预约演示
更新于:2025-07-23
Beclabuvir hydrochloride
贝卡布韦
更新于:2025-07-23
概要
基本信息
药物类型 小分子化药 |
别名 Beclabuvir、Beclabuvir hydrochloride (JAN/USAN)、BMS-791325 + [1] |
作用方式 抑制剂 |
作用机制 NS5B polymerase抑制剂(丙型肝炎病毒NS5B RNA依赖的RNA聚合酶抑制剂) |
在研适应症- |
非在研适应症 |
在研机构- |
权益机构- |
最高研发阶段终止临床3期 |
首次获批日期- |
最高研发阶段(中国)终止 |
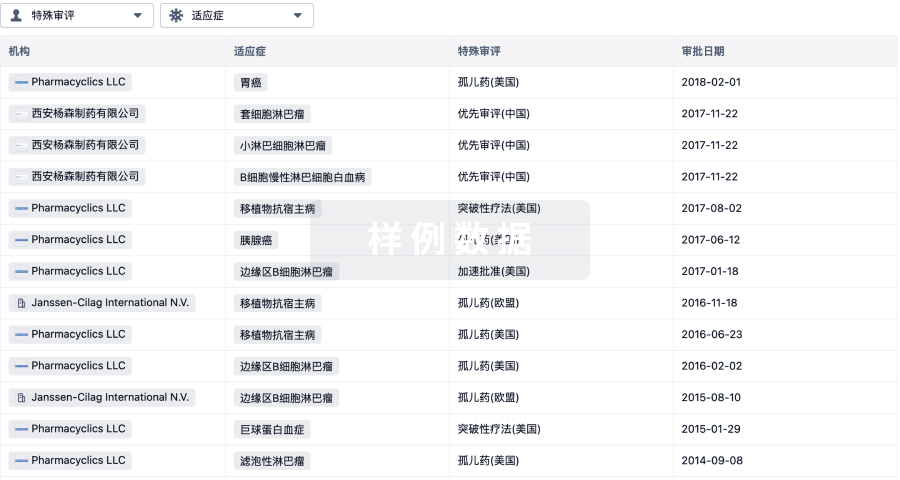
特殊审评- |
登录后查看时间轴
结构/序列
分子式C36H46ClN5O5S |
InChIKeyIHXVACFNNPBRLK-VTFHPOPBSA-N |
CAS号958002-36-3 |
关联
17
项与 贝卡布韦 相关的临床试验NCT03071133
Asunaprevir/Daclatasvir/Beclabuvir Fixed-Dose Combination Safety Surveillance in Japanese Patients With Chronic Hepatitis C (HCV) or Japanese Patients With Compensated Cirrhosis
An observational, postmarketing commitment following the marketing authorization for DCV Trio therapy in Japan
开始日期2018-01-01 |
申办/合作机构 |
JPRN-UMIN000026917
Daclatasvir/Asunaprevir/Beclabuvir regulates innate immunity by inhibition of nonstructural 5B and suppresses liver fibrosis development - Daclatasvir/Asunaprevir/Beclabuvir regulates innate immunity by inhibition of nonstructural 5B and suppresses liver fibrosis development
开始日期2017-09-01 |
申办/合作机构- |
JPRN-UMIN000026909
Evaluation of the efficacy of daclatasvir+asunaprevir+beclabuvir for Japanese hepatitis C patients; a prospective study in real life settings. - The efficacy of daclatasvir+asunaprevir+beclabuvir for Japanese hepatitis C patients.
开始日期2017-03-01 |
申办/合作机构- |
100 项与 贝卡布韦 相关的临床结果
登录后查看更多信息
100 项与 贝卡布韦 相关的转化医学
登录后查看更多信息
100 项与 贝卡布韦 相关的专利(医药)
登录后查看更多信息
134
项与 贝卡布韦 相关的文献(医药)2025-03-24·Journal of biomolecular structure & dynamics
Inhibition of respiratory syncytial virus by Daclatasvir and its derivatives: synthesis of computational derivatives as a new drug development
Article
作者: Mitra, Debanjan ; Abdalla, Mohnad ; Afreen, Shagufta ; Das Mohapatra, Pradeep K.
The most common cause of respiratory tract illness in newborns and young children is the respiratory syncytial virus (RSV). There is no approved vaccination or specific antiviral medication for RSV infections. Here, an attempt has been made to explore the potential of currently marketed drugs as well as their probable derivatives to improve the possibility of developing stronger medications against RSV. From the 100 synthetic drug compounds library, the best drug molecule was identified through drug-likeness properties, toxicity, molecular docking and molecular dynamics simulations. Molecular Mechanics Generalized Born Surface Area (MM-GBSA) was also a method that was applied in this study. Daclatasvir showed the highest binding energy and appeared as the best drug to inhibit matrix protein and a fusion protein of RSV. Based on Daclatasvir, 40 computational derivatives were made. D28, D34 and D40 showed far better results than the actual drug. Changes in lipophilicity character increase the binding energy of derivatives. Molecular dynamic simulations showed their non-deviated, non-fluctuated and stable complex formation with target proteins. The high number of amino acid contacts throughout the trajectory increases the stability and effectiveness of derivatives. The key to producing a novel medicine to eradicate RSV is provided by derivatives. Daclatasvir will be employed as a potential RSV inhibitor up until that point.
2024-09-18·ChemistrySelect
Computational Screening of FDA‐Approved Hepatitis C Drugs for Inhibition of VEGFR2 in Liver Cancer
作者: Wilhelm, Anke ; Tufail, Nasir ; Aluwi, Mohd Fadhlizil Fasihi Mohd ; Issahaku, Abdul Rashid ; Roney, Miah
Abstract:
Liver cancer (LC) is one of the most common tumours and the leading cause of cancer‐related death globally. Amidst the problems associated with existing treatments, such as hepatotoxicity, recurrence, drug resistance, and other adverse effects, researchers are under pressure to find alternatives. Towards a comprehensive rationalisation of the search for new anti‐LC drugs among approved ones, we employed an in‐silico approach to accelerate the selection of the most efficacious LC drugs. The FDA‐approved hepatitis C virus (HCV) drugs were docked with the LC protein using the AutoDock Vina software. Compared to the control compound, two FDA‐approved HCV drugs (DB09102 and DB09027) were selected based on their binding energies and interactions with the target protein, which showed comparable binding energies. Furthermore, these compounds were then subjected to molecular dynamic simulation, principle component analysis, and MMGBSA using the AMBER20 software, and the results showed stable complexes compared to the control complex. All things considered, this study will help the scientific community and society find a novel drug to treat LC.
2024-08-01·REGULATORY TOXICOLOGY AND PHARMACOLOGY
Evaluation of the EMA log kow trigger for fish BCF testing based on data for several human pharmaceuticals
Article
作者: Nfon, Erick ; Tell, Joan ; Davidson, Todd ; Dolan, David G ; Nimrod Perkins, Alison ; Ryan, James J ; Häner, Andreas ; Janer, Gemma ; Constantine, Lisa A ; Burden, Natalie ; Maynard, Samuel K ; Lee, Michael R
In the European Medicines Agency (EMA) "Guideline for Environmental Risk Assessment of Medicinal Products for Human Use," a fish bioconcentration factor (BCF) study is triggered in Phase I for pharmaceuticals having log Kow >4.5, to support Persistence, Bioaccumulation and Toxicity (PBT) screening, and in Phase II to assess secondary poisoning and bioaccumulation ('B') potential when log Kow ≥3. The standard sampling schedule outlined in OECD Test Guideline 305 (TG305) may require assessment of approximately 200 fish following exposure to low- and high-test concentrations and a negative control. We report experimental log Kow and BCF values for 64 human pharmaceuticals that were used to evaluate the current BCF testing trigger of log Kow ≥3, and whether a single BCF exposure concentration allows accurate classification of bioaccumulation potential. Our data support raising the BCF testing trigger to log Kow ≥4, and use of a single test concentration. The resulting reduction in the use of fish is consistent with the 3 R s principle and did not adversely affect classification accuracy. An assessment of potential risk of secondary poisoning was also conducted for three drugs classified as either B or vB, and no risks were identified.
2
项与 贝卡布韦 相关的新闻(医药)2025-02-11
关注并星标CPHI制药在线
近日,东阳光药业1类创新药磷酸萘坦司韦胶囊获NMPA批准上市,用于与艾考磷布韦片联用,治疗初治或干扰素经治的基因1、2、3、6型成人慢性丙型肝炎病毒(HCV)感染,可合并或不合并代偿性肝硬化。
磷酸萘坦司韦是一款NS5A抑制剂,艾考磷布韦是一款NS5B抑制剂,两款药均为东阳光药自主研发的1类新药。该联合疗法治疗慢性丙肝的2期临床研究显示,该方案在基因1型、2型、3型和6型受试者中SVR12(治疗结束后12周时实现持续病毒学应答的受试者百分比)分别为96.2%、100.0%、83.3%、100.0%。代偿期肝硬化和无肝硬化受试者总体SVR12分别为100%和96.2%。全分析集分析结果显示总体SVR12为96.7%。
以此出发,我们回顾一下丙肝药物开发的重要时刻。
干扰素治疗丙肝的困境
HCV于1989年被发现,是一种常见的血源性病原体,主要感染部位是肝脏,一经感染,如果不采取进一步治疗,HCV可能引发急性或慢性肝炎,并增加感染者发展为肝硬化、肝功能衰竭和肝癌的风险。HCV共存在6种基因型(1-6),丙肝患者以基因1型居多,占大约70%。
2020年,三位科学家因发现了HCV获得诺贝尔生理学或医学奖。当年诺奖颁发时,已经有了很多治疗HCV的药物。而在此之前,丙肝的治疗主要采用聚乙二醇干扰素(一种具有抗病毒、抗肿瘤和免疫调节特性的多效性细胞因子)联合利巴韦林(一种广谱抗病毒药物)。
其方案为连续在24至48周内注射聚乙二醇干扰素和服用利巴韦林,但这种疗法不仅费用昂贵且毒副作用强,例如干扰素会引起流感样症状,利巴韦林则可能引起溶血性贫血。在许多情况下,严重的副作用导致患者停止治疗。
在疗效方面,根据《丙型肝炎防治指南》,“干扰素+利巴韦林”治愈率大概只有54%-82%,且治愈者有不小的几率会复发。
20世纪末,随着HCV体外培养难题被攻克,新的机会出现了。科学家们终于有了能用来筛选丙肝药物的工具,这为开发直接抗病毒药物(DAAs)奠定了基础。
NS3/4A蛋白酶抑制剂
宣布HCV治疗进入DAA时代
研究发现,HCV是一条单链正义RNA病毒,基因组包含10个基因,编码膜蛋白E1和E2,核心蛋白、p7以及非结构蛋白NS2、NS3、NS4A、NS4B、NS5A和NS5B。其中NS3/4A、NS5A、NS5B是HCV复制过程中三个重要的作用靶点
DAAs的作用靶点(图片来源:参考来源2)
针对这三个靶点,科学家们研发了HCV特异的直接抗病毒药物:NS3/4A 蛋白酶抑制剂、NS5A 抑制剂及NS5B 聚合酶抑制剂。
NS3/4A蛋白酶抑制剂是最早上市的丙肝特异的DAA。NS3/4A复合体主要催化丙肝病毒非结构蛋白水解成熟,是丙肝病毒生活周期所必须的。
2011年,第一个NS3/4A蛋白酶抑制剂Boceprevir(波西普韦)获FDA批准上市,用于与聚乙二醇干扰素和利巴韦林联合治疗HCV患者,该药由默沙东开发。后续分别上市了Vertex/强生的Incivek(特拉瑞韦,已退市)、强生的Olysio(西美瑞韦,Simeprevir),以及BMS在日本上市的Sunvepra(asunaprevir)。
NS3/4A蛋白酶抑制剂的出现,使“干扰素注射+利巴韦林+DAA”成为丙肝临床标准治疗方案,并将临床治愈率从40%提高到了80%以上。
NS5B 聚合酶抑制剂
使口服治愈丙肝成为可能
NS5B是HCV的RNA聚合酶,抑制它将会直接抑制RNA合成,从而终止病毒复制。而且在不同基因型的HCV中,NS5B 聚合酶均高度保守,且抑制它不会产生宿主毒性,因此,NS5B成为一个理想的抗HCV病毒靶点。
尽管从蛋白编号上来看,NS5B似乎是在NS5A之后。但是从成药速度而言,NS5B抑制剂却早于NS5A抑制剂出现。
2013年,吉利德的小分子药NS5B抑制剂索非布韦(sofosbuvir)获FDA批准上市。索非布韦最初由Pharmasset公司开发,后该公司被吉利德收购,索非布韦被成功纳入麾下。索非布韦对多个基因型HCV感染均疗效惊人,直接把疾病治愈率提高到近乎100%,而且索非布韦是首个获批可用于慢性丙型肝炎全口服治疗方案的药物,在用于特定基因型慢丙肝治疗时,可消除对干扰素的需求。
2014年,索非布韦上市的第一年,销售收入就达102.83亿美元,创下新药上市首年销售突破100亿美元的记录。
市场上另外两种NS5B抑制剂分别是艾伯维的达拉他韦(Viekira Pak,2014年)和BMS的贝卡布韦(Ximency,2016年,仅在日本获批),它们都是复方药物的成分。
NS5A 抑制剂,复合DAA方案中的骨干
与HCV基因组中的许多其他蛋白质不同,NS5A (非结构蛋白5A)实际上没有酶的活性,但它却是HCV RNA复制和病毒粒子组装所必需的,被称为HCV生命周期的主调节器。
2014年,BMS的达卡他韦(Daclatasvir)在日本获批上市,用于治疗基因1型慢丙肝、丙肝代偿性肝硬化和病毒血症。达卡他韦通过结合NS5A蛋白N末端二聚体,抑制NS5A从而抑制HCV复制。达卡他韦上市后,多个NS5A抑制剂陆续推出。
在慢丙肝治疗中,由于单一靶点药物不足以满足全口服、短疗程、多基因型有效、高治愈率的要求,因此一般采用DAA联合治疗方案。目前所有多靶点组合DAA方案中,都含有NS5A抑制剂作为骨干。与之配伍的可以是一个NS3/4A蛋白酶抑制剂(如elbasvir/grazoprevir)、一个NS5B聚合酶抑制剂(如sofosbuvir/ledipasvir)或者两种均有(如sofosbuvir/velpatasvir/voxilaprevir)。这些方案都达到了95%甚至更高的治愈率。
自HCV被发现以来,政府机构以及制药行业均投入大量资源,发现了许多有效的抗丙肝药物。现在可以通过持续病毒学应答(SVR)来实现治愈。由于丙肝的高治愈率,用药患者人数在大幅减少,在患者群体日益萎缩的情况下,丙肝药物市场正在大幅缩水。该赛道相关玩家,早已开始转型。
参考来源:
1.https://www.nmpa.gov.cn/zhuanti/cxylqx/cxypxx/20250208172031130.html.
2.Palese P. Profile of Charles M. Rice, Ralf F. W. Bartenschlager, and Michael J. Sofia, 2016 Lasker-DeBakey Clinical Medical Research Awardees. Proc Natl Acad Sci U S A. 2016 Dec 6;113(49):13934-13937. doi: 10.1073/pnas.1616592113. Epub 2016 Nov 18. PMID: 27864510; PMCID: PMC5150379.
3.Tiffany Benzine, Ryan Brandt, William C. Lovell et al. NS5A inhibitors unmask differences in functional replicase complex half-life between different hepatitis C virus strains. PLoS Pathogens, Published: June 8, 2017, doi:10.1371/journal.ppat.1006343.
扫码领取CPHI & PMEC China 2025展会门票
【智药研习社近期直播】
来源:CPHI制药在线
声明:本文仅代表作者观点,并不代表制药在线立场。本网站内容仅出于传递更多信息之目的。如需转载,请务必注明文章来源和作者。
投稿邮箱:Kelly.Xiao@imsinoexpo.com
▼更多制药资讯,请关注CPHI制药在线▼
点击阅读原文,进入智药研习社~
上市批准申请上市
2023-02-27
关注并星标CPHI制药在线近日,全国各地托幼机构、中小学陆续开学,由诺如病毒感染导致的病毒性胃肠炎进入高发期,而话题"广东诺如病毒进入高发期"一度冲上微博热搜榜首,引发网友热议。 诺如病毒 (norovirus,NoV) 亦称诺瓦克病毒,是引起全球散发性胃肠炎和暴发性急性胃肠炎的主要病原体之一。NoV 发生感染时,可以在小肠上皮细胞内复制,上皮细胞分泌功能受损,小肠消化吸收面积减小,引起消化不良和腹泻。经漏泵机制作用,水和离子由上皮下毛细血管扩散至肠腔内,引起腹泻的发生。此外,NoV 感染机体后,B细胞介导体液免疫反应,同时发生由Th1细胞介导的细胞免疫反应。由于人体对不同基因型的NoV 没有完善的交叉保护作用,所以 NoV 感染后,机体无法形成永 久性免疫,再加上 NoV 的变异或重组发生,会使得 NoV 反复感染机体。 迄今为止,还没有批准用于治疗用途的抗NoV药物或疫苗,唯一完成临床试验的候选药物是硝唑尼特 (nitazoxanide,NTZ)。NTZ能够抑制多种RNA和DNA病毒,包括呼吸道合胞病毒 (RSV)、冠状病毒、轮状病毒 (RV)、乙型肝炎病毒 (HBV)、丙型肝炎病毒(HCV)、登革热病毒 (DENV) 和人类免疫缺陷病毒(HIV) 等,目前FDA批准用于治疗贾第虫 (Giardia)和隐孢子虫 (Crytosporidium)的感染。NTZ已被证实在治疗NoV胃肠炎中有一定疗效,它能够抑制Norwalk复制子。 抗NoV相关靶点药物目前,抗NoV药物的设计主要集中在阻断病毒复制的各个环节,如作用于受体细胞膜上的病毒吸附和侵入抑制剂、基于蛋白酶结合口袋设计的蛋白酶抑制剂以及阻断RNA合成的聚合酶抑制剂。此外,由于免疫调节因子具有提高宿主免疫力以抵御病毒感染的能力,因此也是一类潜在的抗病毒药物。 01吸附和侵入抑制剂NoV无包膜,其附着在细胞表面并与宿主表面的受体识别、结合是进入宿主细胞的第一步,也是病毒发挥感染的限速步骤。根据NoV通过组织血型抗原(HBGAs)受体吸附侵袭和感染宿主这一原理,可以设计阻断病毒和HBGAs相互作用的抗NoV药物。基于酶联免疫吸附测定法,发现人乳寡糖中的2'-岩藻糖基乳糖和3'-岩藻糖基乳糖在浓度 5.5~30.2 mmol·L-1下能够抑制 GII.10 NoV VLPs 与 HBGAs 的结合。随后,2'-岩藻糖基乳糖被证实还可以抑制GII.17和GI.1 VLPs与HBGAs的结合, IC50值分别是13~20和38~50 mmol·L-1。以类似的方式,柠檬酸也被证明可以与NoV GII.10 VLPs结合导致其形态结构的改变,从而阻止VLP-HBGA结合。 单克隆抗体或纳米抗体的被动免疫疗法是另一种预防NoV附着和侵入的抗病毒策略。在黑猩猩模型中,单克隆抗体D8能够阻止Norwalk感染。但到目前为止,产生的单克隆抗体的交叉基因型活性有限,这使得它们对抗原多样的NoV的抗病毒效果不佳。而纳米抗体能够通过与衣壳P结构域相互作用而对VLPs产生广泛的中和活性,且它对特定抗原具有高度亲和力,其中很好的NoV纳米抗体是Nb-85,它可阻止GII.1、GII.2、GII.4、GII.12、GII.17和G1.11 VLPs与HBGAs的结合。 02聚合酶抑制剂RNA依赖性RNA聚合酶 (RdRp) 对病毒的复制至关重要,是极具吸引力的抗病毒靶点。靶向RdRp的抗病毒药物一般分为核苷类似物与非核苷类抑制剂。核苷类似物 (NAs) 作为天然底物的类似物,通过发挥竞争性抑制作用和链终止作用来抑制RNA合成从而发挥抗病毒作用。由于NAs结合在高度保守的RdRp 活性位点,所以与非核苷抑制剂相比,它们通常表现出广谱抗病毒活性。利巴韦林(Ribavirin)是首批在体外和细胞水平上被鉴定为具有 NoV 抑制活性的小分子抑制剂之一。研究报道,利巴韦林在HG23细胞中可有效抑制NoV RNA的复制及蛋白的表达,EC50为40μmol/L,并且I型干扰素和利巴韦林联合运用表现出对NoV复制的累加抑制效应。此外,利巴韦林已被证实可有效降低由常见变异型免疫缺陷诱发的NoV感染。法匹拉韦(favipiravir,T-705)最初是针对流感病毒而开发,后来被证明可作为其他 RNA 病毒的小分子抑制剂。体外实验证明,T-705对人诺如病毒具有比利巴韦林更有效的抗病毒活性,且抗病毒作用温和。2'-C-甲基胞苷(2CM-C)以及前药伐洛他滨( NM283),最初是针对丙型肝炎病毒(HCV)聚合酶而开发的,之后发现2CM-C可以抑制鼠诺如病毒的复制。此外,近些年也报导了一些新型的NoV聚合酶抑制剂,如CMX521、CID-57930781、CID-44396095和CHEMBL167790等。 非核苷抑制剂 (non-nucleoside inhibitors,NNIs) 通常表现出窄谱抗病毒活性,其与病毒聚合酶上的变构位点结合,以防止转录所需的构象变化。苏拉明 (suramin) 作为抗寄生虫药物,是一个对称的聚阴离子萘基脲,它能够有效地对抗HIV和HBV。研究表明,苏拉明能够在体外有效地抑制人和小鼠的NoV 聚合酶,IC50 值分别为24.6和70.0 nmol·L-1。同样,作为苏拉明衍生物,NF023也可抑制人和小鼠的NoV RdRp,IC50分别为71.5和200 nmol·L-1。苏拉明的另两个衍生物,NAF2和PPNDS也被证明能够抑制人诺如病毒 RdRp,IC50值分别为14和0.45 μmol·L-1,PPNDS还可抑制鼠诺如病毒 RdRp,IC50为0.88 μmol·L-1。尽管苏拉明及其衍生物的抑酶活性较好,但这些化合物细胞通透性差,因此它们细胞水平的抗病毒效力大大降低,EC50仅为0.3 μmol·L-1。为进一步提高这些化合物的抗NoV活性,已有科学家开始尝试通过脂质体输送苏拉明来改善其细胞通透性。除苏拉明及其衍生物外,最近几年一些其他非核苷抑制剂也被发现具有抗NoV的功效,如化合物JTK-109、TMC-647055、Beclabuvir和AC-1858,能有效抑制不同种属 NoV。 03蛋白酶抑制剂NoV 蛋白酶在非结构性多元蛋白前体加工过程起着重要的作用,对底物的特异性识别与切割是病毒复制过程中必不可少的过程,因此合成与 NoV 蛋白酶竞争性结合的拟肽抑制剂是设计 NoV 蛋白酶抑制剂的基本策略。芦平曲韦 (rupintrivir) 是第一代NoV蛋白酶抑制剂,最初是作为抗人鼻病毒抑制剂开发的。芦平曲韦对其他小核糖核酸病毒、冠状病毒和杯状病毒具有广谱抗病毒活性。其余的 NoV 蛋白酶抑制剂可分为过渡态模拟物和过渡态抑制剂。其中,过渡态模拟物包括α-羟基膦酸盐,它在低微摩尔浓度下能够有效地抑制 Norwalk 复制子。第一类过渡态抑制剂是一系列肽基醛,其结构含有谷氨酰胺替代物,此后,更多的衍生物被开发出来,如二肽基醛类、三肽基醛类、α-酮基酰胺肽类、α-酮杂环类、亚硫酸氢盐加合物类以及双亚硫酸盐加合物的酯或氨基甲酸酯前药。与最初的肽基醛抑制剂相比,这些衍生物都显示出同等或更高水平的抗病毒活性。 为了改善药物膜渗透性差、口服生物利用度低及抑制活性较差等问题,研究者们对大环抑制剂进行了一系列研究。大环抑制剂是基于醛类抑制剂结构改造而来,主要包括三唑类、恶二唑类和恶唑烷酮类,大环抑制剂在体内和体外同样表现出较好的 NoV抑制作用。 04作用于宿主因子的化合物随着直接抗病毒药物临床应用的增加,耐药问题日益突出,尤其是具有低耐药屏障的非核苷抑制剂和蛋白酶抑制剂。与直接抗病毒药相比,靶向宿主的抗病毒药物通常具有更高的耐药屏障,并且该类药物可以靶向与病毒直接相互作用或辅助病毒复制的关键蛋白,具有较好的抗病毒效力,成为治疗NoV感染的重要类型。 去泛素酶(deubiquitinase,DUB)是一类参与调节泛素-蛋白酶体系统的酶,对病毒复制十分重要。小分子抑制剂WP1130 是一种选择性 DUB 抑制剂,它在病毒复制水平上表现出对鼠诺如病毒和人诺如病毒的显著抗病毒活性,缺点是溶解度不高。之后,研究人员筛选了31个WP1130衍生物希望在体外保留WP1130的抗病毒活性,同时增强水溶性,改善药物特性。其中化合物7具有与WP1130 相似的广谱抗病毒能力,在稳定表达NoV复制子的 HG23细胞中EC50为3. 37 μmol/L,且相比WP1130,细胞毒 性更低,水溶性增强。此 热休克蛋白( heat-shock protein,HSP) Hsp90和Hsp70是在病毒复制中具有重要作用的热休克蛋白家族成员。研究表明,鼠诺如病毒-1和人诺如病毒衣壳蛋白的稳定性与Hsp90 活性密切相关。研究发现热应激对小鼠巨噬细胞RAW264. 7 细胞系中鼠诺如病毒的复制有促进作用,使用Hsp90抑制剂KNK437和Hsp70抑制剂17-AGG 抑制了鼠诺如病毒基因组复制和病毒颗粒的产生。此外还发现KNK437和17-AGG 可以降低鼠诺如病毒感染细胞中 IL-1β、IL-10和TNF-α mRNA的表达水平,提示Hsp90 和Hsp70 抑制剂可以作为潜在的宿主导向的NoV 抑制剂,发挥抗NoV功能。 免疫调节剂干扰素( interferon, IFN)在抵御病毒感染中起着重要作用。研究表明,I型和II 型 IFNs 及其受体能够预防小鼠和人感染 NoV。最近的研究发现III型 IFNs (IFN-λ)也有抗 NoV 感染的作用,IFN-λ可以诱导细胞表达许多由I型和II型IFNs诱导表达的相同基因。然而,与I型和II型IFNs不同的是,单剂量的IFN-λ (1 μg) 可预防和清除小鼠持续的鼠诺如病毒感染并且 IFN-λ 可靶向非造血细胞和肠上皮细胞。在鼠诺如病毒传播模型中,内源性IFN-λ可阻止感染急性CW3 鼠诺如病毒的小鼠的病毒传播,不仅如此,IFN-λ还可防止小鼠肠道CW3复制、炎症和抗体反应。这些研究表明,IFN-λ不仅具有治疗 NoV 感染的潜力,而且 IFN-λ 的预防性用药还可以预防NoV的传播。 免疫调节剂Toll 样受体 (Toll-like receptor,TLR) 激动剂具有抗 NoV 作用。这类化合物已作为疫 苗佐剂使用多年,能够抑制其他病毒如HIV和HBV的复制。TLR激动剂能够诱导先天免疫反应并刺激IFNs的产生,已知其在体外对鼠诺如病毒和人诺如病毒具有抑制作用。其中,几种TLR7激动剂显示出对鼠诺如病毒感染的有效抑制作用,包括 R-848、GS-9620、Gardiquimod和 R-837。 作为引起全球病毒性胃肠炎和腹泻的最主要病原体, NoV 在人群的流行和感染越来越频繁,受到越来越多的关注和重视。目前,还未见获批准上市的抗 NoV 药物。临床上治疗 NoV 感染的方法主要是通过补充电解质缓解病毒感染后引发的腹泻症状。因此,为了应对 NoV在全球范围内的广泛传播,开发针对 NoV 感染的有效抑制剂及其他疗法迫在眉睫。庆幸的是,近年来,针对 NoV 生活周期的各个环节,寻找潜在的抗病毒靶点和有效的抗 NoV 化合物正在有条不紊的展开。小分子抑制剂特别是针对病毒蛋白酶和病毒RdRp 的抑制剂在抵御 NoV 感染的药物研发中发展迅速。此外,宿主导向的抗病毒疗法因其耐药性低、抗病毒活性强以及对宿主伤害较小等优点,日益发展为开发新型抗 NoV 药物的重要方向。深入研究 NoV 的生活特性、完善 NoV 的复制机制、建立有效且简便的 NoV 体外和体内感染模型以及高通量药物筛选模型,是药物开发中的关键突破点,将有助于抗 NoV 感染药物的快速发现。 参考资料 [1]张磊,段梦兰,池杨. 诺如来袭,如何应对[N]. 健康报,2023-02-17(004). [2]董悦,展鹏,刘新泳.抗诺如病毒药物及其疫 苗研究新进展[J].药学学报,2020,55(04):640-651.DOI:10.16438/j.0513-4870.2019-0783. [3]王丽丹,岑山.抗诺如病毒药物研究进展[J].中华实验和临床病毒学杂志,2022,(第5期). 作者简介:小泥沙,食品科技工作者,现就职于国内某大型药物研发公司,从事营养食品及功能性食品的开发与研究。智药研习社3月直播讲座报名智药研习社3月线下研习会报名来源:CPHI制药在线声明:本文仅代表作者观点,并不代表制药在线立场。本网站内容仅出于传递更多信息之目的。如需转载,请务必注明文章来源和作者。投稿邮箱:Kelly.Xiao@imsinoexpo.com▼更多制药资讯,请关注CPHI制药在线▼点击阅读原文,进入智药研习社~
免疫疗法疫苗
100 项与 贝卡布韦 相关的药物交易
登录后查看更多信息
研发状态
10 条进展最快的记录, 后查看更多信息
登录
| 适应症 | 最高研发状态 | 国家/地区 | 公司 | 日期 |
|---|---|---|---|---|
| 慢性丙型肝炎基因型 1 | 临床3期 | 美国 | 2013-12-01 | |
| 慢性丙型肝炎基因型 1 | 临床3期 | 澳大利亚 | 2013-12-01 | |
| 慢性丙型肝炎基因型 1 | 临床3期 | 加拿大 | 2013-12-01 | |
| 慢性丙型肝炎基因型 1 | 临床3期 | 法国 | 2013-12-01 | |
| 纤维化 | 临床3期 | 美国 | 2013-12-01 | |
| 纤维化 | 临床3期 | 澳大利亚 | 2013-12-01 | |
| 纤维化 | 临床3期 | 加拿大 | 2013-12-01 | |
| 纤维化 | 临床3期 | 法国 | 2013-12-01 | |
| 慢性丙型肝炎 | 临床2期 | 波多黎各 | 2011-11-30 | |
| 丙型肝炎 | 临床1期 | 英国 | 2014-04-01 |
登录后查看更多信息
临床结果
临床结果
适应症
分期
评价
查看全部结果
| 研究 | 分期 | 人群特征 | 评价人数 | 分组 | 结果 | 评价 | 发布日期 |
|---|
No Data | |||||||
登录后查看更多信息
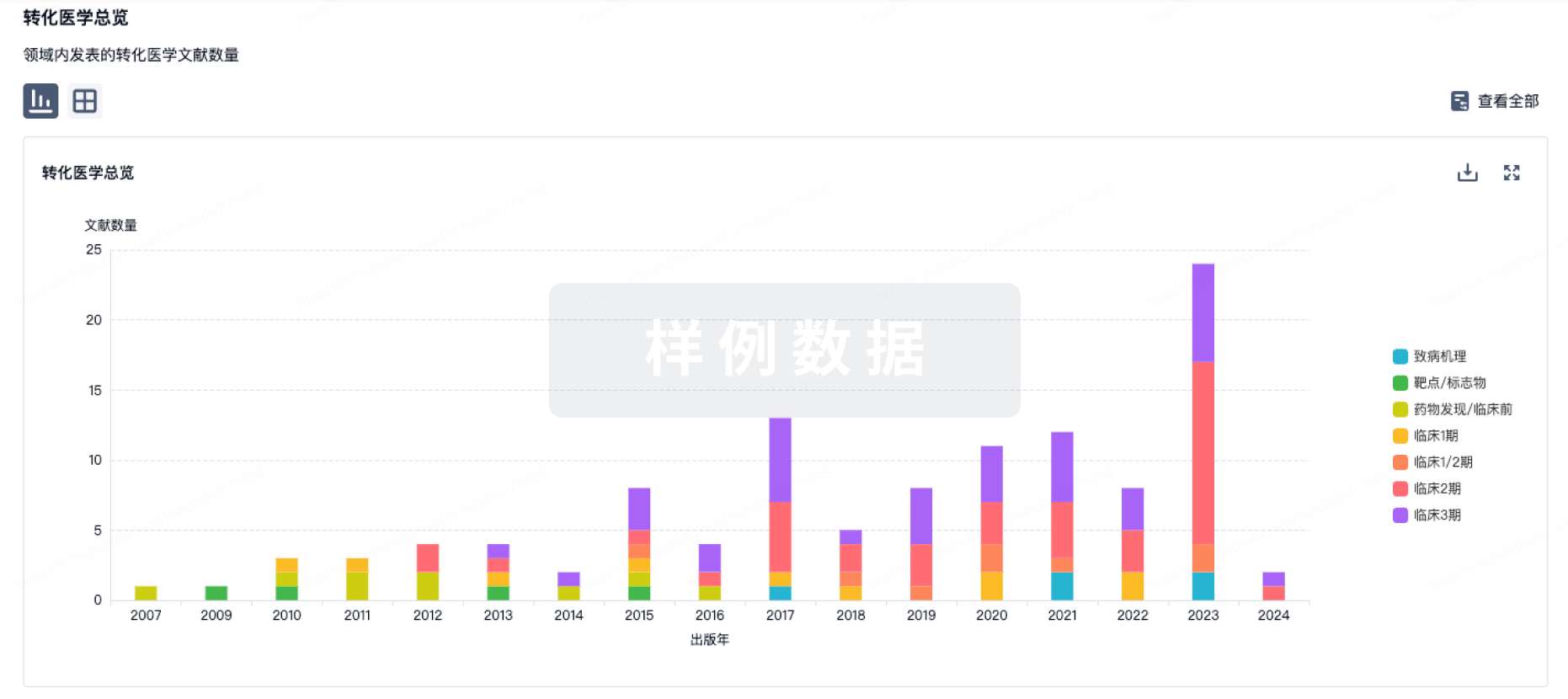
转化医学
使用我们的转化医学数据加速您的研究。
登录
或
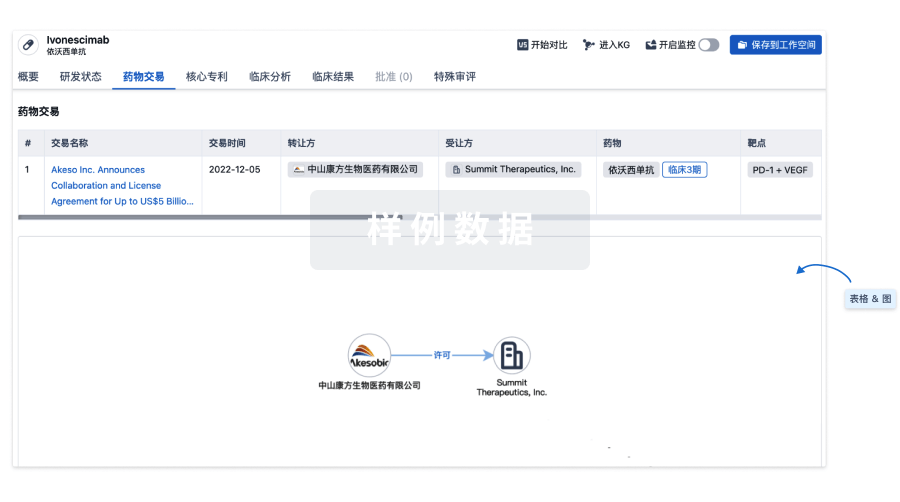
药物交易
使用我们的药物交易数据加速您的研究。
登录
或
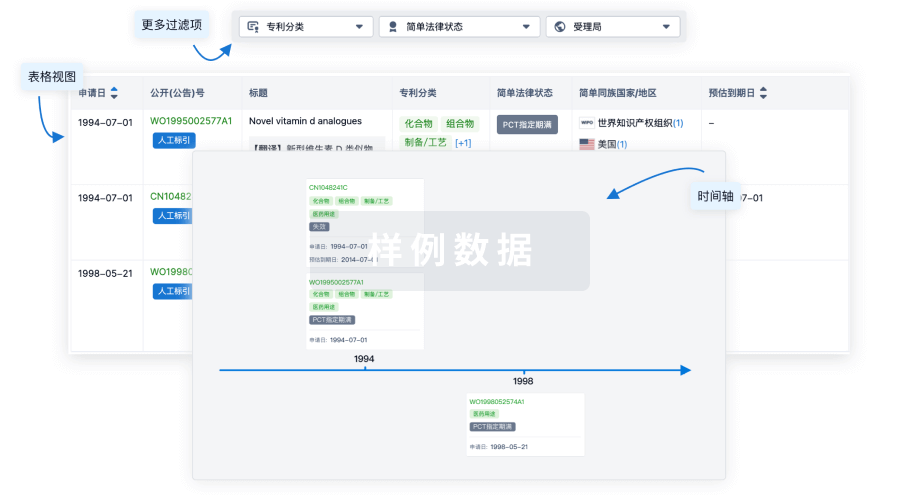
核心专利
使用我们的核心专利数据促进您的研究。
登录
或
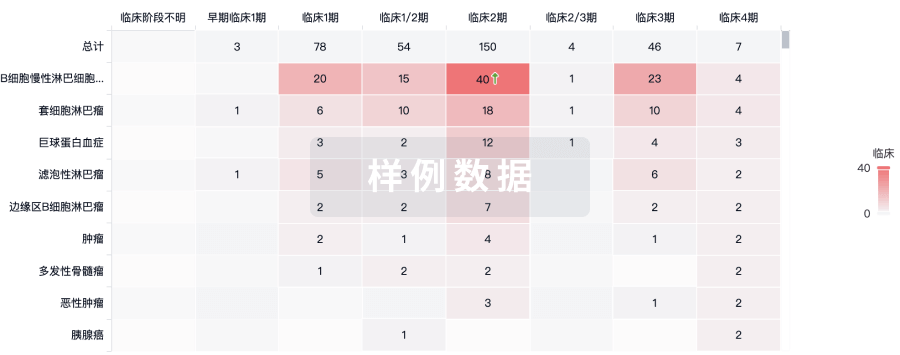
临床分析
紧跟全球注册中心的最新临床试验。
登录
或
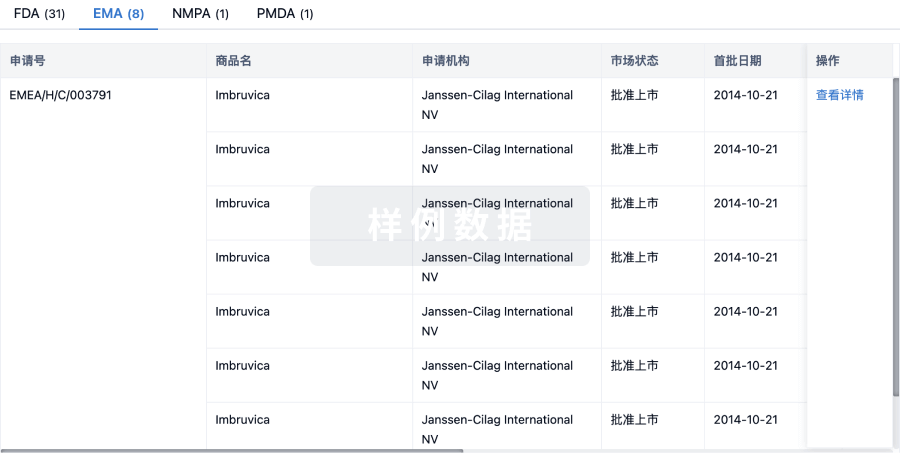
批准
利用最新的监管批准信息加速您的研究。
登录
或
特殊审评
只需点击几下即可了解关键药物信息。
登录
或
Eureka LS:
全新生物医药AI Agent 覆盖科研全链路,让突破性发现快人一步
立即开始免费试用!
智慧芽新药情报库是智慧芽专为生命科学人士构建的基于AI的创新药情报平台,助您全方位提升您的研发与决策效率。
立即开始数据试用!
智慧芽新药库数据也通过智慧芽数据服务平台,以API或者数据包形式对外开放,助您更加充分利用智慧芽新药情报信息。
生物序列数据库
生物药研发创新
免费使用
化学结构数据库
小分子化药研发创新
免费使用
