预约演示
更新于:2025-10-25
Encainide Hydrochloride
盐酸恩卡尼
更新于:2025-10-25
概要
基本信息
非在研机构- |
权益机构- |
最高研发阶段批准上市 |
最高研发阶段(中国)- |
特殊审评- |
登录后查看时间轴
结构/序列
分子式C22H29ClN2O2 |
InChIKeyOJIIZIWOLTYOBS-UHFFFAOYSA-N |
CAS号66794-74-9 |
关联
2
项与 盐酸恩卡尼 相关的临床试验NCT00000526
Cardiac Arrhythmia Suppression Trial (CAST)
To determine whether drug treatment of asymptomatic ventricular arrhythmias in post-myocardial infarction patients reduced the incidence of sudden cardiac death and total mortality.
开始日期1986-08-01 |
NCT00000504
Cardiac Arrhythmia Pilot Study (CAPS)
To compare the effectiveness of various drugs and drug combinations in suppressing complex ventricular arrhythmias, and to evaluate their safety.
开始日期1982-09-01 |
100 项与 盐酸恩卡尼 相关的临床结果
登录后查看更多信息
100 项与 盐酸恩卡尼 相关的转化医学
登录后查看更多信息
100 项与 盐酸恩卡尼 相关的专利(医药)
登录后查看更多信息
450
项与 盐酸恩卡尼 相关的文献(医药)2025-09-01·BRITISH JOURNAL OF CLINICAL PHARMACOLOGY
Propafenone‐ vs. amiodarone‐associated adverse cardiac outcomes in patients with atrial fibrillation and heart failure
Article
作者: Chen, Bi‐Li ; Chien, Li‐Nien ; Lin, Shing‐Jong ; Lin, Yi‐Cheng ; Huang, Chun‐Yao ; Chen, Li‐Ying ; Hsu, Chien‐Yi ; Lip, Gregory Y. H.
Aims:
Clinical trials have shown an increased risk of death in patients with recent myocardial infarction who received antiarrhythmic drugs such as flecainide, encainide or moricizine, especially in the presence of associated structural heart disease such as cardiac dysfunction. This study aimed to evaluate the safety outcomes of propafenone use in atrial fibrillation patients with heart failure when compared to those of amiodarone use.
Methods:
This population‐based cohort study used the National Health Insurance Research Database in Taiwan. Eligible patients were those who had atrial fibrillation or atrial flutter diagnosis, had heart failure diagnosis, and first received propafenone or amiodarone between 2002 and 2018. The primary endpoints were death due to arrhythmia and the composite proarrhythmic outcome, which consisted of sudden cardiac arrest, arrhythmic death, ventricular arrhythmia and implantation of defibrillator.
Results:
After propensity score matching, the study cohort consisted of 7235 propafenone and 14 470 amiodarone users. Compared to amiodarone, propafenone was associated with significantly lower risk of the composite proarrhythmic outcome (adjusted hazard ratio: 0.52; 95% confidence interval: 0.42–0.64; P < .001). Propafenone users also had lower risk of death owing to arrhythmia compared to amiodarone users (adjusted hazard ratio: 0.22; 95% confidence interval: 0.08–0.65; P = .006). Subgroup analysis and sensitivity analysis showed similar trends, favouring propafenone.
Conclusion:
Propafenone was not significantly associated with increased risk of proarrhythmia and mortality when compared to amiodarone in atrial fibrillation patients with heart failure in contemporary real‐world settings. Prospective studies are needed to determine whether propafenone should definitely be avoided in these patients.
2025-06-01·EUROPEAN JOURNAL OF MEDICINAL CHEMISTRY
CUPID: A free drug discovery platform for the explainable multi-ion channel assessment of cardiotoxicity
Article
作者: Mele, Marco ; Belgiovine, Valentina ; Trisciuzzi, Daniela ; Mele, Antonietta ; Imbrici, Paola ; Nicolotti, Orazio ; Tondo, Anna Rita ; Ciriaco, Fulvio ; Altomare, Cosimo Damiano ; Cutropia, Francesca ; Amoroso, Nicola ; Siragusa, Lydia ; Togo, Maria Vittoria ; Liantonio, Antonella ; Mastrolorito, Fabrizio ; Gambacorta, Nicola ; Amenduni, Vincenzo ; De Luca, Annamaria
The withdrawal of numerous approved drugs in late development stages, or even from the market, due to safety concerns remains a major challenge, contributing to the high attrition rate in drug discovery and development. Among these concerns, cardiotoxicity is a critical toxicological issue, particularly in oncology, as drugs can induce heart damage by triggering pathological conditions such as arrhythmia, myocardial infarction, and myocardial hypertrophy. Here, we introduce CUPID (Cardiotox Understanding Platform for Intelligent Drug Discovery), an explainable artificial intelligence (XAI) framework designed to predict cardiotoxicity associated with ERG (ether-à-go-go-related gene) potassium, Nav1.5 sodium, and Cav1.2 calcium ion channels. The framework was trained using three carefully curated interspecies experimental datasets from the latest ChEMBL database (release 34) and the CSFP (Core-Substituent Fingerprint), which encodes molecular fragments derived from the decomposition of drug-like small molecules. By leveraging these experimental datasets, highly accurate explainable machine learning models were developed, achieving approximately 80 % accuracy in 5-fold stratified cross-validation analyses. CUPID provides a comprehensive risk assessment of early cardiotoxicity and a key feature is its interpretability: predictions are annotated with clear applicability domain information, while chemical substructures linked to cardiotoxicity risks are highlighted using SHAP (SHapley Additive exPlanations) values. This enhances molecular understanding and facilitates the rational design of safer bioactive compounds. Last but not least, CUPID is freely accessible at https://prometheus.farmacia.uniba.it/cupid.
2025-04-01·JOURNAL OF PHARMACEUTICAL SCIENCES
Optimization of HPLC method for determination of Abraham solvation parameters of pharmaceuticals
Article
作者: Knašienė, Birutė ; Sazonovas, Andrius ; Sadaunykas, Audrius ; Naujalis, Evaldas ; Balčiūnas, Simonas ; Lanevskij, Kiril
Abraham solvation equation (Absolv) is a popular and well-established approach for modeling solute partitioning between phases of different polarity, which finds its applications in both organic chemistry and biomedical fields. Parameters constituting this equation are good quantitative descriptors of solute hydrogen bonding potential that are useful for QSAR modeling of more complex chemical and biological phenomena. Numerous studies dealing with fast determination of Abraham descriptors using HPLC can be found in the literature, but these mostly focus on small un-ionizable industrial and environmental chemicals, whereas experimental data for pharmaceutical molecules are clearly lacking. In the current study we build upon a previously published chromatographic approach, aiming to adapt the method to ionizable drug-like compounds, and optimize it by reducing the number of required HPLC columns. The analysis involves determination of the overall H-bond acidity (A), H-bond basicity (B) and polarity/polarizability (S) descriptors for 62 pharmaceutical molecules with previously unpublished parameter values.
2
项与 盐酸恩卡尼 相关的新闻(医药)2025-05-22
·赛柏蓝
药物–药物相互作用(DDI)可能会是致命的.....既往教训告诉我们,即使是经过严格研发、被寄予厚望的药物,也可能因为与其他药物的相互作用而导致严重的后果。1997年,某医药公司推出了一款名为西立伐他汀的他汀类药物,这款药物以其良好的降脂效果迅速赢得了市场的青睐,当年就开出了200多万的处方。然而,好景不长。随着西立伐他汀的广泛使用,全球范围内开始陆续出现因服用该药而产生横纹肌溶解所致的死亡报告。据统计,全球600多万患者中,有52例因药物相互作用而丧生。深入调查揭示,几乎所有横纹肌溶解病例都与另一种降胆固醇药物吉非罗齐的联合使用有关。为了追求更快的降脂效果,医生们将这两种药物同时开给患者,却未料到联合使用会导致西立伐他汀血药浓度激增五倍,最终引发致命反应。这场本为治疗疾病的医疗行为,因药物相互作用而变成了夺命的凶手。对此,该公司于2001年8月决定将西立伐他汀退出市场,这一决定既是对逝去生命的悼念,也是对医疗界的一次警示。当下,新冠药物的使用也日益广泛。然而,在使用新冠药物时,我们必须时刻牢记药物相互作用的风险,确保用药安全有效。01药物–药物相互作用是什么?药物-药物相互作用(Drug-Drug Interaction,简称 DDI)是指联合应用两种或两种以上药物时,由于药代动力学(PK)或药效学(PD)的相互影响,导致药物暴露量、药物疗效或不良反应风险发生改变[1]。常见的相互作用结果有协同和拮抗两种。药物协同作用能增加药物的疗效,加快患者的康复速度;而拮抗作用则降低药物疗效,减慢患者康复速度。合理的药物相互作用可以增强疗效或降低药物不良反应,反之可导致疗效降低或毒性增加,还可能发生一些异常反应,干扰治疗,加重病情,甚至死亡。02新冠药物使用中,当心DDI这个隐形杀手在新冠疫情刚开放管控政策初期,特效药市场需求激增,供不应求。其中就包括一款小分子新冠特效药奈玛特韦/利托那韦[2],作用机制属于3CL蛋白酶抑制剂。为什么这款药由两种成分构成?奈玛特韦是这种药物组合中的主要抗病毒成分,角色定位是“战士”。它是一种针对SARS-CoV-2主要蛋白酶Mpro(也称为3C-样蛋白酶,3CLpro)的拟肽类抑制剂。通过抑制这种蛋白酶,奈玛特韦阻止病毒处理多蛋白前体,从而阻断病毒复制。然而,奈玛特韦单独使用时存在一个问题:它在体内的代谢速度较快,导致血浆水平可能不足以达到所需的治疗效果。为了解决这个问题,引入了利托那韦。利托那韦在这种药物组合中扮演辅助角色。它本身并不直接作用于新冠病毒,而是一种肝药酶CYP3A4的抑制剂。CYP3A4是人体内一种重要的药物代谢酶,负责代谢多种药物,包括奈玛特韦。利托那韦通过抑制CYP3A4的活性,减缓了奈玛特韦的代谢速度,从而延长了其在血液中的活性时间,提高了其在体内的药物浓度。奈玛特韦和利托那韦的组合,实现了1+1>2的协同作用。奈玛特韦负责抑制病毒的复制,而利托那韦则确保奈玛特韦在体内保持较高的浓度,增强了整体的抗病毒效果。但是,对于同时服用多种药物的患者来说,奈玛特韦和利托那韦的组合可能会带来一些风险。首先,由于利托那韦是CYP3A4酶的强效抑制剂,它会影响许多依赖该酶进行代谢的药物的清除率。奈玛特韦/利托那韦与这些药物合用时,这些药物的血药浓度可能会显著升高,从而导致毒性反应或不良反应的加剧。此外,奈玛特韦/利托那韦还可能与其他类型的药物产生相互作用。例如,与抗凝药物利伐沙班合用时,可能会增加出血风险;与某些降糖药合用时,可能会增加低血糖风险;与某些镇痛药合用时,可能会增强镇痛效果,导致呼吸抑制等严重不良反应。还有一些药物,会降低奈玛特韦和利托那韦的血药浓度,降低疗效。奈玛特韦/利托那韦说明书中明确罗列了禁止联用的药品:•镇痛剂:哌替啶、吡罗昔康、丙氧芬•抗癌药:Neratinib、Venetoclax•心血管类药物:胺碘酮、苄普地尔、决奈达隆、恩卡尼、氟卡尼、普罗帕酮、奎尼丁、雷诺嗪、辛伐/洛伐他汀•抗感染药物:夫西地酸、伏立康唑、利福布汀、利福平•抗组胺药:阿司咪唑、特非那定•抗精神病/精神安定药/镇静/催眠药:鲁拉西酮、氯氮平、匹莫齐特、喹硫平、氯拉卓酸、地西泮、舒乐安定氟西泮、口服咪达唑仑、三唑仑•麦角衍生物:二氢麦角胺、麦角新碱、麦角胺、甲基麦角新碱•PDE5抑制剂:西地/阿伐/伐地那非•其他:阿呋唑嗪、西沙必利、秋水仙碱、圣约翰草、卡马西平、苯巴比妥、苯妥英、鲁玛卡托/依伐卡托除上述药物外,仍有部分药物与奈玛特韦/利托那韦存在相互作用,合并用药需谨慎。在中国,已获批的新冠治疗药物中,除了奈玛特韦/利托那韦组合外,还包括先诺特韦/利托那韦以及阿泰特韦/利托那韦等药物含有利托那韦。那么,是不是所有新冠治疗药物都无一例外地包含了利托那韦,我们别无他选呢?03阿兹夫定,药物相互作用少除了这些包含利托那韦的药物外,还有其他选择,比如阿兹夫定片。阿兹夫定对CYP450酶系中的多个亚型无诱导或抑制作用,因此它能够与多种临床常见药物类型安全联用,包括降血糖药物、抗凝剂、非阿片类镇痛药、抗组胺药、抗菌药、抗高血压药、心血管药以及抑酸类药等。在与这些通过CYP450酶代谢的药物合并使用时,阿兹夫定不会改变它们的全身暴露量,既不会影响其疗效,也不会增加毒性风险。同时,阿兹夫定片是一种高效且独特的RdRp抑制剂,“标本兼治”抗病毒、提免疫[3,4],为新冠治疗提供了另一种有力武器。具体来讲,阿兹夫定片通过其核苷三磷酸形式精准靶向病毒RdRp,发挥强大的抗病毒作用,这是“标”。“本”是指阿兹夫定及其代谢产物三磷酸盐在人体胸腺和外周血单核细胞中高度富集,这些活性成分在免疫器官进一步激活了关键的免疫细胞——CD4+和CD8+ T细胞,优先清除免疫器官内病毒,促进了宿主免疫系统对新冠病毒的选择性清除能力,强化了人体的免疫防线,实现了从内部对病毒的免疫调节与清除,缩短病程,最大可能避免新冠后遗症的发生。结语在现代医疗实践中,药物治疗是治疗众多疾病不可或缺的一环。然而,随着患者同时使用多种药物的现象日益普遍,药物–药物相互作用的问题也随之凸显。药物–药物相互作用是药物联合使用时可能发生的相互影响,可能导致药物疗效增强、减弱或出现新副作用,甚至危及生命。西立伐他汀与吉非罗齐的联合使用导致的悲剧就是一个典型例子。在新冠治疗中,包含利托那韦的组合药物与其他药物合用不当造成的DDI,不但会导致治疗失败,还可能使药物的毒副反应发生率增高。而阿兹夫定不影响CYP450酶的活性,产生DDI的概率较小,不失为优选药物,值得在临床实践中不断探索和完善其应用。参考文献:1.Sparkes T, Lemonovich TL; AST Infectious Diseases Community of Practice. Interactions between anti-infective agents and immunosuppressants-Guidelines from the American Society of Transplantation Infectious Diseases Community of Practice. Clin Transplant. 2019;33(9):e13510.2.Paxlovid说明书3.Yu, B.; Chang, J. Azvudine (FNC): a promising clinical candidate for COVID-19 treatment. Signal Transduction and Targeted Therapy 2020, 5 (1), 236.4.Zhang JL, et al. Signal Transduct Target Ther. 2021 Dec 6;6(1):414.END声明:此篇文章受众仅为医药、医疗等健康产业专业伙伴,仅供医学药学专业人士使用,不针对普通消费大众。
紧急使用授权临床研究
2023-01-02
小分子抗新型冠状病毒药物老年患者使用指引随着当前疫情的不断变化,防控政策逐渐由“群防群控”转向“防重症、降死亡”的新阶段。奈玛特韦片/利托那韦片和阿兹夫定片这两个用于抗新冠病毒的药物,也一度成为互联网上的热搜词。那么对于感染新冠病毒的老年人,是否需要服用这两种药物,以及都有什么注意事项呢?北京医院药师将为您一一解答。一、什么是小分子新冠“特效药”《新型冠状病毒肺炎诊疗方案(试行第九版)》中推荐的小分子抗病毒药物包括奈玛特韦片/利托那韦片组合包装(即Paxlovid,帕拉维德)和后来补充的阿兹夫定。这两种抗病毒药物早期使用都可能减少重症的发生、缩短病程和排毒的时间。 图1 奈玛特韦/利托那韦片 图2 阿兹夫定片奈玛特韦片/利托那韦片组合包装,于2022年2月11日被我国国家药品监督管理局按照药品特别审批程序,进行应急审评审批,附条件批准进口注册上市,用于治疗成人伴有进展为重症高风险因素的轻至中度新型冠状病毒肺炎(COVID-19)患者。阿兹夫定是我国自主研发的小分子抗病毒药物,用于抗艾滋病病毒治疗。2022年7月25日,国家药监局应急附条件批准了阿兹夫定增加新冠肺炎治疗适应症的注册申请,用于治疗普通型新型冠状病毒肺炎成年患者,是我国第一个拥有完全自主知识产权治疗新冠肺炎小分子口服药物。二、老年新冠病毒感染者是否需要服用“特效药”根据我国《新型冠状病毒肺炎诊疗方案(试行第九版)》,须有以下一种疾病或条件,才定义为重症/危重症高危人群:(1)高龄(≥60岁老年人);(2)有心脑血管疾病(含高血压)、慢性肺部疾病、糖尿病、慢性肝脏疾病、肾脏疾病、肿瘤等基础疾病者;(3)免疫功能缺陷(如艾滋病患者、长期使用糖皮质激素或其他免疫抑制药品导致免疫功能减退状态);(4)肥胖(体重指数≥30);(5)晚期妊娠和围产期女性;(6)重度吸烟者。由此可见,老年人,尤其是患有基础疾病的老年人是新冠重症/危重症的高风险人群,这类人群一旦感染新冠病毒应高度重视,密切观察病情的发展,如若出现疾病加重的情况,建议及时就医对症治疗,以免发展为重症患者。虽然这两种药物对新冠病毒感染具有一定疗效,但并不是新冠的“特效药”,更不是所谓的“神药”。现在一些患者因听信传言造成恐慌情绪,盲目购买甚至囤积这类药物,给自己一种“神药”在手,百毒不侵的错觉,这是万万不可取的。老年人在感染新冠病毒后应首先选择对症治疗,对于确实有可能发展为重症的患者,应及时就医,经专业医师诊断,符合用药标准的才能凭处方购买服用这类药物。三、老年人服用这两种小分子抗病毒药物有哪些注意事项1、需要具有明确的适应症目前这两种药物均为处方药物,需要在医疗机构、在医生的指导下使用。表1 奈玛特韦片/利托那韦片与阿兹夫定的适应症及用法用量奈玛特韦片/利托那韦片阿兹夫定(1)快速抗原诊断检测阳性、或SARS-CoV-2病毒PCR检测阳性;(2)重症高危、≥12岁、≥40kg;高危因素:≥65岁、慢性肺病(如哮喘)、慢性心脏病、慢性肾脏病、慢性肝病、卒中/痴呆、糖尿病/肥胖、镰状细胞贫血/地中海贫血、各种恶性肿瘤、免疫系统受抑制状态、唐氏综合征、精神疾病。(3)轻中度症状。普通型新型冠状病毒肺炎(COVID-19)成年患者。300mg奈玛特韦片与100mg利托那韦片同时服用,每12小时一次。片剂需整片吞服,不得咀嚼、掰开或压碎,疗程不超过5天。空腹整片吞服,每次5mg,每日1次,疗程不超过14天。2、需要结合自己的身体状况选用当老年患者伴有严重的肝、肾功能不全时,应谨慎服用这两种药物。建议患者到医疗机构,由医师或药师对肝肾功能进行评估,调整剂量,个体化的用药才能保证这类患者用药安全。表2 奈玛特韦片/利托那韦片与阿兹夫定在肝肾功能损伤患者中的用法奈玛特韦片/利托那韦片阿兹夫定轻度(Child-Pugh A)或中度(Child-Pugh B)肝功能损害患者不需调整剂量。严重肝功能损害(Child-Pugh C)的患者不建议使用该药。对中重度肾功能损伤(eGFR<70mL/min)者无研究数据,不建议使用。轻度肾功能损伤(60mL/min≤eGFR<90mL/min)无需调整剂量。中度肾功能损伤者(30mL/min≤eGFR<60mL/min)推荐奈玛特韦(1片150mg)和利托那韦(1片100mg),两种片剂一起服用,2次/天,服用5天。严重肾功能损害(eGFR<30mL/min)患者不推荐使用。对中重度(Child-Pugh B)和(Child-Pugh B)肝功能损害患者无研究数据,建议慎用。3、需要警惕药物相互作用当老年患者正在服用其他药物时,应当注意奈玛特韦片/利托那韦片或阿兹夫定与这些药物之间的相互作用。奈玛特韦/利托那韦中的奈玛特韦是抑制病毒的主要成分,但该成分容易被肝脏中的肝药酶CYP3A4快速代谢,导致失去药效。而利托那韦是肝药酶(CYP3A)的抑制剂,可以抑制奈玛特韦的代谢。临床中一部分药物是高度依赖肝药酶CYP3A进行代谢清除的、一部分药物可诱导CYP3A的活性、还有一部分药物可抑制CYP3A的活性。当奈玛特韦/利托那韦与以上几类药物同时使用时,会导致本品或对方药物在体内的浓度产生变化,从而导致不良事件的发生。在此,临床药师对禁止联合使用的药物进行了总结,希望能帮助到更多计划或正在服用奈玛特韦/利托那韦的患者。表3 *禁止与奈玛特韦/利托那韦联用的药品禁止联合使用的药物类别禁止联合使用的具体药物名称相互作用机制及处理方法联合用药导致合用药物的水平升高或降低α1肾上腺素能受体拮抗剂阿呋唑嗪阿呋唑嗪的血药浓度增高,可能导致严重的低血压。镇痛剂哌替啶,吡罗昔康,丙氧芬联合使用会使合用药物的血药浓度增高,从而增加严重呼吸抑制、血液系统异常或这类药物所致的其它严重不良反应的发生风险。抗心绞痛药雷诺嗪雷诺嗪的血浆浓度升高,可能使发生严重和/或危及生命的不良反应的可能性增加,禁止联用。抗心律失常药胺碘酮,苄普地尔,决奈达隆、恩卡尼,氟卡尼,普罗帕酮,奎尼丁,多非利特联合使用会使合用药物的血药浓度增高,从而增加心律失常或这类药物所致的其它严重不良反应。可选择索他洛尔,联合使用时相对安全。抗心衰药依普利酮,伊伐布雷定禁止联合使用。抗凝/抗血小板药利伐沙班,阿哌沙班,替格瑞洛,氯吡格雷联合使用会大幅度降低抗凝/抗血小板药物的效果,增加栓塞风险。阿哌沙班应在奈玛特韦/利托那韦停药3天后回复使用。抗生素夫西地酸夫西地酸和利托那韦的血药浓度增高。抗真菌药伏立康唑利托那韦(400mg,一日两次和更多次)与伏立康唑禁止合用。合用会降低伏立康唑血药浓度,并可能导致失效。抗组胺药阿斯咪唑,特非那定合用导致阿斯咪唑和特非那定的血药浓度增高,从而增加了这些药物所致严重心律失常的发生风险。抗痛风药秋水仙碱对于有肝损伤、肾损伤患者具有严重不良反应或危及生命的潜在风险。抗分枝杆菌药利福布汀利托那韦作为抗反转录病毒药(600mg,一日两次)和利福布汀合用会增加利福布汀的血清浓度和不良反应的发生风险。支气管扩张剂β2受体激动剂沙美特罗联用增加心脏毒性,不宜联用。抗精神病药/精神安定药鲁拉西酮鲁拉西酮的血浆浓度升高,可能使发生严重和/或危及生命的不良反应的可能性增加。氯氮平,匹莫齐特氯氮平和匹莫齐特的血药浓度增高,从而增加了严重血液学异常或这类药物所致的其它严重不良反应的发生风险。喹硫平喹硫平血药浓度增高,从而导致昏迷。禁止与喹硫平联合用药。麦角衍生物二氢麦角胺,麦角新碱,麦角胺,甲基麦角新碱麦角衍生物血药浓度的增高会导致急性麦角碱毒性,包括血管痉挛和缺血。胃肠动力药西沙必利西沙必利的血药浓度增高,将增加该药所致严重心律失常的发生风险。血脂调节剂HMG-CoA还原酶抑制剂洛伐他汀,辛伐他汀,阿托伐他汀,瑞舒伐他汀导致部分他汀类药物的血药浓度升高;因此增加了包括横纹肌溶解在内的肌病的发生风险升高。建议在服用奈玛特韦/利托那韦期间停用左侧四种他汀类,或换用普伐他汀或匹伐他汀代替。PDE5抑制剂阿伐那非合用使阿伐那非的血药浓度升高。西地那非当作为治疗肺动脉高压药物时禁用,合用会使西地那非的血药浓度增高。会增加潜在的西地那非相关不良事件的发生风险。伐地那非合用会使伐地那非的血药浓度升高。镇静/催眠药地西泮,舒乐安定,氟西泮,口服咪达唑仑和三唑仑会使合用药物的血药浓度增高,从而增加这些药物所致过度镇静和呼吸抑制的风险。阿片类药物羟考酮,哌替啶联合使用会增加阿片类药物副作用。免疫抑制剂环孢素,他克莫司,西罗莫司禁止联合使用。肺动脉高压治疗药物波生坦,马西替坦,西地那非,他达拉非禁止联合使用。波生坦为CYP3A底物,联合使用会使波生坦浓度升高。在开始使用奈玛特韦/利托那韦之前至少停用波生坦 ≥36小时。联合用药导致奈玛特韦/利托那韦水平降低中草药制剂圣约翰草(贯叶连翘)由于有降低利托那韦的血药浓度和临床疗效的风险,禁止与含有圣约翰草(贯叶连翘)的中草药制剂合用抗惊厥药卡马西平,苯巴比妥,苯妥英与单独给药相比,联合给药有降低奈玛特韦血药浓度的风险抗感染药利福平利福平是强CYP3A4诱导剂,这可能导致奈玛特韦/利托那韦的暴露减少和病毒学反应的潜在丧失。*不限于上述品种,具体情况请咨询药师或医师目前已知富马酸替诺福韦酯和依非韦伦与阿兹夫定合用可能影响其药代动力学,但对于其他药物与阿兹夫定的相互作用因临床资料较少尚不清楚。如果老年患者正在服用其他药物,建议用药前咨询医师。四、老年人服用奈玛特韦片/利托那韦片的常见问题本着借鉴国外经验、提高对病毒和抗病毒药物了解的目的,近期我国学者邀请到了哈佛大学学者进行交流。会议中外国学者分享了“老年人服用奈玛特韦片/利托那韦片的常见问题”。在此,药师帮助您进行了总结,希望可以帮助到更多60岁以上的老年人。问题1:老人高血脂在服用他汀类药物,怎么使用奈玛特韦片/利托那韦?回答1:奈玛特韦片/利托那韦片可以影响肝功能和酶的代谢,有时增加酶代谢,有时减少酶代谢。比较常见的是增加药物在身体内的时间,延长药物的半衰期。比如我们在服用氨氯地平、硝苯地平或他汀类药物的时候,如果继续按照以前的剂量来吃,同时服用奈玛特韦片/利托那韦片可能带来副作用。所以如果患者正在用他汀类的药物,建议先停止用药,降血脂药停用5天影响不大;如果是降血压药物,则可减少一半剂量。问题2:老人在用控制血糖的药和抗凝药,怎么同时使用奈玛特韦片/利托那韦片?回答2:稀释血液的药物、抗凝药物都需要持续使用,不能让患者停药。这类患者同时使用奈玛特韦片/利托那韦片没有办法完全规避风险,需要平衡风险和好处,同时密切关注。比如患者可能有更高的出血风险,需提前告知患者避免做一些可能导致出血的动作,包括刷牙。如果患者之前服用的是高剂量抗凝药,则需要减少一半剂量。问题3:老人阳了,假设没有其他基础病,什么时候服用奈玛特韦片/利托那韦片更好?回答3:在美国,患者什么时候开始服用奈玛特韦片/利托那韦片遵循三个要求:①新冠检测阳性;②年龄超过12岁且出现症状5天内;③有一个或多个重症风险因素。其中,年龄是主要的重症风险因素,超过60岁就属于有重症风险,使用奈玛特韦片/利托那韦片可以避免80%-90%患者住院或出现重症,只要患者肾功能中度到正常,没有肝脏问题就可以使用。问题4:具体怎么评估老人的肝肾功能是否正常呢?患者服用了是否也要监测?具体监测哪几个指标?回答4:肾功能的评估标准是全世界通用的:一个是肌酐,一个是肾小球滤过率。肾小球滤过率如果超过60毫升/分钟,就给全剂量奈玛特韦片/利托那韦片;如果在30-60毫升/分钟,就给一半剂量;低于30毫升/分钟通常不建议给药。但针对15-30毫升/分钟的患者,如果没做透析,有的医生也会给更小剂量治疗。肝功能的评估标准一般是转氨酶、白蛋白、胆红素。如果这些指标超过正常值1.5-2倍,就需要减少一半奈玛特韦片/利托那韦片的剂量。因为奈玛特韦片/利托那韦片在体内的代谢会延缓肝部代谢的功能,和其他药物产生互相影响。问题5:帕金森老人正在服用美多芭和雷沙吉兰,能同时使用奈玛特韦片/利托那韦吗?要不要停药?回答5:帕金森是运动障碍疾病,如果停药,会导致患者震颤发作,吞咽有问题,从而增加肺部感染的风险。所以这类患者在使用奈玛特韦片/利托那韦时,会继续服用帕金森药物来减少震颤发生,同时密切关注患者症状。问题6:国内老人新冠后,容易并发肺炎。比如一位90岁老人新冠5天了,有肺部感染,国内医生建议使用奈玛特韦片/利托那韦片,您怎么看呢?回答6:我们一般会在患者阳性后的5天内给药。因为5天内是病毒复制的高峰时段,5天后病毒复制明显下降,开始进入免疫应答期,这个时候才使用奈玛特韦片/利托那韦片,效果就会下降。但考虑到患者90岁老人有较高的重症风险,5天后还是可以继续给药,但效果肯定不如5天内开始给药。问题7:用抗生素与Paxlovid治疗新冠,需要有用药间隔吗?回答7:奈玛特韦片/利托那韦片和很多抗生素(如阿奇霉素、阿莫西林、左氧类药物)都没有相互作,可以正常使用;和利福平有相互作用,需要避免使用。患者刚得新冠的5-7天内,是病毒复制的高峰期,5-7天后病毒不再很快复制了,但免疫系统还有个应答期,容易发生肺炎等其他并发症,这些并发症会再持续7-10天。在免疫应答期可以应用免疫抑制类药物,如激素类的强的松、可的松等帮助抑制免疫反应。问题8:老人阳了5天、7天后才找到奈玛特韦片/利托那韦片,这时用药还有意义吗?回答8:如果患者仅有一些很轻的症状,5-7天后病毒已经不再复制了,就不需要再用奈玛特韦片/利托那韦;如果患者出现了比较重的症状,就需要到医院用其他药物来缓解症状。不管患者5-7天后体内还有没有病毒,抗原检测都可能是阳性的,更多的是临床症状观察,包括做血液检测等,而不是做新冠检测来指导治疗。问题9:老年人阳了,但一直没有症状,需不需要服用奈玛特韦片/利托那韦片?回答9:当患者新冠检测阳性,超过65岁,或者没打疫苗,或者有任何一种慢性病,都应该在5天内开始使用奈玛特韦片/利托那韦片,不管有没有症状。问题10:糖尿病患者平时在用控糖的药,感染新冠后使用奈玛特韦片/利托那韦有没有影响?回答10:糖尿病患者不需要停药或减量。但需要注意,降糖类药物可能导致患者出现腹泻,从而引起脱水。在同时用药时,患者需要大量饮水,防止脱水状况的出现。问题11:布洛芬、对乙酰氨基酚,这两种退烧药和奈玛特韦片/利托那韦有冲突吗?回答11:感染新冠后,给患者造成不舒服的主要原因就是脱水,而脱水是发烧和腹泻产生的。这两种退烧药中,对乙酰氨基酚的半衰期更短一些,布洛芬的半衰期更长一些,作用是相似的,没有那种药物更好。我个人比较习惯用泰诺(对乙酰氨基酚),不会刺激肠胃;布洛芬会刺激肠胃。此外,布洛芬是从肾脏代谢的,有肾脏基础病的可以用泰诺替代;而泰诺是肝脏代谢的,有肝脏基础病的可以用布洛芬替代。问题12:发烧两天了,第三天才阳性。是应该一有症状就用奈玛特韦片/利托那韦,还是等阳性再用?回答12:主要看患者是否与新冠患者有密切接触史,如果明确患者本人有比较高的感染风险,那么可以给患者奈玛特韦片/利托那韦来预防。这个药不会给患者带来伤害,而是给他足够的防护。新冠检测也只有75%-80%的准确率,检测也有可能是假的结果。问题13:如果患者指标支持继续用奈玛特韦片/利托那韦,但是新冠症状已经没有了,可以提前停药吗?回答13:建议继续用药。新冠反弹也是常见的情况,如果停药早了,患者可能出现反弹,所以一定要把5天的药吃完。问题14:肿瘤患者化疗期间,能不能使用奈玛特韦片/利托那韦片?回答14:化疗药的种类非常多,代谢机制也不一样。但确实有的化疗药是通过肝脏代谢的,所以需要和肿瘤医生沟通,会不会和新冠药物发生冲突。问题15:有些患者由于身体原因,不能正常口服药物,能不能鼻饲使用奈玛特韦片/利托那韦片?回答15:对于无法吞咽的患者来说,可以鼻饲给药。具体做法是把药物捣碎之后,和苹果酱或者酸奶一起来服用。问题16:第一次感染新冠用了奈玛特韦片/利托那韦片,第二次感染还能使用吗?回答16:可以多次使用。这取决于患者感染的原因:一种可能是之前的感染还没好,可以一直用;还有一种可能是患者感染后有了抗体,几个月内都不会得新冠,除非患者的免疫系统是不健全的,二次感染后,还可以重复使用奈玛特韦片/利托那韦片。最后老年人在选择抗新冠病毒药品时,不但要考虑其治疗属性,还需要关注安全性。以上内容仅简要介绍了应用这类药物的注意事项,对于如何安全用药、合理用药,大多数普通民众仅参考这些信息是难以把握的。此外,很多人在网上购买奈玛特韦片/利托那韦片的仿制药,由于仿制药物未获批在中国境内上市,其质量无法得到国家监管,不建议擅自购买使用。因此,抗新冠病毒药物作为处方药,应经医生或药师的专业评估后,在正规医疗机构获得,切勿擅自用药,以免造成不良的后果。撰稿:那一凡、赵紫楠指导:胡欣审核:金鹏飞、张亚同2022年12月30日来源:北京医院药学部文章信息源于北京医院药学部,登载该文章目的为更广泛的传递行业信息,不代表赞同其观点或对其真实性负责。文章版权归原作者及原出处所有,文章内容仅供参考。本网拥有对此声明的最终解释权,若无意侵犯版权,请联系小编删除。学如逆水行舟,不进则退;心似平原走马,易放难收。行舟Drug每日更新 欢迎订阅+医药大数据|行业动态|政策解读
100 项与 盐酸恩卡尼 相关的药物交易
登录后查看更多信息
研发状态
10 条最早获批的记录, 后查看更多信息
登录
| 适应症 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|
| 室性心动过速 | 美国 | 1986-12-24 | |
| 室性早搏 | 美国 | 1986-12-24 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
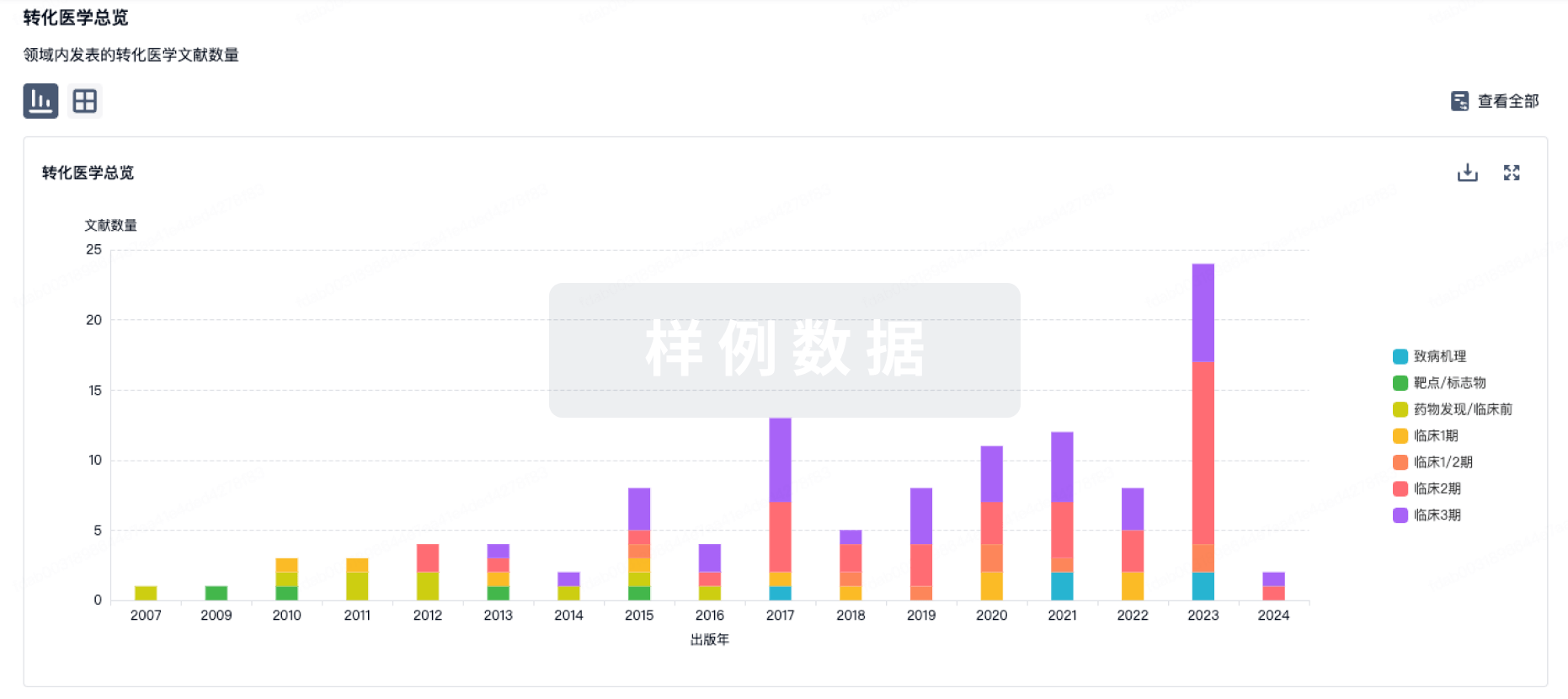
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
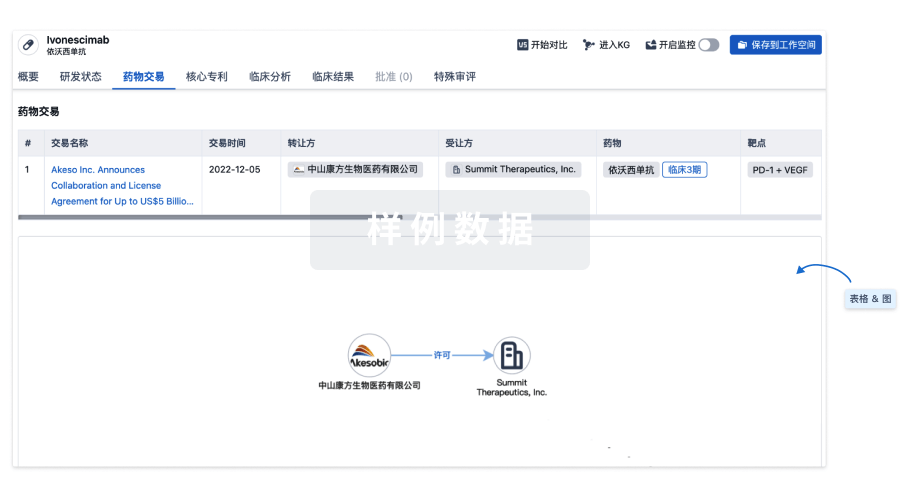
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
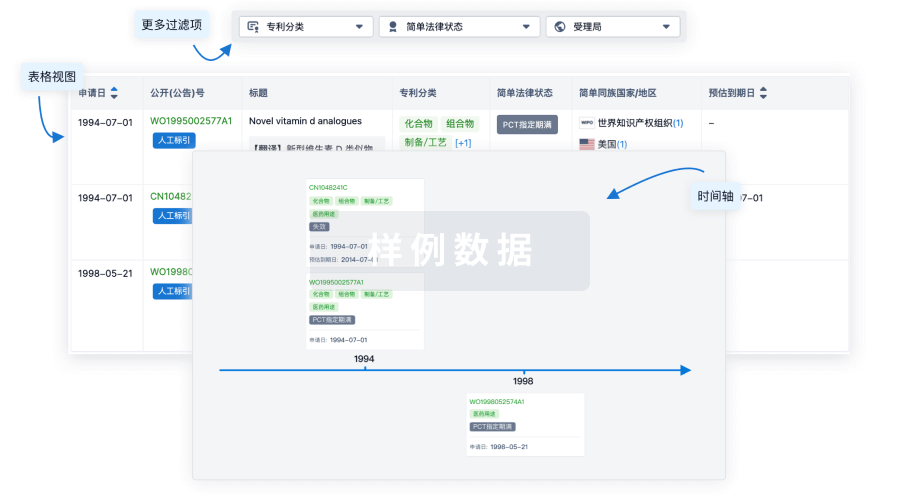
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
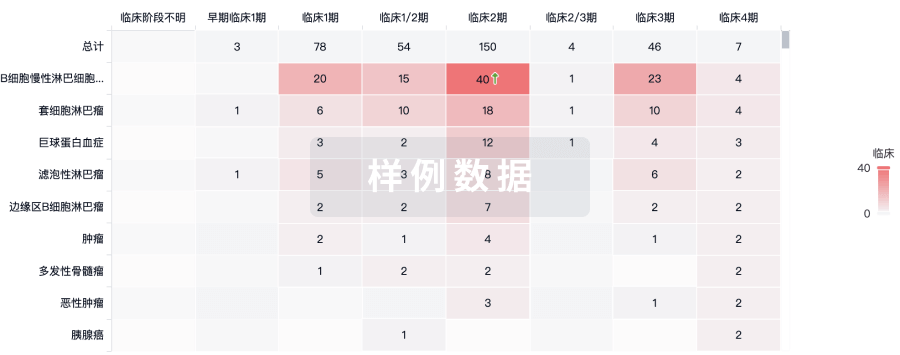
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
批准
利用最新的监管批准信息加速您的研究。
登录
或
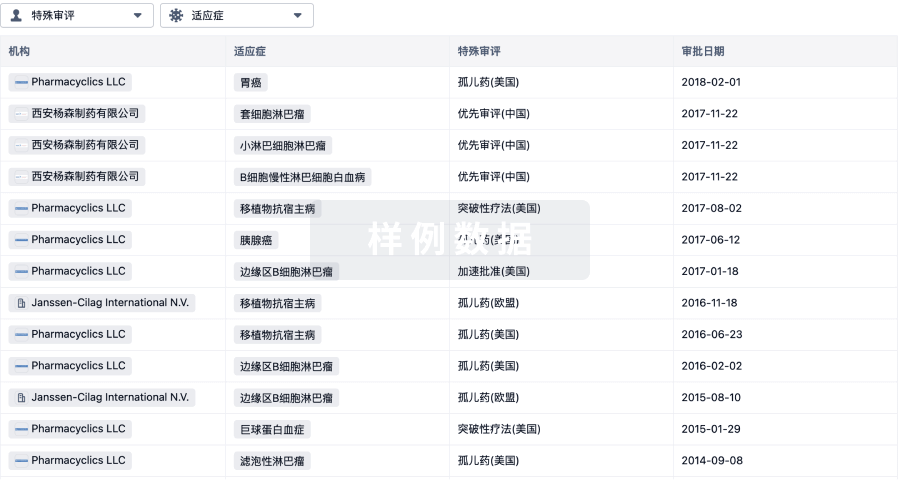
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
生物医药百科问答
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
