UC Berkeley's flagship incubator supporting innovation in life sciences, energy, and materials. Operated by QB3, Bakar Labs provides extensive equipment, lab and office facilities, a network of investors and pharma/biotech companies, and a community of like-minded entrepreneurs to help startups grow. By 2029, Bakar Labs will support more than 100 early-stage companies from around the world focused on translating innovations that promise to improve human and planetary health. No UC Berkeley affiliation is required to join. For information about how to join or form a partnership, visit bakarlabs.org.
Contact:
Kaspar Mossman
[email protected]
SOURCE Bakar Labs
WANT YOUR COMPANY'S NEWS FEATURED ON PRNEWSWIRE.COM?
440k+
Newsrooms &
Influencers
9k+
Digital Media
Outlets
270k+
Journalists
Opted In
GET STARTED
并购
2025-07-15
BOSTON, July 15, 2025 (GLOBE NEWSWIRE) -- Aegle Therapeutics, a clinical-stage dermatology-focused biopharmaceutical company developing novel therapies for those living with rare and severe skin diseases, today announced the appointment of Scott Braunstein, M.D., as Chairman of the Board. The company’s lead product candidate AGLE-102, a composite of mesenchymal stem cell derived extracellular vesicles (MSC-EVs), is currently being evaluated in a Phase 1/2a trial for the treatment of recessive dystrophic epidermolysis bullosa (RDEB), a rare pediatric skin blistering disorder. “We are excited to welcome Dr. Braunstein as Chairman and look forward to his guidance as we advance the development of AGLE-102 as a novel therapeutic platform for rare and severe dermatological disorders,” said Shelley Hartman, CEO of Aegle. “Dr. Braunstein’s extensive industry experience will be a valuable addition to our leadership team as we continue to develop AGLE-102 for RDEB and other subtypes of epidermolysis bullosa.” Dr. Braunstein commented, “Aegle is at the forefront of advancing extracellular vesicle-based therapies which have exciting potential across a broad spectrum of refractory dermatological disorders. I look forward to working with Shelley and the management team to advance their current mission of offering patients and their families a novel and simple biological solution for the treatment of EB.” Scott Braunstein, M.D.Scott Braunstein, M.D., brings over 30 years of knowledge and experience from diverse biotechnology and pharmaceutical industry vantage points. Dr. Braunstein served as President and Chief Executive Officer of Marinus Pharmaceuticals, Inc., a company focused on rare forms of epilepsy, until acquired by Immedica Pharma AB. Prior to that, he served as Senior Vice President, Strategy and Chief Operating Officer at Pacira Pharmaceuticals, Inc., a specialty pharmaceutical company. Prior to Pacira, Dr. Braunstein spent 12 years with J.P. Morgan Asset Management as a Managing Director, Healthcare Analyst and the Portfolio Manager of the JP Morgan Global Healthcare Fund. Dr. Braunstein has been an operating partner at Aisling Capital since 2015. He currently serves on the board of directors for atai Life Sciences N.V., Caribou Biosciences, Inc., RAPT Therapeutics and Site One Therapeutics, a clinical-stage neurology company that is being acquired by Eli Lilly. Dr. Braunstein previously served on the board of directors of Trevena Inc., Esperion Therapeutics, Inc., Ziopharm Oncology Inc., Protara Therapeutics, Inc. (previously known as ArTara Therapeutics, Inc.) and Constellation Pharmaceuticals, Inc. Dr. Braunstein began his career as a practicing physician at the Summit Medical Group and as an Assistant Clinical Professor at Albert Einstein College of Medicine and Columbia University Medical Center. Dr. Braunstein received an M.D. from the Albert Einstein College of Medicine and a BASc in biology from Cornell University. About AGLE-102™ AGLE-102 is a composite of extracellular vesicles isolated from allogeneic mesenchymal stem cells using Aegle's proprietary methods. The product is a composite of native EVs and their associated complex assemblies of biologic molecules such as proteins, peptides, ligands and nucleic acids that induce a wide variety of biological effects in recipient cells, including reducing inflammation, modulating the immune system and promoting regenerative healing. About Aegle Therapeutics CorporationAegle Therapeutics is a clinical-stage dermatology-focused biopharmaceutical company developing novel EV therapies to treat rare and severe dermatological disorders with significant unmet medical need. The company’s lead product candidate, AGLE-102, is currently in a phase 1/2a study for the treatment of recessive dystrophic epidermolysis bullosa and has successfully completed a proof-of-concept study in severe burns. AGLE-102 has the potential to treat all subtypes of epidermolysis bullosa as well as a broad range of other dermatological indications characterized by inflammation resulting from an underlying immune imbalance. For more information about Aegle Therapeutics, please visit www.aegletherapeutics.com. For More Information Contact:info@aegletherapeutics.com
高管变更
2025-07-14
近一年多来,艾伯维频频出手,通过引进或并购的方式扩充自己的TCE及In vivo CAR-T管线及平台。我们可以怀疑艾伯维的研发能力,但我们要相信其战略眼光。读懂艾伯维的研发策略能帮助国内企业更好的布局相关产品及技术。艾伯维的修美乐(TNFα抑制剂)曾经霸榜全球药王11年,但终究由于专利到期而走下了神坛。为了填补专利悬崖带来的销售空白,艾伯维提前十多年布局,通过自研+引进的方式成功的开发出乌帕替尼(JAK1抑制剂)及利生奇珠单抗(IL-23抑制剂),2024年销售分别达59.7及117.2亿美元。肿瘤板块是艾伯维四大业务板块之一,而血液瘤产品则占据了其近年来的主要收入。但艾伯维的两款重磅血液瘤产品,伊布替尼(Imbruvica;BTK抑制剂)和维奈克拉(Venclexta;Bcl-2抑制剂),均面临着专利悬崖及竞争产品的双重压力。为了破局,艾伯维在血液瘤领域也采取了自研+引进的方式。自研方面,艾伯维开发了下一代BTK降解剂(ABBV-101)、Bcl-2抑制剂(ABBV-453)、MALT1抑制剂(ABBV-525)及CD19-ADC(ABBV-319),但目前均处于临床1期。而在引进方面,艾伯维做了广泛的布局。特别是从最近一年的引进及并购交易来看,艾伯维在血液瘤的布局侧重于TCE及In vivo CAR-T。1、TCE其实,艾伯维早在二十年前就开发了自己的双抗平台DVD-Ig,并在十年多前开始尝试研发自己的CD3 TCE,但至今未有自研产品进入临床。2020年6月,艾伯维以7.5亿美元预付款+31.5亿美元里程碑付款的巨额交易,获得Genmab研发的Epcoritamab(CD3/CD20双抗)的全球权益(美国和日本市场由双方共同商业化)(图1)。图1. Epcoritamab结构示意图当时,Epcoritamab处于临床1/2期,而罗氏及再生元的CD3/CD20双抗已进入临床2期。后来,Mosunetzumab、Epcoritamab、Glofitamab分别于2022年12月、2023年5月、2023年6月获FDA批准上市,2024年三款药物的销售分别为0.78、2.81、1.9亿美元。尽管Epcoritamab后来居上,在销售上领先于罗氏的两款CD3/CD20双抗,但距离成为重磅药物还有相当距离。2017年10月,艾伯维与Harpoon Therapeutics达成合作协议,前者利用后者的TriTAC平台开发选定靶向药。2019年11月,艾伯维与Harpoon再次签订独家许可协议扩大合作,前者获得当时处于临床1期的CD3/BCMA/HSA三抗(HPN217)的全球开发权(图2)。图2. HPN217结构示意图2021年6月,艾伯维以1.4亿美元的价格收购了TeneoOne(TeneoBio子公司),获得一款处于临床1期的CD3/BCMA双抗(TNB-383B),TNB-383B双抗采用2+1设计,细胞因子释放效应最小化(图3)。图3. TNB-383B结构示意图2023年10月,艾伯维终止了HPN217的进一步开发而专注于TNB-383B(ABBV-383)的开发。2024年6月,艾伯维启动了ABBV-383的临床3期试验。但是,目前全球已有两款CD3/BCMA双抗上市,分别是强生的Teclistamab及辉瑞的Elranatamab,分别于2022年10月及2023年8月获FDA批准上市。而BMS的CD3/BCMA双抗(Alnuctamab)于2024年1月进入临床3期,但仅几个月后就宣布由于公司策略改变而终止了继续开发(详见双抗 vs CAR-T:BMS的取舍)。因此,对于艾伯维来说,ABBV-383实属鸡肋,获批上市至少还需要2-3年时间,而竞争对手强生的CD3/BCMA双抗的下一代产品CD3/BCMA/GPRC5D三抗(JNJ-79635322)已在临床1期取得了不错的疗效(RP2D ORR 100%;预期1年PFS 95%)。这就是艾伯维于今年1月以10.55亿美元的价格由先声再明引进CD3/BCMA/GPRC5D三抗(SIM0500)的重要原因(图4)。图4. SIM0500结构示意图然而,研究MM患者样本显示,GPRC5D的表达存在显著差异,CD38、CD138、BCMA和GPRC5D的中位数和标准差分别为89和113、69和134、58和132、10和166。因此,CD3/BCMA/GPRC5D三抗针对BCMA/GPRC5D均低表达的患者是否有效、药效能否持久存在不确定性。这就能解释为什么艾伯维在引进CD3/BCMA/GPRC5D三抗半年后又以首付7亿美元、总额19.25亿美元从IGI Therapeutics引进了CD3/BCMA/CD38三抗(ISB2001)(图5)。图5. ISB2001结构示意图强生的达雷妥尤(CD38)单抗是MM领域目前的药王,2024年销售达116.7亿美元。而ISB2001主要针对达雷妥尤单抗耐药的患者进行设计,其CD38臂与达雷妥尤单抗的抗原结合域有着显著区别并且对CD38弱表达的细胞仍有杀伤功能,而CD38表达降低是达雷妥尤单抗耐药的重要机制之一。ISB2001目前处于临床1期剂量爬坡最后阶段,据今年ASCO大会上公开信息显示,ISB2001在极难治RRMM患者中展现出高缓解率(ORR 74%)、深度反应(包括MRD阴性75%)及良好安全性(图6)。图6. ISB2001治疗末线或CD38耐药MM患者预测艾伯维可能同时开发SIM0500及ISB2001,并通过找到临床Biomarker(如GPRC5D)用来区分两款药物不同的适用患者,甚至效仿强生的Teclistamab+Talquetamab(CD3/GPRC5D双抗)联用以及JNJ-79635322+达雷妥尤单抗联用,开发两款三抗的联用。当然从艾伯维过往的行事方式来看,其还有可能通过在临床上比较SIM0500及ISB2001的疗效而二选一。除了直接引进临床阶段TCE产品外,艾伯维还通过与拥有下一代TCE技术的Biotech合作,开发临床前产品。而从其近一年的两起交易来看,艾伯维对包含共刺激信号的TCE很感兴趣。2024年10月,艾伯维与EvolveImmune Therapeutics达成协议,前者以6500万美元的首付款、14亿美元的里程碑付款利用后者的TCE平台开发多款肿瘤药物(详见下一代TCE之共刺激信号的选择:我们从CAR-T学到了什么?)。2025年1月,艾伯维与AbCellera扩大早期达成的合作,共同开发TCE。其中,AbCellera将负责相关发现工作,而艾伯维则拥有发现和商业化由此产生的抗体的权利。通过分析AbCellera相关平台推测,双方合作的焦点可能是AbCellera的CD3/CD28/TAA平台(图7)。图7. AbCellera的CD28平台示意图2、In vivo CAR-T自2017年首次获批以来,自体CAR-T细胞疗法彻底改变了血液瘤的治疗,给对以往疗法失败的患者带来了希望。与此同时,近年来自体CAR-T细胞疗法在自免领域也取得了一系列的突破。然而,目前的自体CAR-T疗法需要采集患者的细胞,并在遥远的生产基地进行数周甚至数月的改造。因此,这些疗法虽然能够显著改善患者的预后,但往往负担过重,未得到广泛的应用。艾伯维作为血液瘤及自免领域的领导者,虽然错过了第一批进入细胞治疗的机会,但希望通过引进项目及技术平台实现弯道超车。2021年2月,艾伯维向Caribou预付了4000万美元,为开发两款新靶点的通用型CAR-T疗法提供3亿美元的研发和注册里程碑付款。艾伯维使用Caribou的下一代Cas12a CRISPR杂合RNA-DNA(chRDNA)基因组编辑和细胞治疗技术,来研究和开发两种针对艾伯维指定靶点的新型CAR-T细胞疗法。Caribou负责临床前研究、开发和制造,艾伯维负责细胞疗法的临床开发、商业化和制造。尽管在2023年,Caribou公布了其通用型CD19 CAR-T疗法CB-010的1期临床试验的积极数据,接受治疗的患者的ORR达94%(15/16),完全缓解(CR)率达69%(11/16),但艾伯维于2023年10月终止了与Caribou的合作。有趣的是,仅仅过了三个月后的2024年1月,艾伯维与Umoja宣布达成合作,利用后者的VivoVec平台(图8)开发针对CD19和其它靶点的In vivo CAR-T疗法,用于治疗恶性血液肿瘤和难治型实体瘤。图8. VivoVec平台示意图2025年6月,艾伯维宣布以21亿美元现金收购成立仅三年、处于临床阶段的生物技术公司Capstan Therapeutics。根据协议,艾伯维此次收购 Capstan,不仅将获得处于临床1期用于治疗B细胞自免疾病的In vivo CD19-CAR-T(CPTX2309)这一极具潜力的在研产品,还将获得Capstan专有的tLNP平台技术(图9)。图9. CPTX2309结构示意图不难看出,通过引进及并购,艾伯维已经获得了In vivo CAR-T的两大主流技术路线,通过慢病毒递送(以Umoja、EsoBiotec为代表)或通过LNP递送(以Capstan为代表),很可能实现弯道超车成为该领域的领导者。结合自研的BTK降解剂(ABBV-101)、CD19-ADC(ABBV-319)等产品,通过TCE及In vivo CAR-T的引进或并购的加持,艾伯维正在B细胞相关疾病领域进行广泛布局并构筑了深深的护城河,而这些药物的单药或联用将继续巩固艾伯维在自免及肿瘤领域的领导地位。
免疫疗法细胞疗法并购抗体药物偶联物临床1期
100 项与 Caribou Biosciences, Inc. 相关的药物交易
登录后查看更多信息
100 项与 Caribou Biosciences, Inc. 相关的转化医学
登录后查看更多信息
组织架构
使用我们的机构树数据加速您的研究。
登录
或

管线布局
2025年10月04日管线快照
管线布局中药物为当前组织机构及其子机构作为药物机构进行统计,早期临床1期并入临床1期,临床1/2期并入临床2期,临床2/3期并入临床3期
药物发现
3
2
临床1期
其他
2
登录后查看更多信息
当前项目
| 药物(靶点) | 适应症 | 全球最高研发状态 |
|---|---|---|
CB-010 ( CD19 ) | 难治性B细胞淋巴瘤 更多 | 临床1期 |
CB-011 ( B2M x BCMA x HLA-E ) | 难治性多发性骨髓瘤 更多 | 临床1期 |
WO2024228989 ( BCMA )专利挖掘 | 罕见病 更多 | 药物发现 |
WO2024107646 ( CLL-1 )专利挖掘 | 血液及淋巴系统疾病 更多 | 药物发现 |
WO2025006880 ( ROR1 )专利挖掘 | 肿瘤 更多 | 药物发现 |
登录后查看更多信息
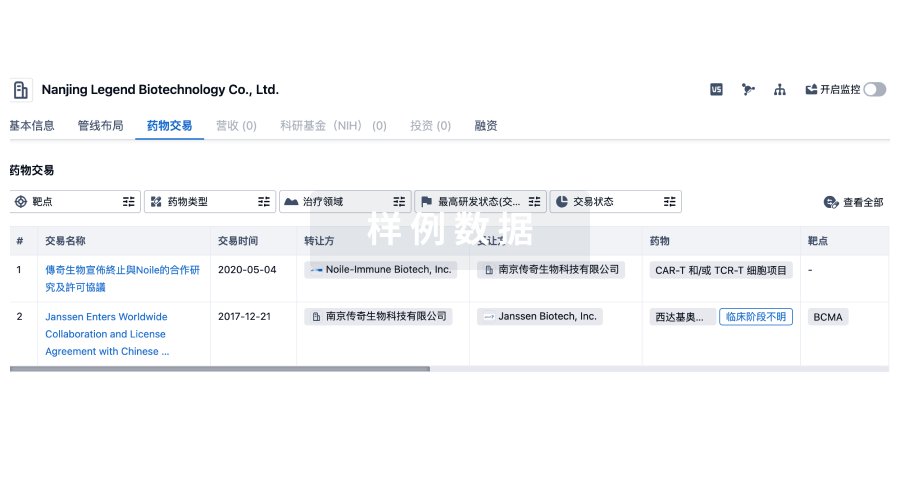
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
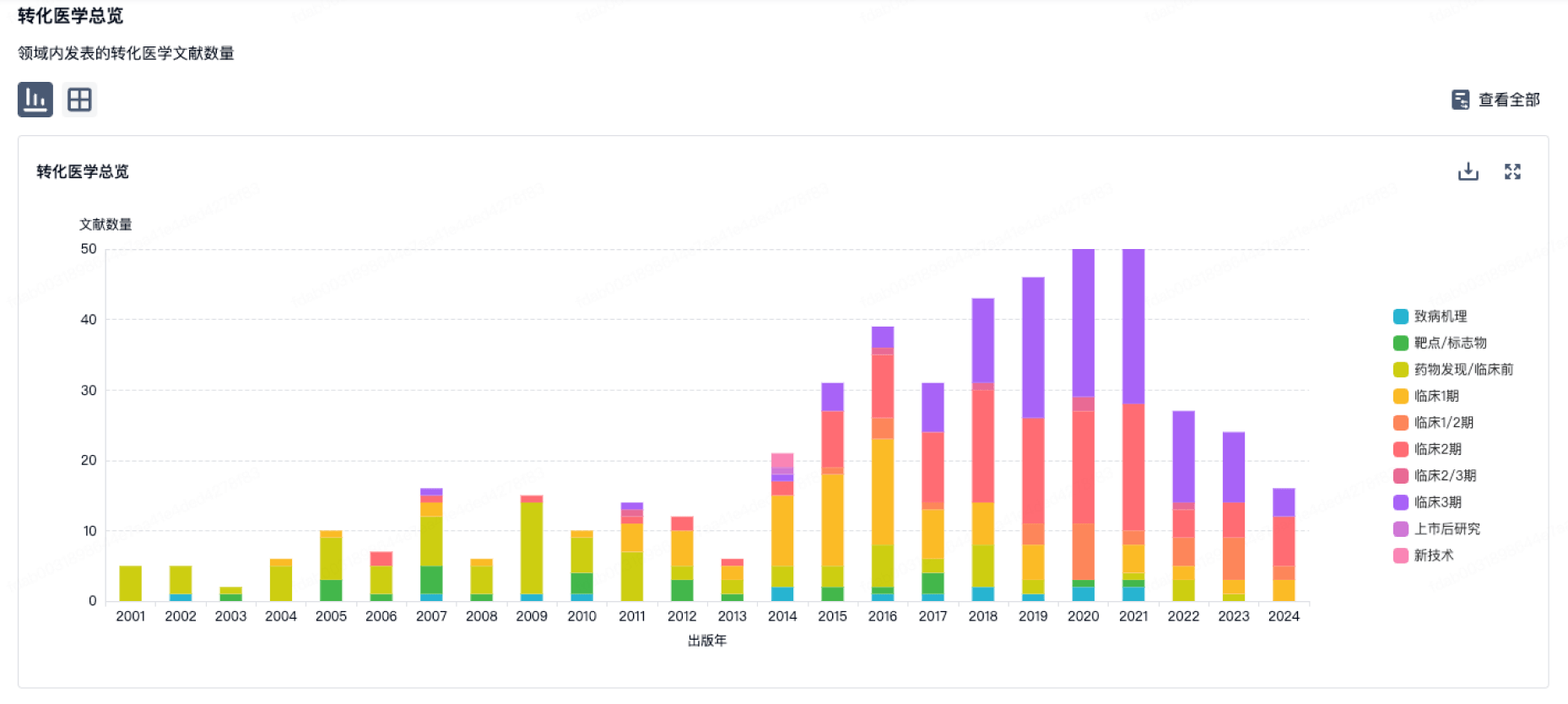
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
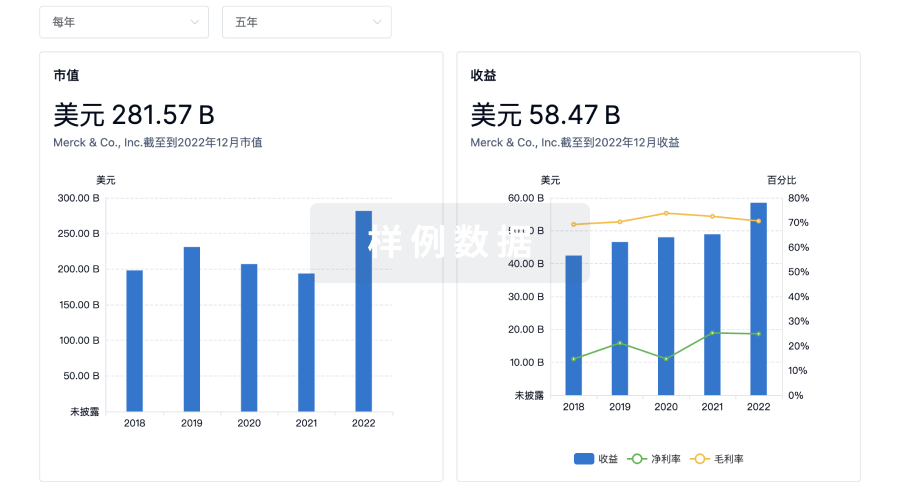
营收
使用 Synapse 探索超过 36 万个组织的财务状况。
登录
或
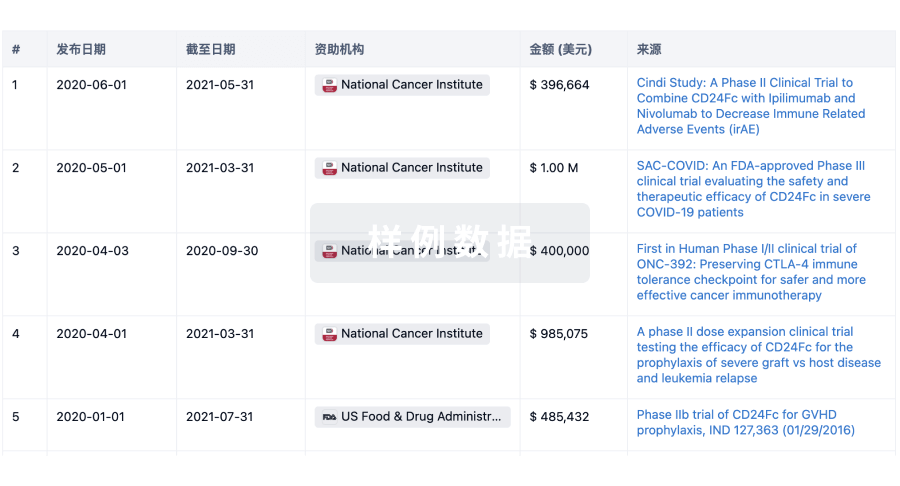
科研基金(NIH)
访问超过 200 万项资助和基金信息,以提升您的研究之旅。
登录
或
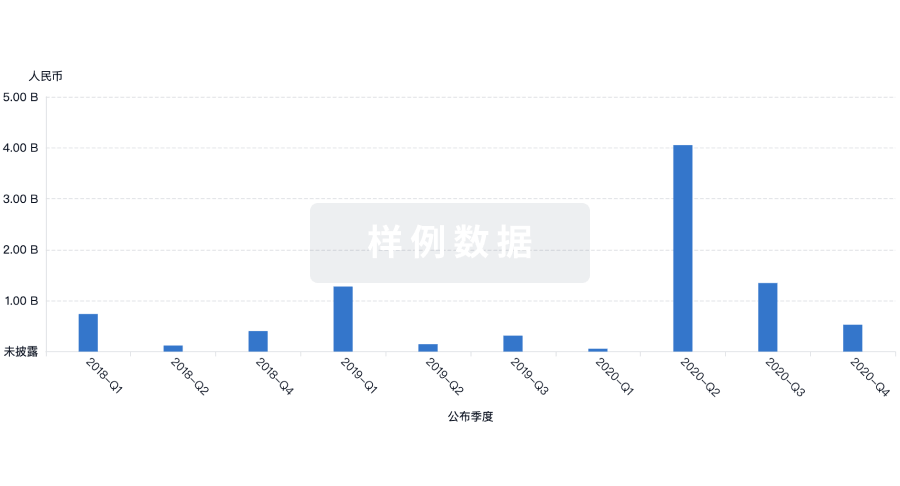
投资
深入了解从初创企业到成熟企业的最新公司投资动态。
登录
或
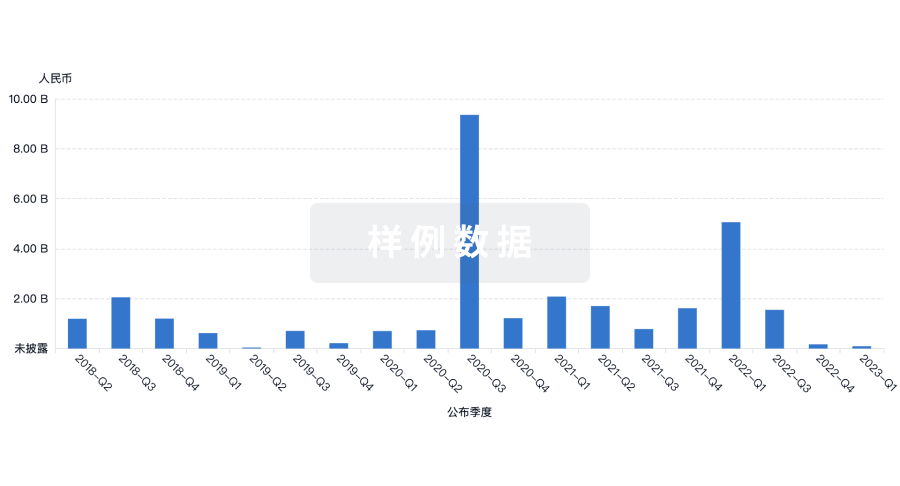
融资
发掘融资趋势以验证和推进您的投资机会。
登录
或
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用